Patrick S. Schnable
FloraForge: LLM-Assisted Procedural Generation of Editable and Analysis-Ready 3D Plant Geometric Models For Agricultural Applications
Dec 11, 2025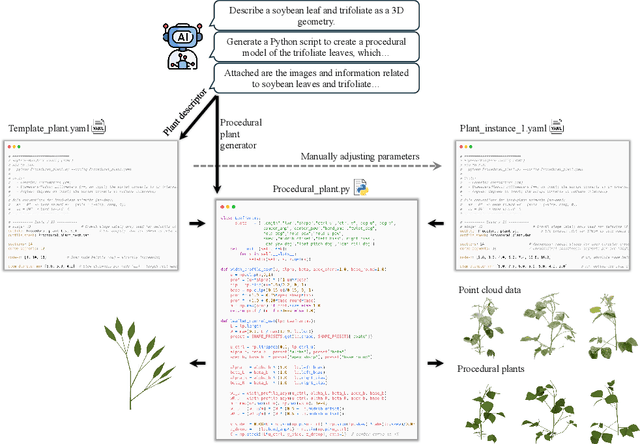
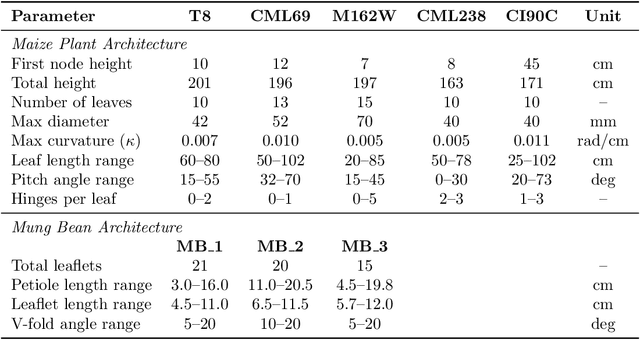
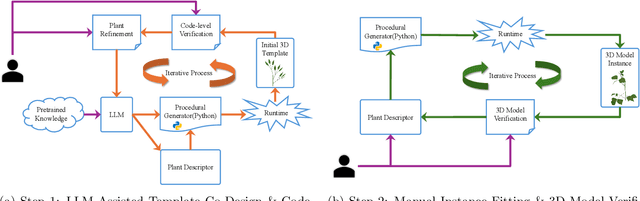
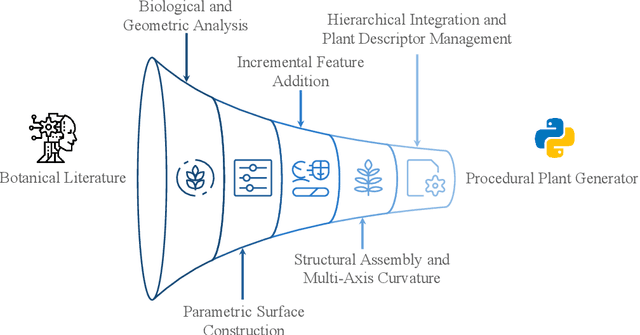
Abstract:Accurate 3D plant models are crucial for computational phenotyping and physics-based simulation; however, current approaches face significant limitations. Learning-based reconstruction methods require extensive species-specific training data and lack editability. Procedural modeling offers parametric control but demands specialized expertise in geometric modeling and an in-depth understanding of complex procedural rules, making it inaccessible to domain scientists. We present FloraForge, an LLM-assisted framework that enables domain experts to generate biologically accurate, fully parametric 3D plant models through iterative natural language Plant Refinements (PR), minimizing programming expertise. Our framework leverages LLM-enabled co-design to refine Python scripts that generate parameterized plant geometries as hierarchical B-spline surface representations with botanical constraints with explicit control points and parametric deformation functions. This representation can be easily tessellated into polygonal meshes with arbitrary precision, ensuring compatibility with functional structural plant analysis workflows such as light simulation, computational fluid dynamics, and finite element analysis. We demonstrate the framework on maize, soybean, and mung bean, fitting procedural models to empirical point cloud data through manual refinement of the Plant Descriptor (PD), human-readable files. The pipeline generates dual outputs: triangular meshes for visualization and triangular meshes with additional parametric metadata for quantitative analysis. This approach uniquely combines LLM-assisted template creation, mathematically continuous representations enabling both phenotyping and rendering, and direct parametric control through PD. The framework democratizes sophisticated geometric modeling for plant science while maintaining mathematical rigor.
Accessing the Effect of Phyllotaxy and Planting Density on Light Use Efficiency in Field-Grown Maize using 3D Reconstructions
Mar 10, 2025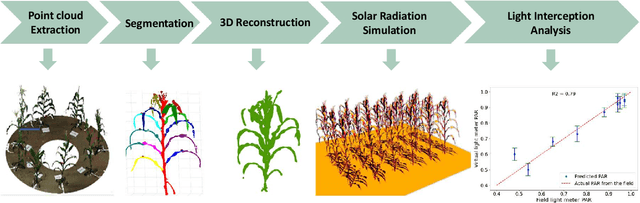
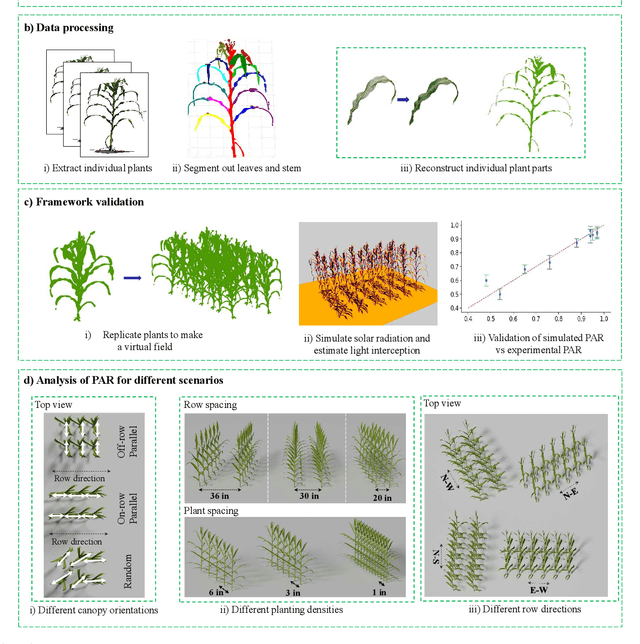
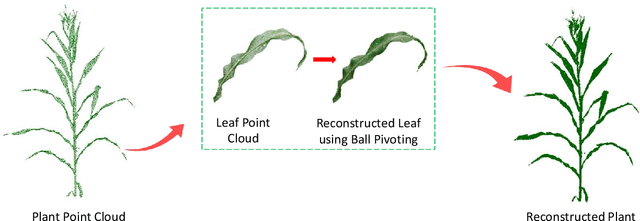
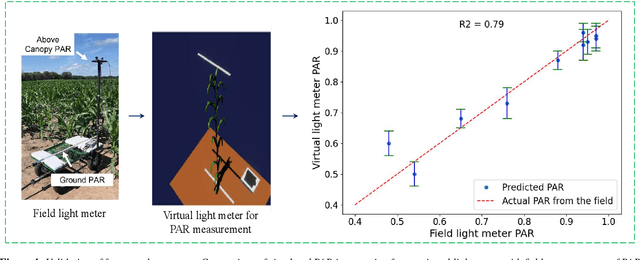
Abstract:High-density planting is a widely adopted strategy to enhance maize productivity, yet it introduces challenges such as increased interplant competition and shading, which can limit light capture and overall yield potential. In response, some maize plants naturally reorient their canopies to optimize light capture, a process known as canopy reorientation. Understanding this adaptive response and its impact on light capture is crucial for maximizing agricultural yield potential. This study introduces an end-to-end framework that integrates realistic 3D reconstructions of field-grown maize with photosynthetically active radiation (PAR) modeling to assess the effects of phyllotaxy and planting density on light interception. In particular, using 3D point clouds derived from field data, virtual fields for a diverse set of maize genotypes were constructed and validated against field PAR measurements. Using this framework, we present detailed analyses of the impact of canopy orientations, plant and row spacings, and planting row directions on PAR interception throughout a typical growing season. Our findings highlight significant variations in light interception efficiency across different planting densities and canopy orientations. By elucidating the relationship between canopy architecture and light capture, this study offers valuable guidance for optimizing maize breeding and cultivation strategies across diverse agricultural settings.
AgriField3D: A Curated 3D Point Cloud and Procedural Model Dataset of Field-Grown Maize from a Diversity Panel
Mar 10, 2025



Abstract:The application of artificial intelligence (AI) in three-dimensional (3D) agricultural research, particularly for maize, has been limited by the scarcity of large-scale, diverse datasets. While 2D image datasets are abundant, they fail to capture essential structural details such as leaf architecture, plant volume, and spatial arrangements that 3D data provide. To address this limitation, we present AgriField3D (https://baskargroup.github.io/AgriField3D/), a curated dataset of 3D point clouds of field-grown maize plants from a diverse genetic panel, designed to be AI-ready for advancing agricultural research. Our dataset comprises over 1,000 high-quality point clouds collected using a Terrestrial Laser Scanner, complemented by procedural models that provide structured, parametric representations of maize plants. These procedural models, generated using Non-Uniform Rational B-Splines (NURBS) and optimized via a two-step process combining Particle Swarm Optimization (PSO) and differentiable programming, enable precise, scalable reconstructions of leaf surfaces and plant architectures. To enhance usability, we performed graph-based segmentation to isolate individual leaves and stalks, ensuring consistent labeling across all samples. We also conducted rigorous manual quality control on all datasets, correcting errors in segmentation, ensuring accurate leaf ordering, and validating metadata annotations. The dataset further includes metadata detailing plant morphology and quality, alongside multi-resolution subsampled versions (100k, 50k, 10k points) optimized for various computational needs. By integrating point cloud data of field grown plants with high-fidelity procedural models and ensuring meticulous manual validation, AgriField3D provides a comprehensive foundation for AI-driven phenotyping, plant structural analysis, and 3D applications in agricultural research.
MaizeEar-SAM: Zero-Shot Maize Ear Phenotyping
Feb 19, 2025Abstract:Quantifying the variation in yield component traits of maize (Zea mays L.), which together determine the overall productivity of this globally important crop, plays a critical role in plant genetics research, plant breeding, and the development of improved farming practices. Grain yield per acre is calculated by multiplying the number of plants per acre, ears per plant, number of kernels per ear, and the average kernel weight. The number of kernels per ear is determined by the number of kernel rows per ear multiplied by the number of kernels per row. Traditional manual methods for measuring these two traits are time-consuming, limiting large-scale data collection. Recent automation efforts using image processing and deep learning encounter challenges such as high annotation costs and uncertain generalizability. We tackle these issues by exploring Large Vision Models for zero-shot, annotation-free maize kernel segmentation. By using an open-source large vision model, the Segment Anything Model (SAM), we segment individual kernels in RGB images of maize ears and apply a graph-based algorithm to calculate the number of kernels per row. Our approach successfully identifies the number of kernels per row across a wide range of maize ears, showing the potential of zero-shot learning with foundation vision models combined with image processing techniques to improve automation and reduce subjectivity in agronomic data collection. All our code is open-sourced to make these affordable phenotyping methods accessible to everyone.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge