Zaifu Zhan
PRIME: Prototype-Driven Multimodal Pretraining for Cancer Prognosis with Missing Modalities
Apr 05, 2026Abstract:Multimodal self-supervised pretraining offers a promising route to cancer prognosis by integrating histopathology whole-slide images, gene expression, and pathology reports, yet most existing approaches require fully paired and complete inputs. In practice, clinical cohorts are fragmented and often miss one or more modalities, limiting both supervised fusion and scalable multimodal pretraining. We propose PRIME, a missing-aware multimodal self-supervised pretraining framework that learns robust and transferable representations from partially observed cohorts. PRIME maps heterogeneous modality embeddings into a unified token space and introduces a shared prototype memory bank for latent-space semantic imputation via patient-level consensus retrieval, producing structurally aligned tokens without reconstructing raw signals. Two complementary pretraining objectives: inter-modality alignment and post-fusion consistency under structured missingness augmentation, jointly learn representations that remain predictive under arbitrary modality subsets. We evaluate PRIME on The Cancer Genome Atlas with label-free pretraining on 32 cancer types and downstream 5-fold evaluation on five cohorts across overall survival prediction, 3-year mortality classification, and 3-year recurrence classification. PRIME achieves the best macro-average performance among all compared methods, reaching 0.653 C-index, 0.689 AUROC, and 0.637 AUROC on the three tasks, respectively, while improving robustness under test-time missingness and supporting parameter-efficient and label-efficient adaptation. These results support missing-aware multimodal pretraining as a practical strategy for prognosis modeling in fragmented clinical data settings.
Can Large Language Models Self-Correct in Medical Question Answering? An Exploratory Study
Apr 02, 2026Abstract:Large language models (LLMs) have achieved strong performance on medical question answering (medical QA), and chain-of-thought (CoT) prompting has further improved results by eliciting explicit intermediate reasoning; meanwhile, self-reflective (self-corrective) prompting has been widely claimed to enhance model reliability by prompting LLMs to critique and revise their own reasoning, yet its effectiveness in safety-critical medical settings remains unclear. In this work, we conduct an exploratory analysis of self-reflective reasoning for medical multiple-choice question answering: using GPT-4o and GPT-4o-mini, we compare standard CoT prompting with an iterative self-reflection loop and track how predictions evolve across reflection steps on three widely used medical QA benchmarks (MedQA, HeadQA, and PubMedQA). We analyze whether self-reflection leads to error correction, error persistence, or the introduction of new errors. Our results show that self-reflective prompting does not consistently improve accuracy and its impact is highly dataset- and model-dependent: it yields modest gains on MedQA but provides limited or negative benefits on HeadQA and PubMedQA, and increasing the number of reflection steps does not guarantee better performance. These findings highlight a gap between reasoning transparency and reasoning correctness, suggesting that self-reflective reasoning is better viewed as an analytical tool for understanding model behavior rather than a standalone solution for improving medical QA reliability.
EpiScreen: Early Epilepsy Detection from Electronic Health Records with Large Language Models
Mar 30, 2026Abstract:Epilepsy and psychogenic non-epileptic seizures often present with similar seizure-like manifestations but require fundamentally different management strategies. Misdiagnosis is common and can lead to prolonged diagnostic delays, unnecessary treatments, and substantial patient morbidity. Although prolonged video-electroencephalography is the diagnostic gold standard, its high cost and limited accessibility hinder timely diagnosis. Here, we developed a low-cost, effective approach, EpiScreen, for early epilepsy detection by utilizing routinely collected clinical notes from electronic health records. Through fine-tuning large language models on labeled notes, EpiScreen achieved an AUC of up to 0.875 on the MIMIC-IV dataset and 0.980 on a private cohort of the University of Minnesota. In a clinician-AI collaboration setting, EpiScreen-assisted neurologists outperformed unaided experts by up to 10.9%. Overall, this study demonstrates that EpiScreen supports early epilepsy detection, facilitating timely and cost-effective screening that may reduce diagnostic delays and avoid unnecessary interventions, particularly in resource-limited regions.
MedCL-Bench: Benchmarking stability-efficiency trade-offs and scaling in biomedical continual learning
Mar 17, 2026Abstract:Medical language models must be updated as evidence and terminology evolve, yet sequential updating can trigger catastrophic forgetting. Although biomedical NLP has many static benchmarks, no unified, task-diverse benchmark exists for evaluating continual learning under standardized protocols, robustness to task order and compute-aware reporting. We introduce MedCL-Bench, which streams ten biomedical NLP datasets spanning five task families and evaluates eleven continual learning strategies across eight task orders, reporting retention, transfer, and GPU-hour cost. Across backbones and task orders, direct sequential fine-tuning on incoming tasks induces catastrophic forgetting, causing update-induced performance regressions on prior tasks. Continual learning methods occupy distinct retention-compute frontiers: parameter-isolation provides the best retention per GPU-hour, replay offers strong protection at higher cost, and regularization yields limited benefit. Forgetting is task-dependent, with multi-label topic classification most vulnerable and constrained-output tasks more robust. MedCL-Bench provides a reproducible framework for auditing model updates before deployment.
HeartAgent: An Autonomous Agent System for Explainable Differential Diagnosis in Cardiology
Mar 11, 2026Abstract:Heart diseases remain a leading cause of morbidity and mortality worldwide, necessitating accurate and trustworthy differential diagnosis. However, existing artificial intelligence-based diagnostic methods are often limited by insufficient cardiology knowledge, inadequate support for complex reasoning, and poor interpretability. Here we present HeartAgent, a cardiology-specific agent system designed to support a reliable and explainable differential diagnosis. HeartAgent integrates customized tools and curated data resources and orchestrates multiple specialized sub-agents to perform complex reasoning while generating transparent reasoning trajectories and verifiable supporting references. Evaluated on the MIMIC dataset and a private electronic health records cohort, HeartAgent achieved over 36% and 20% improvements over established comparative methods, in top-3 diagnostic accuracy, respectively. Additionally, clinicians assisted by HeartAgent demonstrated gains of 26.9% in diagnostic accuracy and 22.7% in explanatory quality compared with unaided experts. These results demonstrate that HeartAgent provides reliable, explainable, and clinically actionable decision support for cardiovascular care.
To Reason or Not to: Selective Chain-of-Thought in Medical Question Answering
Feb 23, 2026Abstract:Objective: To improve the efficiency of medical question answering (MedQA) with large language models (LLMs) by avoiding unnecessary reasoning while maintaining accuracy. Methods: We propose Selective Chain-of-Thought (Selective CoT), an inference-time strategy that first predicts whether a question requires reasoning and generates a rationale only when needed. Two open-source LLMs (Llama-3.1-8B and Qwen-2.5-7B) were evaluated on four biomedical QA benchmarks-HeadQA, MedQA-USMLE, MedMCQA, and PubMedQA. Metrics included accuracy, total generated tokens, and inference time. Results: Selective CoT reduced inference time by 13-45% and token usage by 8-47% with minimal accuracy loss ($\leq$4\%). In some model-task pairs, it achieved both higher accuracy and greater efficiency than standard CoT. Compared with fixed-length CoT, Selective CoT reached similar or superior accuracy at substantially lower computational cost. Discussion: Selective CoT dynamically balances reasoning depth and efficiency by invoking explicit reasoning only when beneficial, reducing redundancy on recall-type questions while preserving interpretability. Conclusion: Selective CoT provides a simple, model-agnostic, and cost-effective approach for medical QA, aligning reasoning effort with question complexity to enhance real-world deployability of LLM-based clinical systems.
Data-Efficient Biomedical In-Context Learning: A Diversity-Enhanced Submodular Perspective
Aug 11, 2025Abstract:Recent progress in large language models (LLMs) has leveraged their in-context learning (ICL) abilities to enable quick adaptation to unseen biomedical NLP tasks. By incorporating only a few input-output examples into prompts, LLMs can rapidly perform these new tasks. While the impact of these demonstrations on LLM performance has been extensively studied, most existing approaches prioritize representativeness over diversity when selecting examples from large corpora. To address this gap, we propose Dual-Div, a diversity-enhanced data-efficient framework for demonstration selection in biomedical ICL. Dual-Div employs a two-stage retrieval and ranking process: First, it identifies a limited set of candidate examples from a corpus by optimizing both representativeness and diversity (with optional annotation for unlabeled data). Second, it ranks these candidates against test queries to select the most relevant and non-redundant demonstrations. Evaluated on three biomedical NLP tasks (named entity recognition (NER), relation extraction (RE), and text classification (TC)) using LLaMA 3.1 and Qwen 2.5 for inference, along with three retrievers (BGE-Large, BMRetriever, MedCPT), Dual-Div consistently outperforms baselines-achieving up to 5% higher macro-F1 scores-while demonstrating robustness to prompt permutations and class imbalance. Our findings establish that diversity in initial retrieval is more critical than ranking-stage optimization, and limiting demonstrations to 3-5 examples maximizes performance efficiency.
Automating Expert-Level Medical Reasoning Evaluation of Large Language Models
Jul 10, 2025Abstract:As large language models (LLMs) become increasingly integrated into clinical decision-making, ensuring transparent and trustworthy reasoning is essential. However, existing evaluation strategies of LLMs' medical reasoning capability either suffer from unsatisfactory assessment or poor scalability, and a rigorous benchmark remains lacking. To address this, we introduce MedThink-Bench, a benchmark designed for rigorous, explainable, and scalable assessment of LLMs' medical reasoning. MedThink-Bench comprises 500 challenging questions across ten medical domains, each annotated with expert-crafted step-by-step rationales. Building on this, we propose LLM-w-Ref, a novel evaluation framework that leverages fine-grained rationales and LLM-as-a-Judge mechanisms to assess intermediate reasoning with expert-level fidelity while maintaining scalability. Experiments show that LLM-w-Ref exhibits a strong positive correlation with expert judgments. Benchmarking twelve state-of-the-art LLMs, we find that smaller models (e.g., MedGemma-27B) can surpass larger proprietary counterparts (e.g., OpenAI-o3). Overall, MedThink-Bench offers a foundational tool for evaluating LLMs' medical reasoning, advancing their safe and responsible deployment in clinical practice.
Uncertainty-Aware Large Language Models for Explainable Disease Diagnosis
May 06, 2025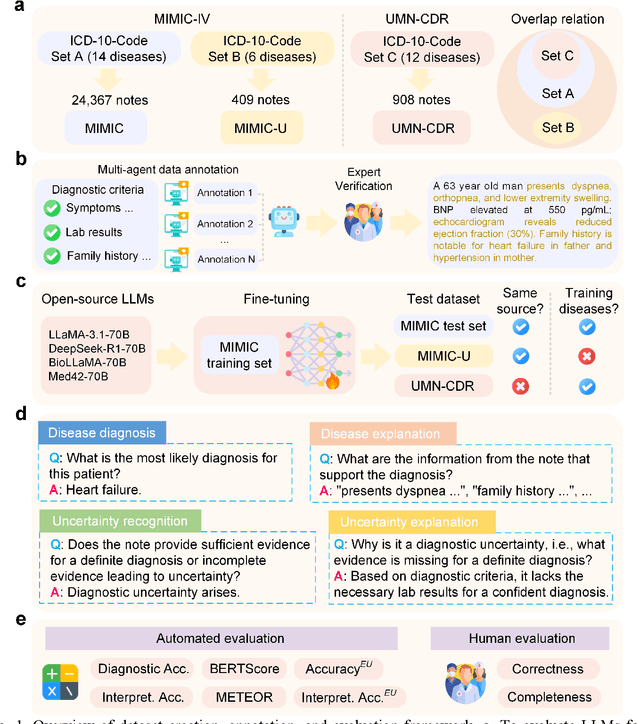
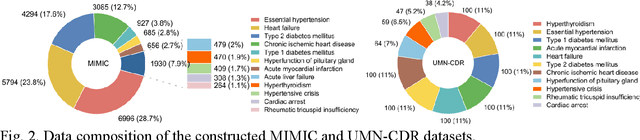
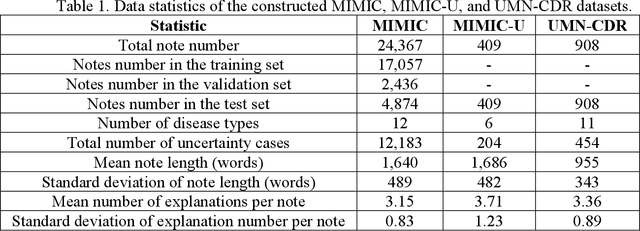

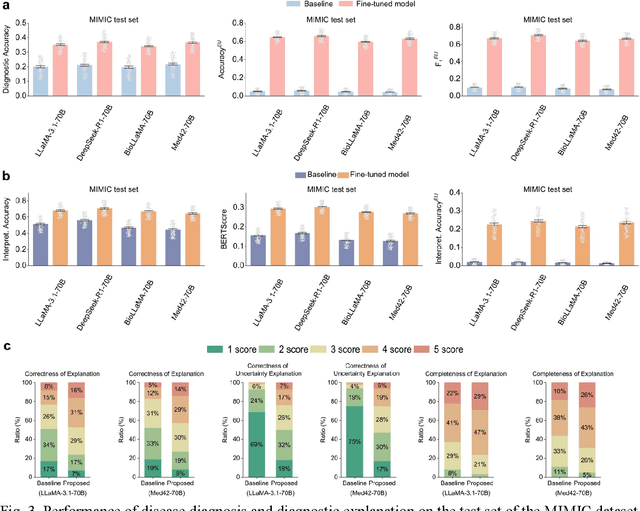
Abstract:Explainable disease diagnosis, which leverages patient information (e.g., signs and symptoms) and computational models to generate probable diagnoses and reasonings, offers clear clinical values. However, when clinical notes encompass insufficient evidence for a definite diagnosis, such as the absence of definitive symptoms, diagnostic uncertainty usually arises, increasing the risk of misdiagnosis and adverse outcomes. Although explicitly identifying and explaining diagnostic uncertainties is essential for trustworthy diagnostic systems, it remains under-explored. To fill this gap, we introduce ConfiDx, an uncertainty-aware large language model (LLM) created by fine-tuning open-source LLMs with diagnostic criteria. We formalized the task and assembled richly annotated datasets that capture varying degrees of diagnostic ambiguity. Evaluating ConfiDx on real-world datasets demonstrated that it excelled in identifying diagnostic uncertainties, achieving superior diagnostic performance, and generating trustworthy explanations for diagnoses and uncertainties. To our knowledge, this is the first study to jointly address diagnostic uncertainty recognition and explanation, substantially enhancing the reliability of automatic diagnostic systems.
Retrieval-augmented in-context learning for multimodal large language models in disease classification
May 04, 2025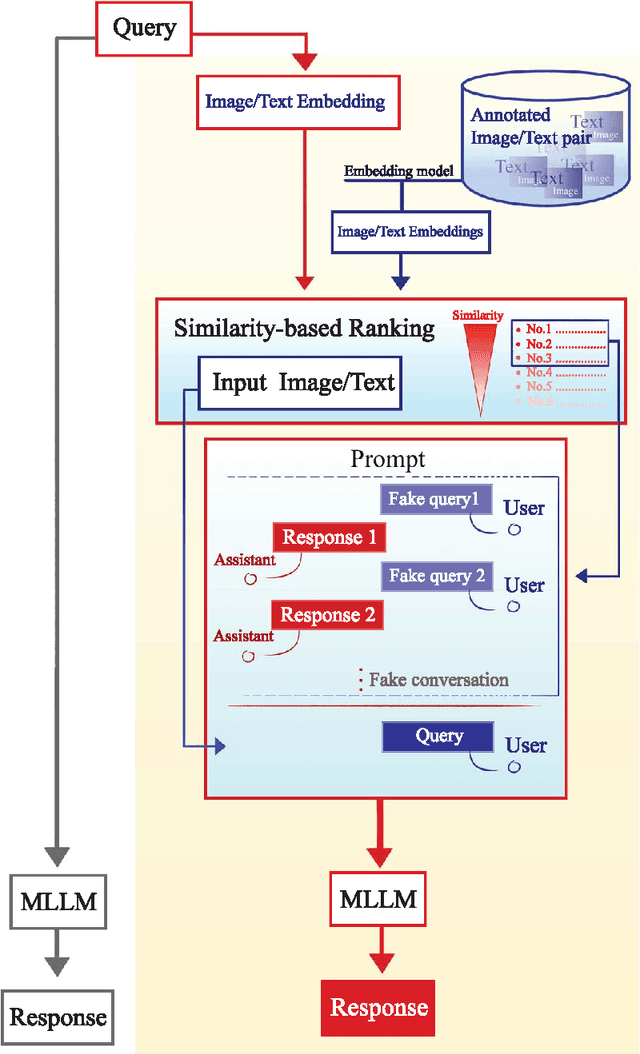
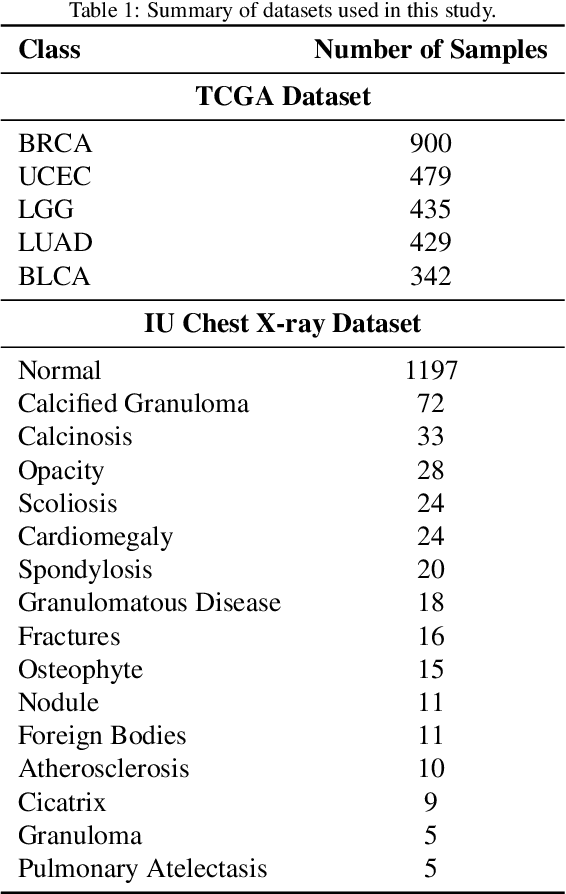
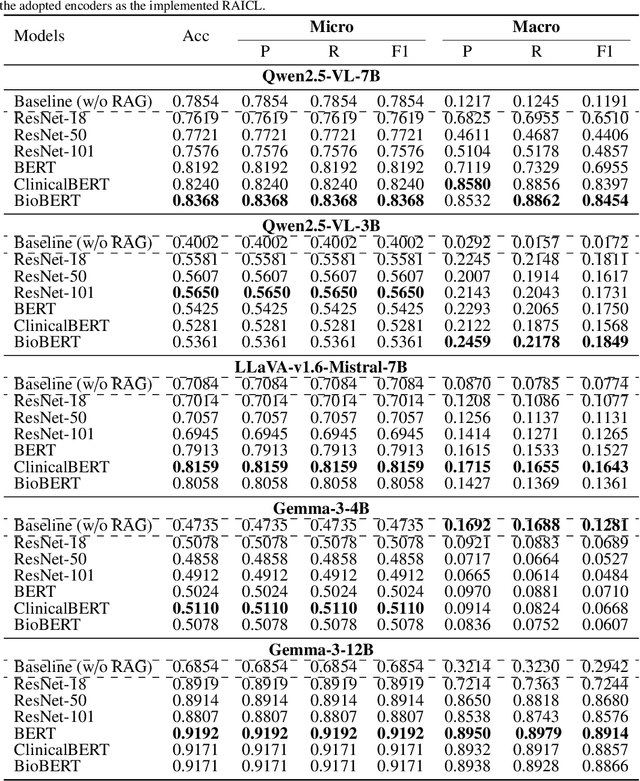
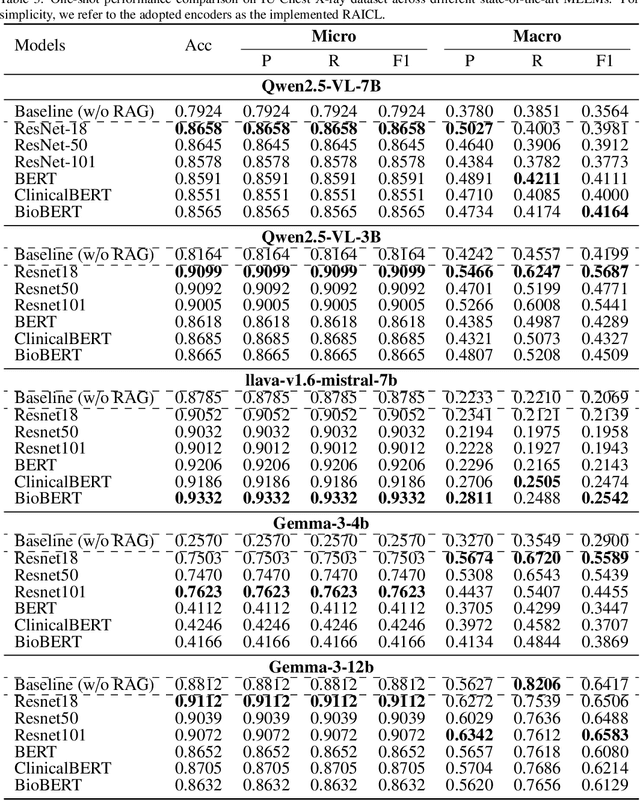
Abstract:Objectives: We aim to dynamically retrieve informative demonstrations, enhancing in-context learning in multimodal large language models (MLLMs) for disease classification. Methods: We propose a Retrieval-Augmented In-Context Learning (RAICL) framework, which integrates retrieval-augmented generation (RAG) and in-context learning (ICL) to adaptively select demonstrations with similar disease patterns, enabling more effective ICL in MLLMs. Specifically, RAICL examines embeddings from diverse encoders, including ResNet, BERT, BioBERT, and ClinicalBERT, to retrieve appropriate demonstrations, and constructs conversational prompts optimized for ICL. We evaluated the framework on two real-world multi-modal datasets (TCGA and IU Chest X-ray), assessing its performance across multiple MLLMs (Qwen, Llava, Gemma), embedding strategies, similarity metrics, and varying numbers of demonstrations. Results: RAICL consistently improved classification performance. Accuracy increased from 0.7854 to 0.8368 on TCGA and from 0.7924 to 0.8658 on IU Chest X-ray. Multi-modal inputs outperformed single-modal ones, with text-only inputs being stronger than images alone. The richness of information embedded in each modality will determine which embedding model can be used to get better results. Few-shot experiments showed that increasing the number of retrieved examples further enhanced performance. Across different similarity metrics, Euclidean distance achieved the highest accuracy while cosine similarity yielded better macro-F1 scores. RAICL demonstrated consistent improvements across various MLLMs, confirming its robustness and versatility. Conclusions: RAICL provides an efficient and scalable approach to enhance in-context learning in MLLMs for multimodal disease classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge