Aditya Balu
Artifact Removal and Image Restoration in AFM:A Structured Mask-Guided Directional Inpainting Approach
Feb 03, 2026Abstract:Atomic Force Microscopy (AFM) enables high-resolution surface imaging at the nanoscale, yet the output is often degraded by artifacts introduced by environmental noise, scanning imperfections, and tip-sample interactions. To address this challenge, a lightweight and fully automated framework for artifact detection and restoration in AFM image analysis is presented. The pipeline begins with a classification model that determines whether an AFM image contains artifacts. If necessary, a lightweight semantic segmentation network, custom-designed and trained on AFM data, is applied to generate precise artifact masks. These masks are adaptively expanded based on their structural orientation and then inpainted using a directional neighbor-based interpolation strategy to preserve 3D surface continuity. A localized Gaussian smoothing operation is then applied for seamless restoration. The system is integrated into a user-friendly GUI that supports real-time parameter adjustments and batch processing. Experimental results demonstrate the effective artifact removal while preserving nanoscale structural details, providing a robust, geometry-aware solution for high-fidelity AFM data interpretation.
Trustworthy LLM-Mediated Communication: Evaluating Information Fidelity in LLM as a Communicator (LAAC) Framework in Multiple Application Domains
Nov 06, 2025Abstract:The proliferation of AI-generated content has created an absurd communication theater where senders use LLMs to inflate simple ideas into verbose content, recipients use LLMs to compress them back into summaries, and as a consequence neither party engage with authentic content. LAAC (LLM as a Communicator) proposes a paradigm shift - positioning LLMs as intelligent communication intermediaries that capture the sender's intent through structured dialogue and facilitate genuine knowledge exchange with recipients. Rather than perpetuating cycles of AI-generated inflation and compression, LAAC enables authentic communication across diverse contexts including academic papers, proposals, professional emails, and cross-platform content generation. However, deploying LLMs as trusted communication intermediaries raises critical questions about information fidelity, consistency, and reliability. This position paper systematically evaluates the trustworthiness requirements for LAAC's deployment across multiple communication domains. We investigate three fundamental dimensions: (1) Information Capture Fidelity - accuracy of intent extraction during sender interviews across different communication types, (2) Reproducibility - consistency of structured knowledge across multiple interaction instances, and (3) Query Response Integrity - reliability of recipient-facing responses without hallucination, source conflation, or fabrication. Through controlled experiments spanning multiple LAAC use cases, we assess these trust dimensions using LAAC's multi-agent architecture. Preliminary findings reveal measurable trust gaps that must be addressed before LAAC can be reliably deployed in high-stakes communication scenarios.
Balancing Utility and Privacy: Dynamically Private SGD with Random Projection
Sep 11, 2025Abstract:Stochastic optimization is a pivotal enabler in modern machine learning, producing effective models for various tasks. However, several existing works have shown that model parameters and gradient information are susceptible to privacy leakage. Although Differentially Private SGD (DPSGD) addresses privacy concerns, its static noise mechanism impacts the error bounds for model performance. Additionally, with the exponential increase in model parameters, efficient learning of these models using stochastic optimizers has become more challenging. To address these concerns, we introduce the Dynamically Differentially Private Projected SGD (D2P2-SGD) optimizer. In D2P2-SGD, we combine two important ideas: (i) dynamic differential privacy (DDP) with automatic gradient clipping and (ii) random projection with SGD, allowing dynamic adjustment of the tradeoff between utility and privacy of the model. It exhibits provably sub-linear convergence rates across different objective functions, matching the best available rate. The theoretical analysis further suggests that DDP leads to better utility at the cost of privacy, while random projection enables more efficient model learning. Extensive experiments across diverse datasets show that D2P2-SGD remarkably enhances accuracy while maintaining privacy. Our code is available here.
DeCAF: Decentralized Consensus-And-Factorization for Low-Rank Adaptation of Foundation Models
May 27, 2025Abstract:Low-Rank Adaptation (LoRA) has emerged as one of the most effective, computationally tractable fine-tuning approaches for training Vision-Language Models (VLMs) and Large Language Models (LLMs). LoRA accomplishes this by freezing the pre-trained model weights and injecting trainable low-rank matrices, allowing for efficient learning of these foundation models even on edge devices. However, LoRA in decentralized settings still remains under explored, particularly for the theoretical underpinnings due to the lack of smoothness guarantee and model consensus interference (defined formally below). This work improves the convergence rate of decentralized LoRA (DLoRA) to match the rate of decentralized SGD by ensuring gradient smoothness. We also introduce DeCAF, a novel algorithm integrating DLoRA with truncated singular value decomposition (TSVD)-based matrix factorization to resolve consensus interference. Theoretical analysis shows TSVD's approximation error is bounded and consensus differences between DLoRA and DeCAF vanish as rank increases, yielding DeCAF's matching convergence rate. Extensive experiments across vision/language tasks demonstrate our algorithms outperform local training and rivals federated learning under both IID and non-IID data distributions.
Towards Large Reasoning Models for Agriculture
May 25, 2025Abstract:Agricultural decision-making involves complex, context-specific reasoning, where choices about crops, practices, and interventions depend heavily on geographic, climatic, and economic conditions. Traditional large language models (LLMs) often fall short in navigating this nuanced problem due to limited reasoning capacity. We hypothesize that recent advances in large reasoning models (LRMs) can better handle such structured, domain-specific inference. To investigate this, we introduce AgReason, the first expert-curated open-ended science benchmark with 100 questions for agricultural reasoning. Evaluations across thirteen open-source and proprietary models reveal that LRMs outperform conventional ones, though notable challenges persist, with the strongest Gemini-based baseline achieving 36% accuracy. We also present AgThoughts, a large-scale dataset of 44.6K question-answer pairs generated with human oversight and equipped with synthetically generated reasoning traces. Using AgThoughts, we develop AgThinker, a suite of small reasoning models that can be run on consumer-grade GPUs, and show that our dataset can be effective in unlocking agricultural reasoning abilities in LLMs. Our project page is here: https://baskargroup.github.io/Ag_reasoning/
WeedNet: A Foundation Model-Based Global-to-Local AI Approach for Real-Time Weed Species Identification and Classification
May 25, 2025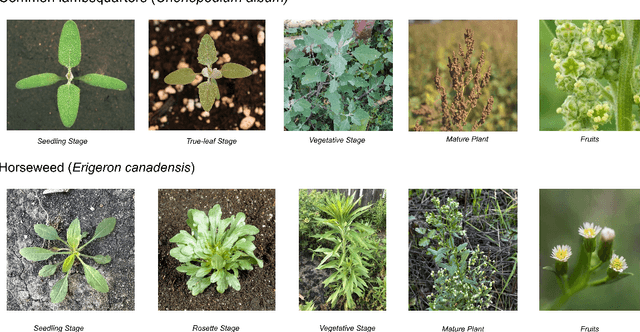
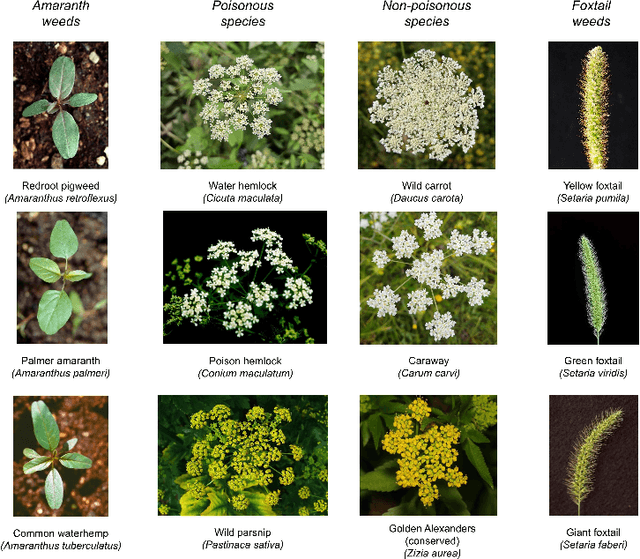
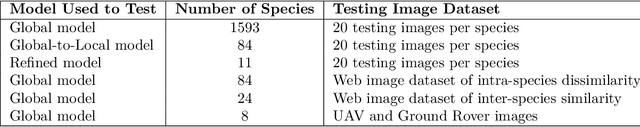
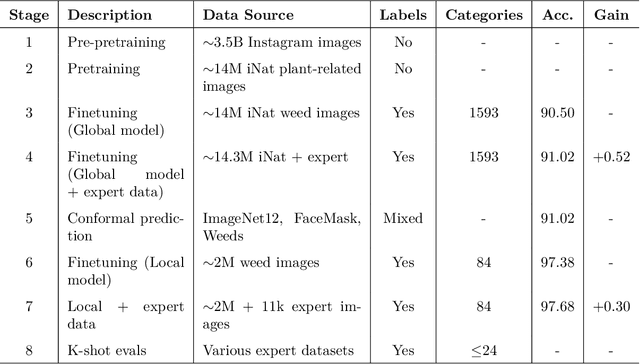


Abstract:Early identification of weeds is essential for effective management and control, and there is growing interest in automating the process using computer vision techniques coupled with AI methods. However, challenges associated with training AI-based weed identification models, such as limited expert-verified data and complexity and variability in morphological features, have hindered progress. To address these issues, we present WeedNet, the first global-scale weed identification model capable of recognizing an extensive set of weed species, including noxious and invasive plant species. WeedNet is an end-to-end real-time weed identification pipeline and uses self-supervised learning, fine-tuning, and enhanced trustworthiness strategies. WeedNet achieved 91.02% accuracy across 1,593 weed species, with 41% species achieving 100% accuracy. Using a fine-tuning strategy and a Global-to-Local approach, the local Iowa WeedNet model achieved an overall accuracy of 97.38% for 85 Iowa weeds, most classes exceeded a 90% mean accuracy per class. Testing across intra-species dissimilarity (developmental stages) and inter-species similarity (look-alike species) suggests that diversity in the images collected, spanning all the growth stages and distinguishable plant characteristics, is crucial in driving model performance. The generalizability and adaptability of the Global WeedNet model enable it to function as a foundational model, with the Global-to-Local strategy allowing fine-tuning for region-specific weed communities. Additional validation of drone- and ground-rover-based images highlights the potential of WeedNet for integration into robotic platforms. Furthermore, integration with AI for conversational use provides intelligent agricultural and ecological conservation consulting tools for farmers, agronomists, researchers, land managers, and government agencies across diverse landscapes.
Bidirectional Linear Recurrent Models for Sequence-Level Multisource Fusion
Apr 11, 2025Abstract:Sequence modeling is a critical yet challenging task with wide-ranging applications, especially in time series forecasting for domains like weather prediction, temperature monitoring, and energy load forecasting. Transformers, with their attention mechanism, have emerged as state-of-the-art due to their efficient parallel training, but they suffer from quadratic time complexity, limiting their scalability for long sequences. In contrast, recurrent neural networks (RNNs) offer linear time complexity, spurring renewed interest in linear RNNs for more computationally efficient sequence modeling. In this work, we introduce BLUR (Bidirectional Linear Unit for Recurrent network), which uses forward and backward linear recurrent units (LRUs) to capture both past and future dependencies with high computational efficiency. BLUR maintains the linear time complexity of traditional RNNs, while enabling fast parallel training through LRUs. Furthermore, it offers provably stable training and strong approximation capabilities, making it highly effective for modeling long-term dependencies. Extensive experiments on sequential image and time series datasets reveal that BLUR not only surpasses transformers and traditional RNNs in accuracy but also significantly reduces computational costs, making it particularly suitable for real-world forecasting tasks. Our code is available here.
NeRF-based Point Cloud Reconstruction using a Stationary Camera for Agricultural Applications
Mar 27, 2025Abstract:This paper presents a NeRF-based framework for point cloud (PCD) reconstruction, specifically designed for indoor high-throughput plant phenotyping facilities. Traditional NeRF-based reconstruction methods require cameras to move around stationary objects, but this approach is impractical for high-throughput environments where objects are rapidly imaged while moving on conveyors or rotating pedestals. To address this limitation, we develop a variant of NeRF-based PCD reconstruction that uses a single stationary camera to capture images as the object rotates on a pedestal. Our workflow comprises COLMAP-based pose estimation, a straightforward pose transformation to simulate camera movement, and subsequent standard NeRF training. A defined Region of Interest (ROI) excludes irrelevant scene data, enabling the generation of high-resolution point clouds (10M points). Experimental results demonstrate excellent reconstruction fidelity, with precision-recall analyses yielding an F-score close to 100.00 across all evaluated plant objects. Although pose estimation remains computationally intensive with a stationary camera setup, overall training and reconstruction times are competitive, validating the method's feasibility for practical high-throughput indoor phenotyping applications. Our findings indicate that high-quality NeRF-based 3D reconstructions are achievable using a stationary camera, eliminating the need for complex camera motion or costly imaging equipment. This approach is especially beneficial when employing expensive and delicate instruments, such as hyperspectral cameras, for 3D plant phenotyping. Future work will focus on optimizing pose estimation techniques and further streamlining the methodology to facilitate seamless integration into automated, high-throughput 3D phenotyping pipelines.
Accessing the Effect of Phyllotaxy and Planting Density on Light Use Efficiency in Field-Grown Maize using 3D Reconstructions
Mar 10, 2025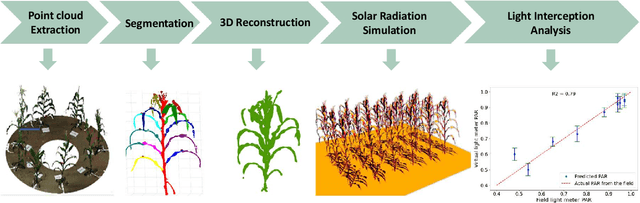
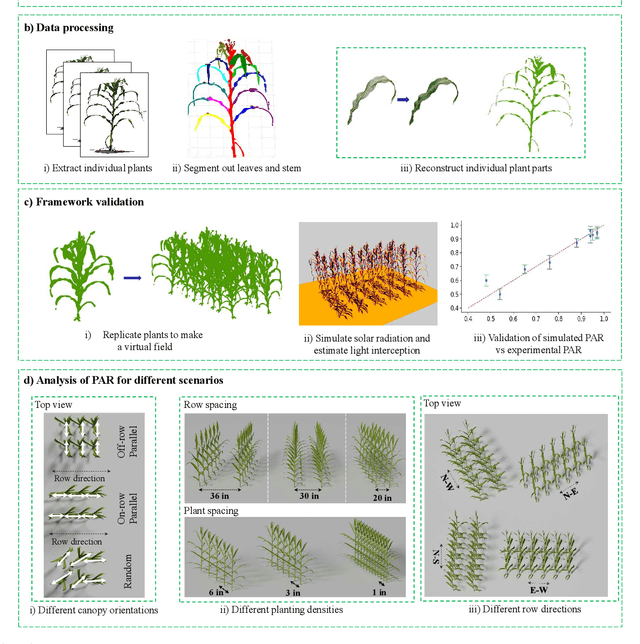
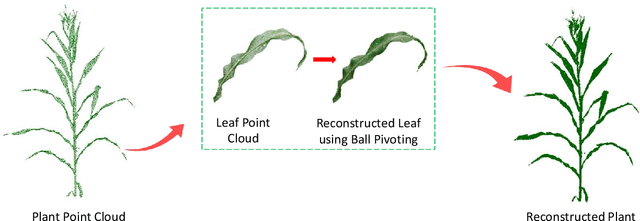
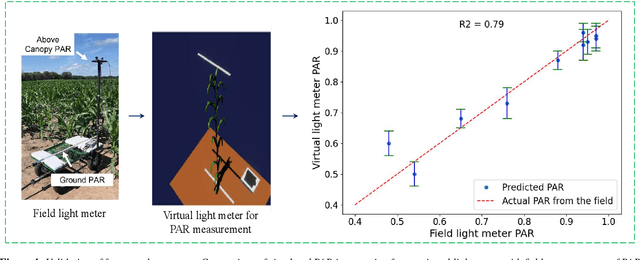
Abstract:High-density planting is a widely adopted strategy to enhance maize productivity, yet it introduces challenges such as increased interplant competition and shading, which can limit light capture and overall yield potential. In response, some maize plants naturally reorient their canopies to optimize light capture, a process known as canopy reorientation. Understanding this adaptive response and its impact on light capture is crucial for maximizing agricultural yield potential. This study introduces an end-to-end framework that integrates realistic 3D reconstructions of field-grown maize with photosynthetically active radiation (PAR) modeling to assess the effects of phyllotaxy and planting density on light interception. In particular, using 3D point clouds derived from field data, virtual fields for a diverse set of maize genotypes were constructed and validated against field PAR measurements. Using this framework, we present detailed analyses of the impact of canopy orientations, plant and row spacings, and planting row directions on PAR interception throughout a typical growing season. Our findings highlight significant variations in light interception efficiency across different planting densities and canopy orientations. By elucidating the relationship between canopy architecture and light capture, this study offers valuable guidance for optimizing maize breeding and cultivation strategies across diverse agricultural settings.
AgriField3D: A Curated 3D Point Cloud and Procedural Model Dataset of Field-Grown Maize from a Diversity Panel
Mar 10, 2025



Abstract:The application of artificial intelligence (AI) in three-dimensional (3D) agricultural research, particularly for maize, has been limited by the scarcity of large-scale, diverse datasets. While 2D image datasets are abundant, they fail to capture essential structural details such as leaf architecture, plant volume, and spatial arrangements that 3D data provide. To address this limitation, we present AgriField3D (https://baskargroup.github.io/AgriField3D/), a curated dataset of 3D point clouds of field-grown maize plants from a diverse genetic panel, designed to be AI-ready for advancing agricultural research. Our dataset comprises over 1,000 high-quality point clouds collected using a Terrestrial Laser Scanner, complemented by procedural models that provide structured, parametric representations of maize plants. These procedural models, generated using Non-Uniform Rational B-Splines (NURBS) and optimized via a two-step process combining Particle Swarm Optimization (PSO) and differentiable programming, enable precise, scalable reconstructions of leaf surfaces and plant architectures. To enhance usability, we performed graph-based segmentation to isolate individual leaves and stalks, ensuring consistent labeling across all samples. We also conducted rigorous manual quality control on all datasets, correcting errors in segmentation, ensuring accurate leaf ordering, and validating metadata annotations. The dataset further includes metadata detailing plant morphology and quality, alongside multi-resolution subsampled versions (100k, 50k, 10k points) optimized for various computational needs. By integrating point cloud data of field grown plants with high-fidelity procedural models and ensuring meticulous manual validation, AgriField3D provides a comprehensive foundation for AI-driven phenotyping, plant structural analysis, and 3D applications in agricultural research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge