Shenghao Zhu
No Modality Left Behind: Adapting to Missing Modalities via Knowledge Distillation for Brain Tumor Segmentation
Sep 18, 2025
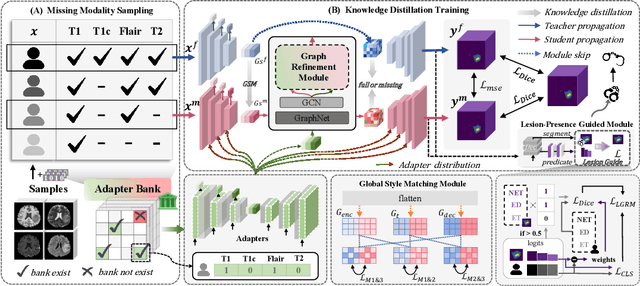
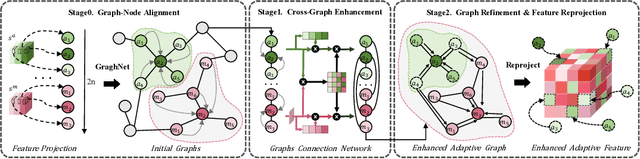
Abstract:Accurate brain tumor segmentation is essential for preoperative evaluation and personalized treatment. Multi-modal MRI is widely used due to its ability to capture complementary tumor features across different sequences. However, in clinical practice, missing modalities are common, limiting the robustness and generalizability of existing deep learning methods that rely on complete inputs, especially under non-dominant modality combinations. To address this, we propose AdaMM, a multi-modal brain tumor segmentation framework tailored for missing-modality scenarios, centered on knowledge distillation and composed of three synergistic modules. The Graph-guided Adaptive Refinement Module explicitly models semantic associations between generalizable and modality-specific features, enhancing adaptability to modality absence. The Bi-Bottleneck Distillation Module transfers structural and textural knowledge from teacher to student models via global style matching and adversarial feature alignment. The Lesion-Presence-Guided Reliability Module predicts prior probabilities of lesion types through an auxiliary classification task, effectively suppressing false positives under incomplete inputs. Extensive experiments on the BraTS 2018 and 2024 datasets demonstrate that AdaMM consistently outperforms existing methods, exhibiting superior segmentation accuracy and robustness, particularly in single-modality and weak-modality configurations. In addition, we conduct a systematic evaluation of six categories of missing-modality strategies, confirming the superiority of knowledge distillation and offering practical guidance for method selection and future research. Our source code is available at https://github.com/Quanato607/AdaMM.
Bridging the Gap in Missing Modalities: Leveraging Knowledge Distillation and Style Matching for Brain Tumor Segmentation
Jul 30, 2025Abstract:Accurate and reliable brain tumor segmentation, particularly when dealing with missing modalities, remains a critical challenge in medical image analysis. Previous studies have not fully resolved the challenges of tumor boundary segmentation insensitivity and feature transfer in the absence of key imaging modalities. In this study, we introduce MST-KDNet, aimed at addressing these critical issues. Our model features Multi-Scale Transformer Knowledge Distillation to effectively capture attention weights at various resolutions, Dual-Mode Logit Distillation to improve the transfer of knowledge, and a Global Style Matching Module that integrates feature matching with adversarial learning. Comprehensive experiments conducted on the BraTS and FeTS 2024 datasets demonstrate that MST-KDNet surpasses current leading methods in both Dice and HD95 scores, particularly in conditions with substantial modality loss. Our approach shows exceptional robustness and generalization potential, making it a promising candidate for real-world clinical applications. Our source code is available at https://github.com/Quanato607/MST-KDNet.
* 11 pages, 2 figures
XLSTM-HVED: Cross-Modal Brain Tumor Segmentation and MRI Reconstruction Method Using Vision XLSTM and Heteromodal Variational Encoder-Decoder
Dec 09, 2024



Abstract:Neurogliomas are among the most aggressive forms of cancer, presenting considerable challenges in both treatment and monitoring due to their unpredictable biological behavior. Magnetic resonance imaging (MRI) is currently the preferred method for diagnosing and monitoring gliomas. However, the lack of specific imaging techniques often compromises the accuracy of tumor segmentation during the imaging process. To address this issue, we introduce the XLSTM-HVED model. This model integrates a hetero-modal encoder-decoder framework with the Vision XLSTM module to reconstruct missing MRI modalities. By deeply fusing spatial and temporal features, it enhances tumor segmentation performance. The key innovation of our approach is the Self-Attention Variational Encoder (SAVE) module, which improves the integration of modal features. Additionally, it optimizes the interaction of features between segmentation and reconstruction tasks through the Squeeze-Fusion-Excitation Cross Awareness (SFECA) module. Our experiments using the BraTS 2024 dataset demonstrate that our model significantly outperforms existing advanced methods in handling cases where modalities are missing. Our source code is available at https://github.com/Quanato607/XLSTM-HVED.
Toward Robust Early Detection of Alzheimer's Disease via an Integrated Multimodal Learning Approach
Aug 29, 2024


Abstract:Alzheimer's Disease (AD) is a complex neurodegenerative disorder marked by memory loss, executive dysfunction, and personality changes. Early diagnosis is challenging due to subtle symptoms and varied presentations, often leading to misdiagnosis with traditional unimodal diagnostic methods due to their limited scope. This study introduces an advanced multimodal classification model that integrates clinical, cognitive, neuroimaging, and EEG data to enhance diagnostic accuracy. The model incorporates a feature tagger with a tabular data coding architecture and utilizes the TimesBlock module to capture intricate temporal patterns in Electroencephalograms (EEG) data. By employing Cross-modal Attention Aggregation module, the model effectively fuses Magnetic Resonance Imaging (MRI) spatial information with EEG temporal data, significantly improving the distinction between AD, Mild Cognitive Impairment, and Normal Cognition. Simultaneously, we have constructed the first AD classification dataset that includes three modalities: EEG, MRI, and tabular data. Our innovative approach aims to facilitate early diagnosis and intervention, potentially slowing the progression of AD. The source code and our private ADMC dataset are available at https://github.com/JustlfC03/MSTNet.
TC-KANRecon: High-Quality and Accelerated MRI Reconstruction via Adaptive KAN Mechanisms and Intelligent Feature Scaling
Aug 11, 2024



Abstract:Magnetic Resonance Imaging (MRI) has become essential in clinical diagnosis due to its high resolution and multiple contrast mechanisms. However, the relatively long acquisition time limits its broader application. To address this issue, this study presents an innovative conditional guided diffusion model, named as TC-KANRecon, which incorporates the Multi-Free U-KAN (MF-UKAN) module and a dynamic clipping strategy. TC-KANRecon model aims to accelerate the MRI reconstruction process through deep learning methods while maintaining the quality of the reconstructed images. The MF-UKAN module can effectively balance the tradeoff between image denoising and structure preservation. Specifically, it presents the multi-head attention mechanisms and scalar modulation factors, which significantly enhances the model's robustness and structure preservation capabilities in complex noise environments. Moreover, the dynamic clipping strategy in TC-KANRecon adjusts the cropping interval according to the sampling steps, thereby mitigating image detail loss typically caused by traditional cropping methods and enriching the visual features of the images. Furthermore, the MC-Model module incorporates full-sampling k-space information, realizing efficient fusion of conditional information, enhancing the model's ability to process complex data, and improving the realism and detail richness of reconstructed images. Experimental results demonstrate that the proposed method outperforms other MRI reconstruction methods in both qualitative and quantitative evaluations. Notably, TC-KANRecon method exhibits excellent reconstruction results when processing high-noise, low-sampling-rate MRI data. Our source code is available at https://github.com/lcbkmm/TC-KANRecon.
GFE-Mamba: Mamba-based AD Multi-modal Progression Assessment via Generative Feature Extraction from MCI
Jul 22, 2024Abstract:Alzheimer's Disease (AD) is an irreversible neurodegenerative disorder that often progresses from Mild Cognitive Impairment (MCI), leading to memory loss and significantly impacting patients' lives. Clinical trials indicate that early targeted interventions for MCI patients can potentially slow or halt the development and progression of AD. Previous research has shown that accurate medical classification requires the inclusion of extensive multimodal data, such as assessment scales and various neuroimaging techniques like Magnetic Resonance Imaging (MRI) and Positron Emission Tomography (PET). However, consistently tracking the diagnosis of the same individual over time and simultaneously collecting multimodal data poses significant challenges. To address this issue, we introduce GFE-Mamba, a classifier based on Generative Feature Extraction (GFE). This classifier effectively integrates data from assessment scales, MRI, and PET, enabling deeper multimodal fusion. It efficiently extracts both long and short sequence information and incorporates additional information beyond the pixel space. This approach not only improves classification accuracy but also enhances the interpretability and stability of the model. We constructed datasets of over 3000 samples based on the Alzheimer's Disease Neuroimaging Initiative (ADNI) for a two-step training process. Our experimental results demonstrate that the GFE-Mamba model is effective in predicting the conversion from MCI to AD and outperforms several state-of-the-art methods. Our source code and ADNI dataset processing code are available at https://github.com/Tinysqua/GFE-Mamba.
SCKansformer: Fine-Grained Classification of Bone Marrow Cells via Kansformer Backbone and Hierarchical Attention Mechanisms
Jun 14, 2024



Abstract:The incidence and mortality rates of malignant tumors, such as acute leukemia, have risen significantly. Clinically, hospitals rely on cytological examination of peripheral blood and bone marrow smears to diagnose malignant tumors, with accurate blood cell counting being crucial. Existing automated methods face challenges such as low feature expression capability, poor interpretability, and redundant feature extraction when processing high-dimensional microimage data. We propose a novel fine-grained classification model, SCKansformer, for bone marrow blood cells, which addresses these challenges and enhances classification accuracy and efficiency. The model integrates the Kansformer Encoder, SCConv Encoder, and Global-Local Attention Encoder. The Kansformer Encoder replaces the traditional MLP layer with the KAN, improving nonlinear feature representation and interpretability. The SCConv Encoder, with its Spatial and Channel Reconstruction Units, enhances feature representation and reduces redundancy. The Global-Local Attention Encoder combines Multi-head Self-Attention with a Local Part module to capture both global and local features. We validated our model using the Bone Marrow Blood Cell Fine-Grained Classification Dataset (BMCD-FGCD), comprising over 10,000 samples and nearly 40 classifications, developed with a partner hospital. Comparative experiments on our private dataset, as well as the publicly available PBC and ALL-IDB datasets, demonstrate that SCKansformer outperforms both typical and advanced microcell classification methods across all datasets. Our source code and private BMCD-FGCD dataset are available at https://github.com/JustlfC03/SCKansformer.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge