Mingze Yin
A Generalist Cross-Domain Molecular Learning Framework for Structure-Based Drug Discovery
Mar 06, 2025Abstract:Structure-based drug discovery (SBDD) is a systematic scientific process that develops new drugs by leveraging the detailed physical structure of the target protein. Recent advancements in pre-trained models for biomolecules have demonstrated remarkable success across various biochemical applications, including drug discovery and protein engineering. However, in most approaches, the pre-trained models primarily focus on the characteristics of either small molecules or proteins, without delving into their binding interactions which are essential cross-domain relationships pivotal to SBDD. To fill this gap, we propose a general-purpose foundation model named BIT (an abbreviation for Biomolecular Interaction Transformer), which is capable of encoding a range of biochemical entities, including small molecules, proteins, and protein-ligand complexes, as well as various data formats, encompassing both 2D and 3D structures. Specifically, we introduce Mixture-of-Domain-Experts (MoDE) to handle the biomolecules from diverse biochemical domains and Mixture-of-Structure-Experts (MoSE) to capture positional dependencies in the molecular structures. The proposed mixture-of-experts approach enables BIT to achieve both deep fusion and domain-specific encoding, effectively capturing fine-grained molecular interactions within protein-ligand complexes. Then, we perform cross-domain pre-training on the shared Transformer backbone via several unified self-supervised denoising tasks. Experimental results on various benchmarks demonstrate that BIT achieves exceptional performance in downstream tasks, including binding affinity prediction, structure-based virtual screening, and molecular property prediction.
ProtCLIP: Function-Informed Protein Multi-Modal Learning
Dec 28, 2024Abstract:Multi-modality pre-training paradigm that aligns protein sequences and biological descriptions has learned general protein representations and achieved promising performance in various downstream applications. However, these works were still unable to replicate the extraordinary success of language-supervised visual foundation models due to the ineffective usage of aligned protein-text paired data and the lack of an effective function-informed pre-training paradigm. To address these issues, this paper curates a large-scale protein-text paired dataset called ProtAnno with a property-driven sampling strategy, and introduces a novel function-informed protein pre-training paradigm. Specifically, the sampling strategy determines selecting probability based on the sample confidence and property coverage, balancing the data quality and data quantity in face of large-scale noisy data. Furthermore, motivated by significance of the protein specific functional mechanism, the proposed paradigm explicitly model protein static and dynamic functional segments by two segment-wise pre-training objectives, injecting fine-grained information in a function-informed manner. Leveraging all these innovations, we develop ProtCLIP, a multi-modality foundation model that comprehensively represents function-aware protein embeddings. On 22 different protein benchmarks within 5 types, including protein functionality classification, mutation effect prediction, cross-modal transformation, semantic similarity inference and protein-protein interaction prediction, our ProtCLIP consistently achieves SOTA performance, with remarkable improvements of 75% on average in five cross-modal transformation benchmarks, 59.9% in GO-CC and 39.7% in GO-BP protein function prediction. The experimental results verify the extraordinary potential of ProtCLIP serving as the protein multi-modality foundation model.
S$^2$ALM: Sequence-Structure Pre-trained Large Language Model for Comprehensive Antibody Representation Learning
Nov 20, 2024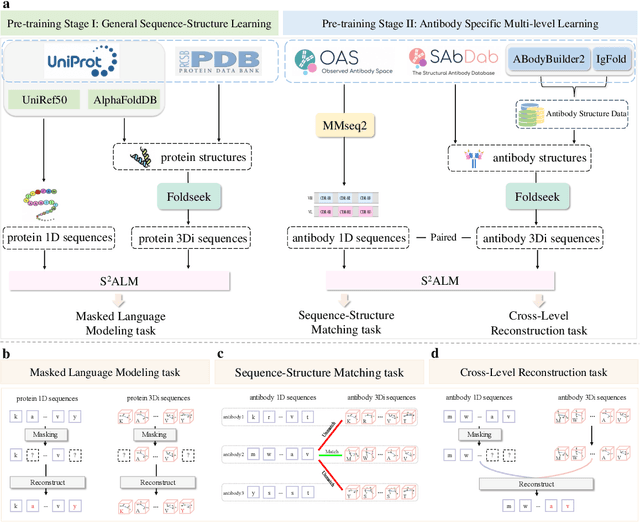
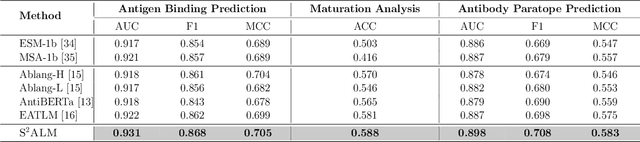
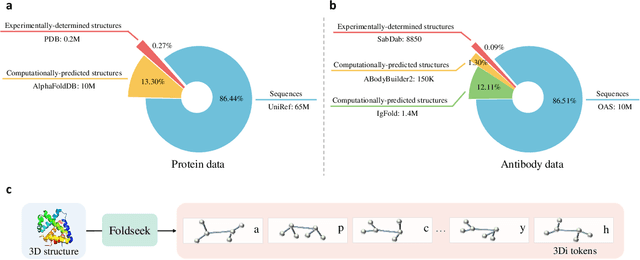
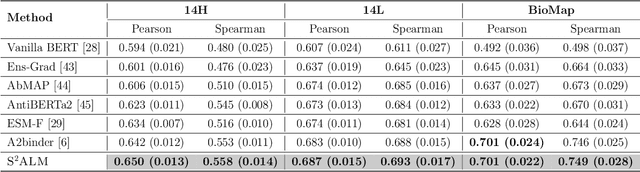
Abstract:Antibodies safeguard our health through their precise and potent binding to specific antigens, demonstrating promising therapeutic efficacy in the treatment of numerous diseases, including COVID-19. Recent advancements in biomedical language models have shown the great potential to interpret complex biological structures and functions. However, existing antibody specific models have a notable limitation that they lack explicit consideration for antibody structural information, despite the fact that both 1D sequence and 3D structure carry unique and complementary insights into antibody behavior and functionality. This paper proposes Sequence-Structure multi-level pre-trained Antibody Language Model (S$^2$ALM), combining holistic sequential and structural information in one unified, generic antibody foundation model. We construct a hierarchical pre-training paradigm incorporated with two customized multi-level training objectives to facilitate the modeling of comprehensive antibody representations. S$^2$ALM's representation space uncovers inherent functional binding mechanisms, biological evolution properties and structural interaction patterns. Pre-trained over 75 million sequences and 11.7 million structures, S$^2$ALM can be adopted for diverse downstream tasks: accurately predicting antigen-antibody binding affinities, precisely distinguishing B cell maturation stages, identifying antibody crucial binding positions, and specifically designing novel coronavirus-binding antibodies. Remarkably, S$^2$ALM outperforms well-established and renowned baselines and sets new state-of-the-art performance across extensive antibody specific understanding and generation tasks. S$^2$ALM's ability to model comprehensive and generalized representations further positions its potential to advance real-world therapeutic antibody development, potentially addressing unmet academic, industrial, and clinical needs.
Bridge-IF: Learning Inverse Protein Folding with Markov Bridges
Nov 04, 2024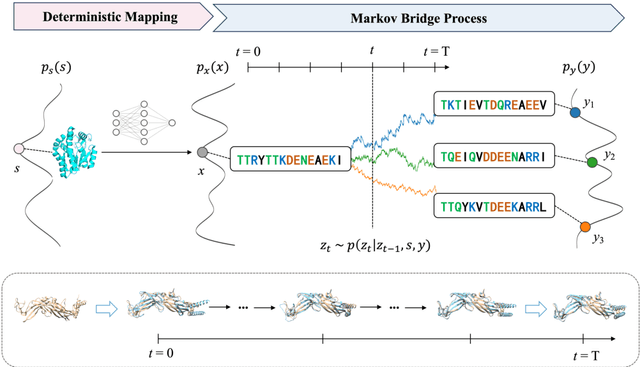
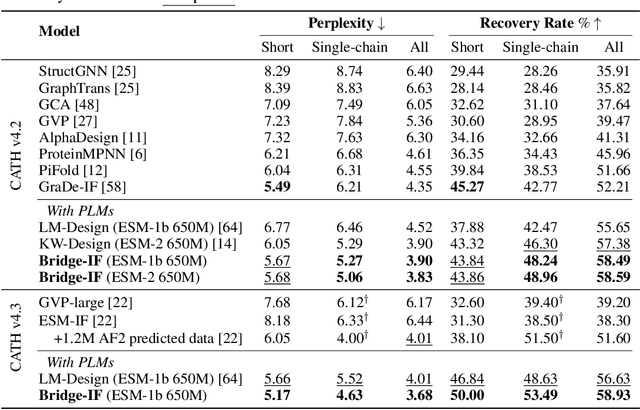
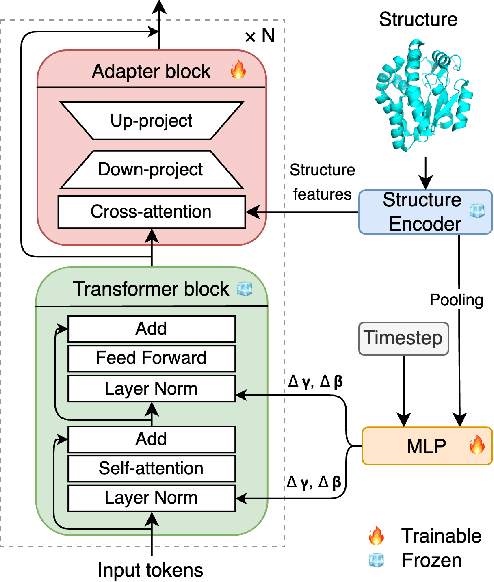
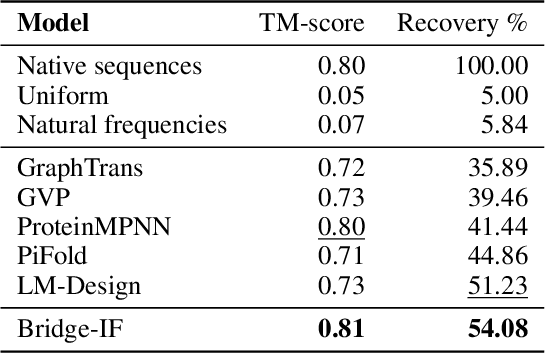
Abstract:Inverse protein folding is a fundamental task in computational protein design, which aims to design protein sequences that fold into the desired backbone structures. While the development of machine learning algorithms for this task has seen significant success, the prevailing approaches, which predominantly employ a discriminative formulation, frequently encounter the error accumulation issue and often fail to capture the extensive variety of plausible sequences. To fill these gaps, we propose Bridge-IF, a generative diffusion bridge model for inverse folding, which is designed to learn the probabilistic dependency between the distributions of backbone structures and protein sequences. Specifically, we harness an expressive structure encoder to propose a discrete, informative prior derived from structures, and establish a Markov bridge to connect this prior with native sequences. During the inference stage, Bridge-IF progressively refines the prior sequence, culminating in a more plausible design. Moreover, we introduce a reparameterization perspective on Markov bridge models, from which we derive a simplified loss function that facilitates more effective training. We also modulate protein language models (PLMs) with structural conditions to precisely approximate the Markov bridge process, thereby significantly enhancing generation performance while maintaining parameter-efficient training. Extensive experiments on well-established benchmarks demonstrate that Bridge-IF predominantly surpasses existing baselines in sequence recovery and excels in the design of plausible proteins with high foldability. The code is available at https://github.com/violet-sto/Bridge-IF.
Structure-Enhanced Protein Instruction Tuning: Towards General-Purpose Protein Understanding
Oct 04, 2024Abstract:Proteins, as essential biomolecules, play a central role in biological processes, including metabolic reactions and DNA replication. Accurate prediction of their properties and functions is crucial in biological applications. Recent development of protein language models (pLMs) with supervised fine tuning provides a promising solution to this problem. However, the fine-tuned model is tailored for particular downstream prediction task, and achieving general-purpose protein understanding remains a challenge. In this paper, we introduce Structure-Enhanced Protein Instruction Tuning (SEPIT) framework to bridge this gap. Our approach integrates a noval structure-aware module into pLMs to inform them with structural knowledge, and then connects these enhanced pLMs to large language models (LLMs) to generate understanding of proteins. In this framework, we propose a novel two-stage instruction tuning pipeline that first establishes a basic understanding of proteins through caption-based instructions and then refines this understanding using a mixture of experts (MoEs) to learn more complex properties and functional information with the same amount of activated parameters. Moreover, we construct the largest and most comprehensive protein instruction dataset to date, which allows us to train and evaluate the general-purpose protein understanding model. Extensive experimental results on open-ended generation and closed-set answer tasks demonstrate the superior performance of SEPIT over both closed-source general LLMs and open-source LLMs trained with protein knowledge.
Multi-Modal CLIP-Informed Protein Editing
Jul 27, 2024Abstract:Proteins govern most biological functions essential for life, but achieving controllable protein discovery and optimization remains challenging. Recently, machine learning-assisted protein editing (MLPE) has shown promise in accelerating optimization cycles and reducing experimental workloads. However, current methods struggle with the vast combinatorial space of potential protein edits and cannot explicitly conduct protein editing using biotext instructions, limiting their interactivity with human feedback. To fill these gaps, we propose a novel method called ProtET for efficient CLIP-informed protein editing through multi-modality learning. Our approach comprises two stages: in the pretraining stage, contrastive learning aligns protein-biotext representations encoded by two large language models (LLMs), respectively. Subsequently, during the protein editing stage, the fused features from editing instruction texts and original protein sequences serve as the final editing condition for generating target protein sequences. Comprehensive experiments demonstrated the superiority of ProtET in editing proteins to enhance human-expected functionality across multiple attribute domains, including enzyme catalytic activity, protein stability and antibody specific binding ability. And ProtET improves the state-of-the-art results by a large margin, leading to significant stability improvements of 16.67% and 16.90%. This capability positions ProtET to advance real-world artificial protein editing, potentially addressing unmet academic, industrial, and clinical needs.
Group-On: Boosting One-Shot Segmentation with Supportive Query
Apr 18, 2024Abstract:One-shot semantic segmentation aims to segment query images given only ONE annotated support image of the same class. This task is challenging because target objects in the support and query images can be largely different in appearance and pose (i.e., intra-class variation). Prior works suggested that incorporating more annotated support images in few-shot settings boosts performances but increases costs due to additional manual labeling. In this paper, we propose a novel approach for ONE-shot semantic segmentation, called Group-On, which packs multiple query images in batches for the benefit of mutual knowledge support within the same category. Specifically, after coarse segmentation masks of the batch of queries are predicted, query-mask pairs act as pseudo support data to enhance mask predictions mutually, under the guidance of a simple Group-On Voting module. Comprehensive experiments on three standard benchmarks show that, in the ONE-shot setting, our Group-On approach significantly outperforms previous works by considerable margins. For example, on the COCO-20i dataset, we increase mIoU scores by 8.21% and 7.46% on ASNet and HSNet baselines, respectively. With only one support image, Group-On can be even competitive with the counterparts using 5 annotated support images.
Generative AI for Controllable Protein Sequence Design: A Survey
Feb 16, 2024

Abstract:The design of novel protein sequences with targeted functionalities underpins a central theme in protein engineering, impacting diverse fields such as drug discovery and enzymatic engineering. However, navigating this vast combinatorial search space remains a severe challenge due to time and financial constraints. This scenario is rapidly evolving as the transformative advancements in AI, particularly in the realm of generative models and optimization algorithms, have been propelling the protein design field towards an unprecedented revolution. In this survey, we systematically review recent advances in generative AI for controllable protein sequence design. To set the stage, we first outline the foundational tasks in protein sequence design in terms of the constraints involved and present key generative models and optimization algorithms. We then offer in-depth reviews of each design task and discuss the pertinent applications. Finally, we identify the unresolved challenges and highlight research opportunities that merit deeper exploration.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge