Zhaojie Fang
DGSAN: Dual-Graph Spatiotemporal Attention Network for Pulmonary Nodule Malignancy Prediction
Dec 24, 2025Abstract:Lung cancer continues to be the leading cause of cancer-related deaths globally. Early detection and diagnosis of pulmonary nodules are essential for improving patient survival rates. Although previous research has integrated multimodal and multi-temporal information, outperforming single modality and single time point, the fusion methods are limited to inefficient vector concatenation and simple mutual attention, highlighting the need for more effective multimodal information fusion. To address these challenges, we introduce a Dual-Graph Spatiotemporal Attention Network, which leverages temporal variations and multimodal data to enhance the accuracy of predictions. Our methodology involves developing a Global-Local Feature Encoder to better capture the local, global, and fused characteristics of pulmonary nodules. Additionally, a Dual-Graph Construction method organizes multimodal features into inter-modal and intra-modal graphs. Furthermore, a Hierarchical Cross-Modal Graph Fusion Module is introduced to refine feature integration. We also compiled a novel multimodal dataset named the NLST-cmst dataset as a comprehensive source of support for related research. Our extensive experiments, conducted on both the NLST-cmst and curated CSTL-derived datasets, demonstrate that our DGSAN significantly outperforms state-of-the-art methods in classifying pulmonary nodules with exceptional computational efficiency.
LPUWF-LDM: Enhanced Latent Diffusion Model for Precise Late-phase UWF-FA Generation on Limited Dataset
Sep 01, 2024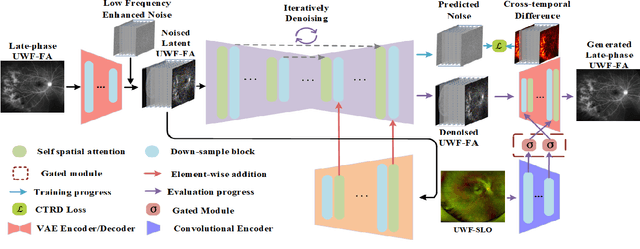
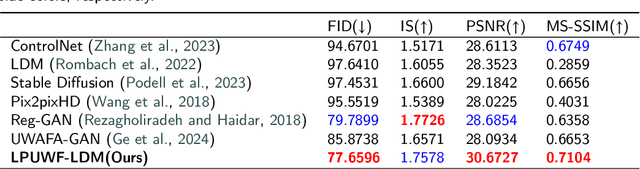
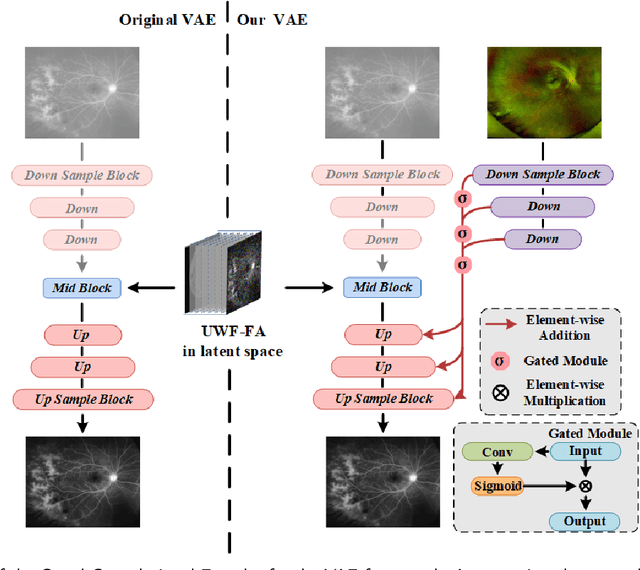

Abstract:Ultra-Wide-Field Fluorescein Angiography (UWF-FA) enables precise identification of ocular diseases using sodium fluorescein, which can be potentially harmful. Existing research has developed methods to generate UWF-FA from Ultra-Wide-Field Scanning Laser Ophthalmoscopy (UWF-SLO) to reduce the adverse reactions associated with injections. However, these methods have been less effective in producing high-quality late-phase UWF-FA, particularly in lesion areas and fine details. Two primary challenges hinder the generation of high-quality late-phase UWF-FA: the scarcity of paired UWF-SLO and early/late-phase UWF-FA datasets, and the need for realistic generation at lesion sites and potential blood leakage regions. This study introduces an improved latent diffusion model framework to generate high-quality late-phase UWF-FA from limited paired UWF images. To address the challenges as mentioned earlier, our approach employs a module utilizing Cross-temporal Regional Difference Loss, which encourages the model to focus on the differences between early and late phases. Additionally, we introduce a low-frequency enhanced noise strategy in the diffusion forward process to improve the realism of medical images. To further enhance the mapping capability of the variational autoencoder module, especially with limited datasets, we implement a Gated Convolutional Encoder to extract additional information from conditional images. Our Latent Diffusion Model for Ultra-Wide-Field Late-Phase Fluorescein Angiography (LPUWF-LDM) effectively reconstructs fine details in late-phase UWF-FA and achieves state-of-the-art results compared to other existing methods when working with limited datasets. Our source code is available at: https://github.com/Tinysqua/****.
Toward Robust Early Detection of Alzheimer's Disease via an Integrated Multimodal Learning Approach
Aug 29, 2024


Abstract:Alzheimer's Disease (AD) is a complex neurodegenerative disorder marked by memory loss, executive dysfunction, and personality changes. Early diagnosis is challenging due to subtle symptoms and varied presentations, often leading to misdiagnosis with traditional unimodal diagnostic methods due to their limited scope. This study introduces an advanced multimodal classification model that integrates clinical, cognitive, neuroimaging, and EEG data to enhance diagnostic accuracy. The model incorporates a feature tagger with a tabular data coding architecture and utilizes the TimesBlock module to capture intricate temporal patterns in Electroencephalograms (EEG) data. By employing Cross-modal Attention Aggregation module, the model effectively fuses Magnetic Resonance Imaging (MRI) spatial information with EEG temporal data, significantly improving the distinction between AD, Mild Cognitive Impairment, and Normal Cognition. Simultaneously, we have constructed the first AD classification dataset that includes three modalities: EEG, MRI, and tabular data. Our innovative approach aims to facilitate early diagnosis and intervention, potentially slowing the progression of AD. The source code and our private ADMC dataset are available at https://github.com/JustlfC03/MSTNet.
GFE-Mamba: Mamba-based AD Multi-modal Progression Assessment via Generative Feature Extraction from MCI
Jul 22, 2024Abstract:Alzheimer's Disease (AD) is an irreversible neurodegenerative disorder that often progresses from Mild Cognitive Impairment (MCI), leading to memory loss and significantly impacting patients' lives. Clinical trials indicate that early targeted interventions for MCI patients can potentially slow or halt the development and progression of AD. Previous research has shown that accurate medical classification requires the inclusion of extensive multimodal data, such as assessment scales and various neuroimaging techniques like Magnetic Resonance Imaging (MRI) and Positron Emission Tomography (PET). However, consistently tracking the diagnosis of the same individual over time and simultaneously collecting multimodal data poses significant challenges. To address this issue, we introduce GFE-Mamba, a classifier based on Generative Feature Extraction (GFE). This classifier effectively integrates data from assessment scales, MRI, and PET, enabling deeper multimodal fusion. It efficiently extracts both long and short sequence information and incorporates additional information beyond the pixel space. This approach not only improves classification accuracy but also enhances the interpretability and stability of the model. We constructed datasets of over 3000 samples based on the Alzheimer's Disease Neuroimaging Initiative (ADNI) for a two-step training process. Our experimental results demonstrate that the GFE-Mamba model is effective in predicting the conversion from MCI to AD and outperforms several state-of-the-art methods. Our source code and ADNI dataset processing code are available at https://github.com/Tinysqua/GFE-Mamba.
SCKansformer: Fine-Grained Classification of Bone Marrow Cells via Kansformer Backbone and Hierarchical Attention Mechanisms
Jun 14, 2024



Abstract:The incidence and mortality rates of malignant tumors, such as acute leukemia, have risen significantly. Clinically, hospitals rely on cytological examination of peripheral blood and bone marrow smears to diagnose malignant tumors, with accurate blood cell counting being crucial. Existing automated methods face challenges such as low feature expression capability, poor interpretability, and redundant feature extraction when processing high-dimensional microimage data. We propose a novel fine-grained classification model, SCKansformer, for bone marrow blood cells, which addresses these challenges and enhances classification accuracy and efficiency. The model integrates the Kansformer Encoder, SCConv Encoder, and Global-Local Attention Encoder. The Kansformer Encoder replaces the traditional MLP layer with the KAN, improving nonlinear feature representation and interpretability. The SCConv Encoder, with its Spatial and Channel Reconstruction Units, enhances feature representation and reduces redundancy. The Global-Local Attention Encoder combines Multi-head Self-Attention with a Local Part module to capture both global and local features. We validated our model using the Bone Marrow Blood Cell Fine-Grained Classification Dataset (BMCD-FGCD), comprising over 10,000 samples and nearly 40 classifications, developed with a partner hospital. Comparative experiments on our private dataset, as well as the publicly available PBC and ALL-IDB datasets, demonstrate that SCKansformer outperforms both typical and advanced microcell classification methods across all datasets. Our source code and private BMCD-FGCD dataset are available at https://github.com/JustlfC03/SCKansformer.
UWAFA-GAN: Ultra-Wide-Angle Fluorescein Angiography Transformation via Multi-scale Generation and Registration Enhancement
May 01, 2024



Abstract:Fundus photography, in combination with the ultra-wide-angle fundus (UWF) techniques, becomes an indispensable diagnostic tool in clinical settings by offering a more comprehensive view of the retina. Nonetheless, UWF fluorescein angiography (UWF-FA) necessitates the administration of a fluorescent dye via injection into the patient's hand or elbow unlike UWF scanning laser ophthalmoscopy (UWF-SLO). To mitigate potential adverse effects associated with injections, researchers have proposed the development of cross-modality medical image generation algorithms capable of converting UWF-SLO images into their UWF-FA counterparts. Current image generation techniques applied to fundus photography encounter difficulties in producing high-resolution retinal images, particularly in capturing minute vascular lesions. To address these issues, we introduce a novel conditional generative adversarial network (UWAFA-GAN) to synthesize UWF-FA from UWF-SLO. This approach employs multi-scale generators and an attention transmit module to efficiently extract both global structures and local lesions. Additionally, to counteract the image blurriness issue that arises from training with misaligned data, a registration module is integrated within this framework. Our method performs non-trivially on inception scores and details generation. Clinical user studies further indicate that the UWF-FA images generated by UWAFA-GAN are clinically comparable to authentic images in terms of diagnostic reliability. Empirical evaluations on our proprietary UWF image datasets elucidate that UWAFA-GAN outperforms extant methodologies. The code is accessible at https://github.com/Tinysqua/UWAFA-GAN.
UWAT-GAN: Fundus Fluorescein Angiography Synthesis via Ultra-wide-angle Transformation Multi-scale GAN
Jul 21, 2023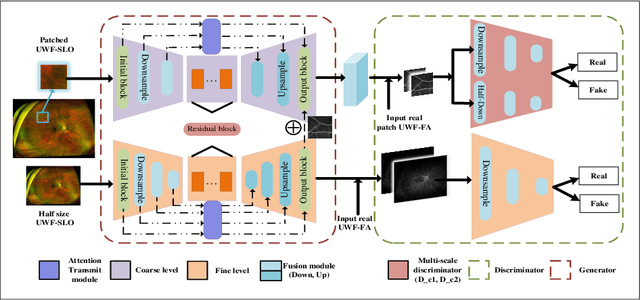



Abstract:Fundus photography is an essential examination for clinical and differential diagnosis of fundus diseases. Recently, Ultra-Wide-angle Fundus (UWF) techniques, UWF Fluorescein Angiography (UWF-FA) and UWF Scanning Laser Ophthalmoscopy (UWF-SLO) have been gradually put into use. However, Fluorescein Angiography (FA) and UWF-FA require injecting sodium fluorescein which may have detrimental influences. To avoid negative impacts, cross-modality medical image generation algorithms have been proposed. Nevertheless, current methods in fundus imaging could not produce high-resolution images and are unable to capture tiny vascular lesion areas. This paper proposes a novel conditional generative adversarial network (UWAT-GAN) to synthesize UWF-FA from UWF-SLO. Using multi-scale generators and a fusion module patch to better extract global and local information, our model can generate high-resolution images. Moreover, an attention transmit module is proposed to help the decoder learn effectively. Besides, a supervised approach is used to train the network using multiple new weighted losses on different scales of data. Experiments on an in-house UWF image dataset demonstrate the superiority of the UWAT-GAN over the state-of-the-art methods. The source code is available at: https://github.com/Tinysqua/UWAT-GAN.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge