Michael B. Wallace
Cyst-X: AI-Powered Pancreatic Cancer Risk Prediction from Multicenter MRI in Centralized and Federated Learning
Jul 29, 2025

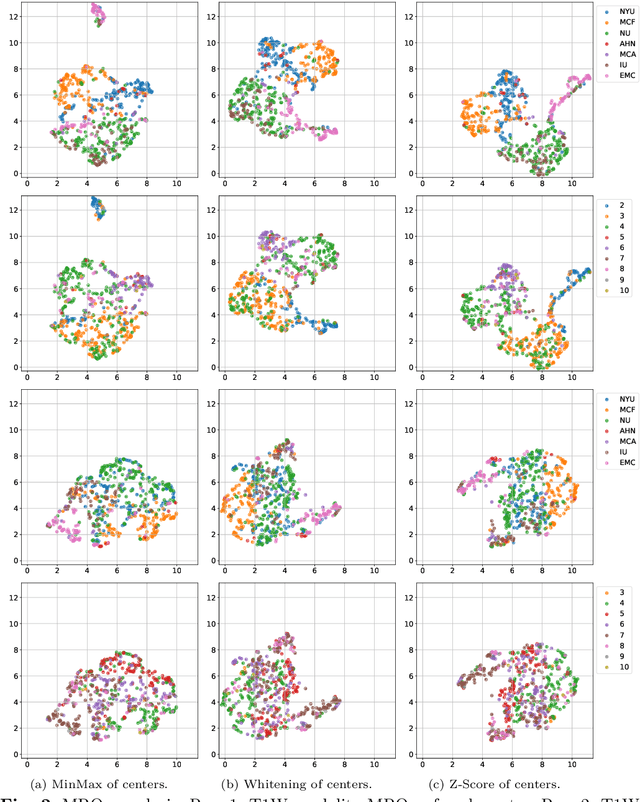
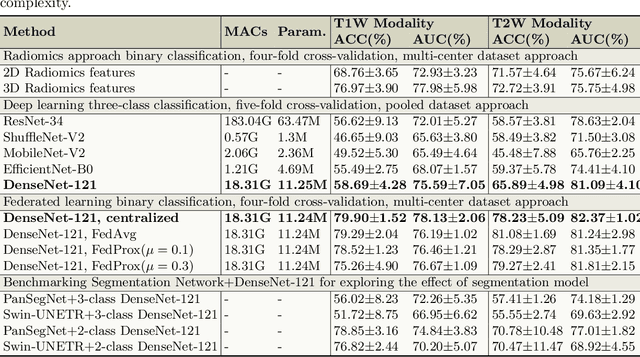
Abstract:Pancreatic cancer is projected to become the second-deadliest malignancy in Western countries by 2030, highlighting the urgent need for better early detection. Intraductal papillary mucinous neoplasms (IPMNs), key precursors to pancreatic cancer, are challenging to assess with current guidelines, often leading to unnecessary surgeries or missed malignancies. We present Cyst-X, an AI framework that predicts IPMN malignancy using multicenter MRI data, leveraging MRI's superior soft tissue contrast over CT. Trained on 723 T1- and 738 T2-weighted scans from 764 patients across seven institutions, our models (AUC=0.82) significantly outperform both Kyoto guidelines (AUC=0.75) and expert radiologists. The AI-derived imaging features align with known clinical markers and offer biologically meaningful insights. We also demonstrate strong performance in a federated learning setting, enabling collaborative training without sharing patient data. To promote privacy-preserving AI development and improve IPMN risk stratification, the Cyst-X dataset is released as the first large-scale, multi-center pancreatic cysts MRI dataset.
IPMN Risk Assessment under Federated Learning Paradigm
Nov 08, 2024



Abstract:Accurate classification of Intraductal Papillary Mucinous Neoplasms (IPMN) is essential for identifying high-risk cases that require timely intervention. In this study, we develop a federated learning framework for multi-center IPMN classification utilizing a comprehensive pancreas MRI dataset. This dataset includes 653 T1-weighted and 656 T2-weighted MRI images, accompanied by corresponding IPMN risk scores from 7 leading medical institutions, making it the largest and most diverse dataset for IPMN classification to date. We assess the performance of DenseNet-121 in both centralized and federated settings for training on distributed data. Our results demonstrate that the federated learning approach achieves high classification accuracy comparable to centralized learning while ensuring data privacy across institutions. This work marks a significant advancement in collaborative IPMN classification, facilitating secure and high-accuracy model training across multiple centers.
Adaptive Aggregation Weights for Federated Segmentation of Pancreas MRI
Oct 29, 2024

Abstract:Federated learning (FL) enables collaborative model training across institutions without sharing sensitive data, making it an attractive solution for medical imaging tasks. However, traditional FL methods, such as Federated Averaging (FedAvg), face difficulties in generalizing across domains due to variations in imaging protocols and patient demographics across institutions. This challenge is particularly evident in pancreas MRI segmentation, where anatomical variability and imaging artifacts significantly impact performance. In this paper, we conduct a comprehensive evaluation of FL algorithms for pancreas MRI segmentation and introduce a novel approach that incorporates adaptive aggregation weights. By dynamically adjusting the contribution of each client during model aggregation, our method accounts for domain-specific differences and improves generalization across heterogeneous datasets. Experimental results demonstrate that our approach enhances segmentation accuracy and reduces the impact of domain shift compared to conventional FL methods while maintaining privacy-preserving capabilities. Significant performance improvements are observed across multiple hospitals (centers).
Optimizing Synthetic Data for Enhanced Pancreatic Tumor Segmentation
Jul 27, 2024



Abstract:Pancreatic cancer remains one of the leading causes of cancer-related mortality worldwide. Precise segmentation of pancreatic tumors from medical images is a bottleneck for effective clinical decision-making. However, achieving a high accuracy is often limited by the small size and availability of real patient data for training deep learning models. Recent approaches have employed synthetic data generation to augment training datasets. While promising, these methods may not yet meet the performance benchmarks required for real-world clinical use. This study critically evaluates the limitations of existing generative-AI based frameworks for pancreatic tumor segmentation. We conduct a series of experiments to investigate the impact of synthetic \textit{tumor size} and \textit{boundary definition} precision on model performance. Our findings demonstrate that: (1) strategically selecting a combination of synthetic tumor sizes is crucial for optimal segmentation outcomes, and (2) generating synthetic tumors with precise boundaries significantly improves model accuracy. These insights highlight the importance of utilizing refined synthetic data augmentation for enhancing the clinical utility of segmentation models in pancreatic cancer decision making including diagnosis, prognosis, and treatment plans. Our code will be available at https://github.com/lkpengcs/SynTumorAnalyzer.
Large-Scale Multi-Center CT and MRI Segmentation of Pancreas with Deep Learning
May 20, 2024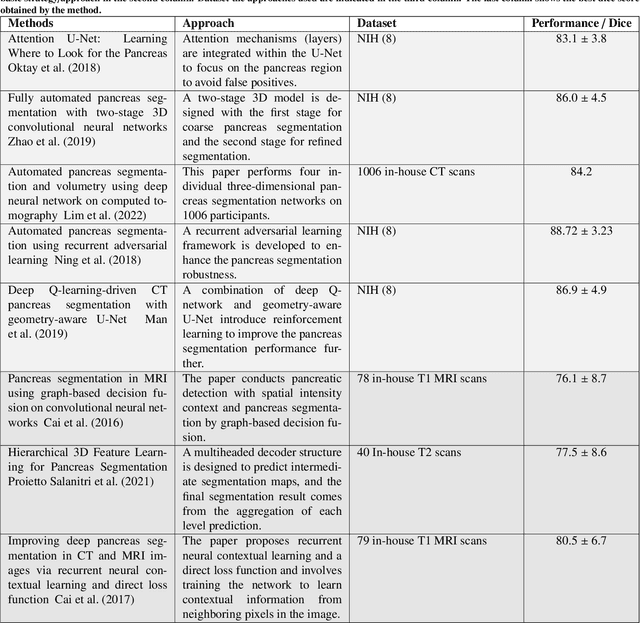
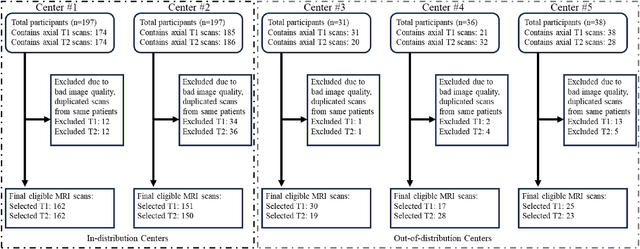
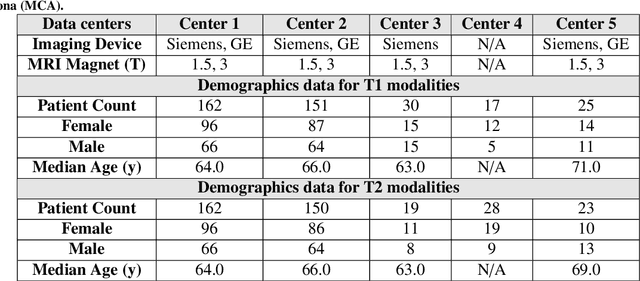
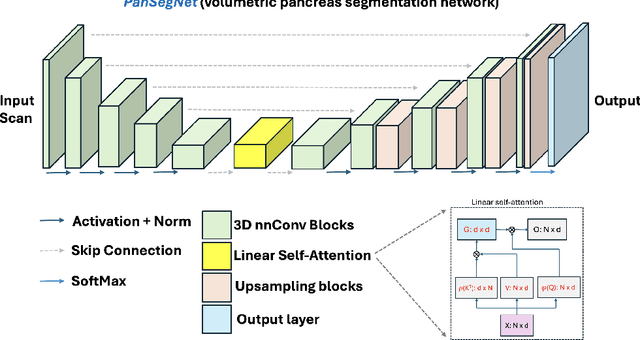
Abstract:Automated volumetric segmentation of the pancreas on cross-sectional imaging is needed for diagnosis and follow-up of pancreatic diseases. While CT-based pancreatic segmentation is more established, MRI-based segmentation methods are understudied, largely due to a lack of publicly available datasets, benchmarking research efforts, and domain-specific deep learning methods. In this retrospective study, we collected a large dataset (767 scans from 499 participants) of T1-weighted (T1W) and T2-weighted (T2W) abdominal MRI series from five centers between March 2004 and November 2022. We also collected CT scans of 1,350 patients from publicly available sources for benchmarking purposes. We developed a new pancreas segmentation method, called PanSegNet, combining the strengths of nnUNet and a Transformer network with a new linear attention module enabling volumetric computation. We tested PanSegNet's accuracy in cross-modality (a total of 2,117 scans) and cross-center settings with Dice and Hausdorff distance (HD95) evaluation metrics. We used Cohen's kappa statistics for intra and inter-rater agreement evaluation and paired t-tests for volume and Dice comparisons, respectively. For segmentation accuracy, we achieved Dice coefficients of 88.3% (std: 7.2%, at case level) with CT, 85.0% (std: 7.9%) with T1W MRI, and 86.3% (std: 6.4%) with T2W MRI. There was a high correlation for pancreas volume prediction with R^2 of 0.91, 0.84, and 0.85 for CT, T1W, and T2W, respectively. We found moderate inter-observer (0.624 and 0.638 for T1W and T2W MRI, respectively) and high intra-observer agreement scores. All MRI data is made available at https://osf.io/kysnj/. Our source code is available at https://github.com/NUBagciLab/PaNSegNet.
Neural Transformers for Intraductal Papillary Mucosal Neoplasms (IPMN) Classification in MRI images
Jun 21, 2022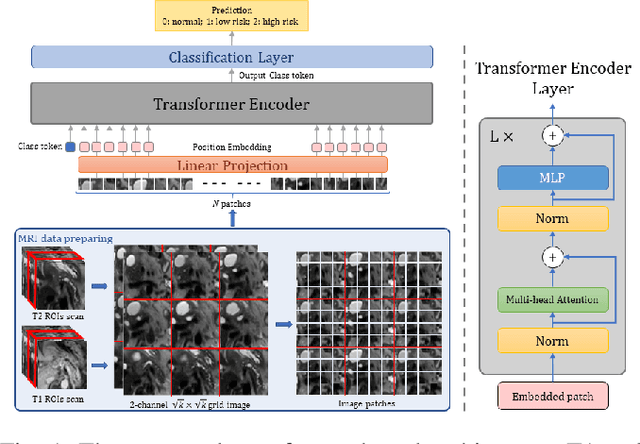
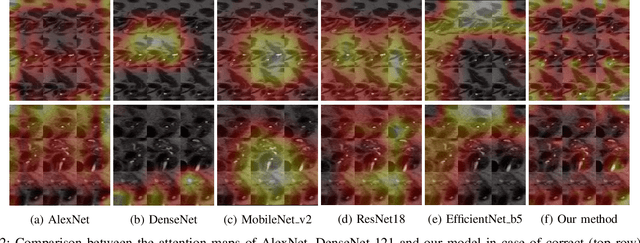

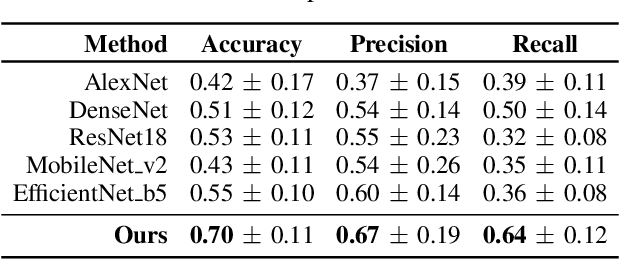
Abstract:Early detection of precancerous cysts or neoplasms, i.e., Intraductal Papillary Mucosal Neoplasms (IPMN), in pancreas is a challenging and complex task, and it may lead to a more favourable outcome. Once detected, grading IPMNs accurately is also necessary, since low-risk IPMNs can be under surveillance program, while high-risk IPMNs have to be surgically resected before they turn into cancer. Current standards (Fukuoka and others) for IPMN classification show significant intra- and inter-operator variability, beside being error-prone, making a proper diagnosis unreliable. The established progress in artificial intelligence, through the deep learning paradigm, may provide a key tool for an effective support to medical decision for pancreatic cancer. In this work, we follow this trend, by proposing a novel AI-based IPMN classifier that leverages the recent success of transformer networks in generalizing across a wide variety of tasks, including vision ones. We specifically show that our transformer-based model exploits pre-training better than standard convolutional neural networks, thus supporting the sought architectural universalism of transformers in vision, including the medical image domain and it allows for a better interpretation of the obtained results.
Diagnosing Colorectal Polyps in the Wild with Capsule Networks
Jan 10, 2020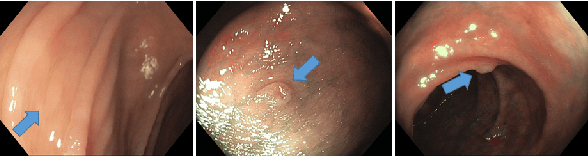
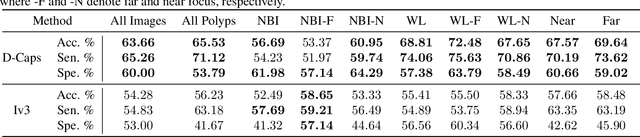
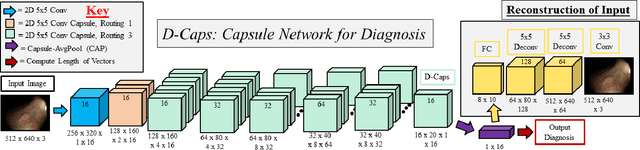
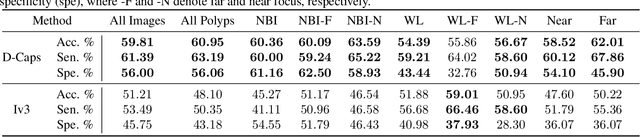
Abstract:Colorectal cancer, largely arising from precursor lesions called polyps, remains one of the leading causes of cancer-related death worldwide. Current clinical standards require the resection and histopathological analysis of polyps due to test accuracy and sensitivity of optical biopsy methods falling substantially below recommended levels. In this study, we design a novel capsule network architecture (D-Caps) to improve the viability of optical biopsy of colorectal polyps. Our proposed method introduces several technical novelties including a novel capsule architecture with a capsule-average pooling (CAP) method to improve efficiency in large-scale image classification. We demonstrate improved results over the previous state-of-the-art convolutional neural network (CNN) approach by as much as 43%. This work provides an important benchmark on the new Mayo Polyp dataset, a significantly more challenging and larger dataset than previous polyp studies, with results stratified across all available categories, imaging devices and modalities, and focus modes to promote future direction into AI-driven colorectal cancer screening systems. Code is publicly available at https://github.com/lalonderodney/D-Caps .
INN: Inflated Neural Networks for IPMN Diagnosis
Jun 30, 2019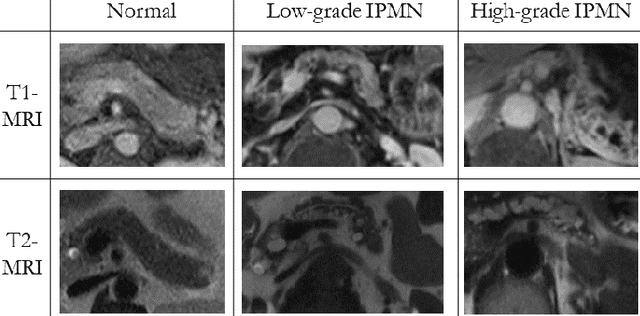
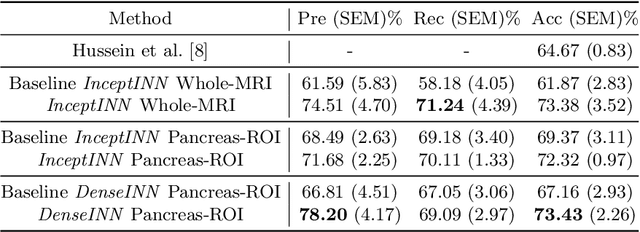
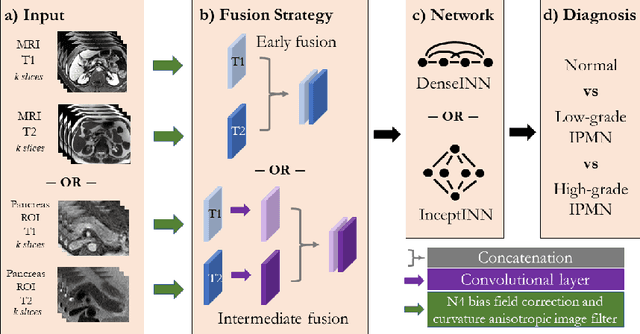

Abstract:Intraductal papillary mucinous neoplasm (IPMN) is a precursor to pancreatic ductal adenocarcinoma. While over half of patients are diagnosed with pancreatic cancer at a distant stage, patients who are diagnosed early enjoy a much higher 5-year survival rate of $34\%$ compared to $3\%$ in the former; hence, early diagnosis is key. Unique challenges in the medical imaging domain such as extremely limited annotated data sets and typically large 3D volumetric data have made it difficult for deep learning to secure a strong foothold. In this work, we construct two novel "inflated" deep network architectures, $\textit{InceptINN}$ and $\textit{DenseINN}$, for the task of diagnosing IPMN from multisequence (T1 and T2) MRI. These networks inflate their 2D layers to 3D and bootstrap weights from their 2D counterparts (Inceptionv3 and DenseNet121 respectively) trained on ImageNet to the new 3D kernels. We also extend the inflation process by further expanding the pre-trained kernels to handle any number of input modalities and different fusion strategies. This is one of the first studies to train an end-to-end deep network on multisequence MRI for IPMN diagnosis, and shows that our proposed novel inflated network architectures are able to handle the extremely limited training data (139 MRI scans), while providing an absolute improvement of $8.76\%$ in accuracy for diagnosing IPMN over the current state-of-the-art. Code is publicly available at https://github.com/lalonderodney/INN-Inflated-Neural-Nets.
Supervised and Unsupervised Tumor Characterization in the Deep Learning Era
Jul 29, 2018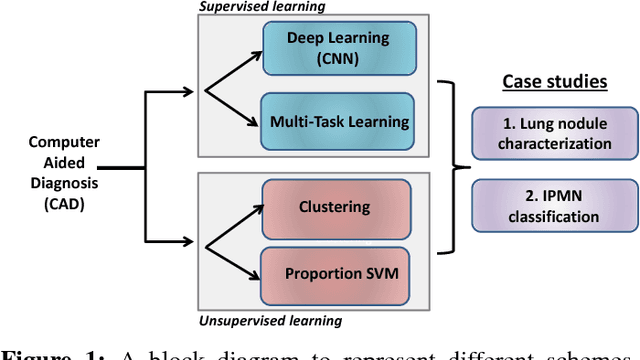

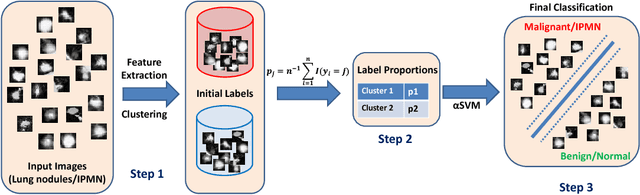
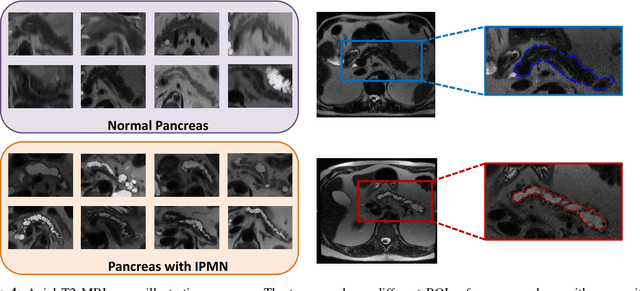
Abstract:Cancer is among the leading causes of death worldwide. Risk stratification of cancer tumors in radiology images can be improved with computer-aided diagnosis (CAD) tools which can be made faster and more accurate. Tumor characterization through CADs can enable non-invasive cancer staging and prognosis, and foster personalized treatment planning as a part of precision medicine. In this study, we propose both supervised and unsupervised machine learning strategies to improve tumor characterization. Our first approach is based on supervised learning for which we demonstrate significant gains in deep learning algorithms, particularly by utilizing a 3D Convolutional Neural Network along with transfer learning. Motivated by the radiologists' interpretations of the scans, we then show how to incorporate task dependent feature representations into a CAD system via a "graph-regularized sparse Multi-Task Learning (MTL)" framework. In the second approach, we explore an unsupervised scheme to address the limited availability of labeled training data, a common problem in medical imaging applications. Inspired by learning from label proportion (LLP) approaches, we propose a new algorithm, proportion-SVM, to characterize tumor types. We also seek the answer to the fundamental question about the goodness of "deep features" for unsupervised tumor classification. We evaluate our proposed approaches (both supervised and unsupervised) on two different tumor diagnosis challenges: lung and pancreas with 1018 CT and 171 MRI scans respectively.
Deep Multi-Modal Classification of Intraductal Papillary Mucinous Neoplasms (IPMN) with Canonical Correlation Analysis
Apr 27, 2018
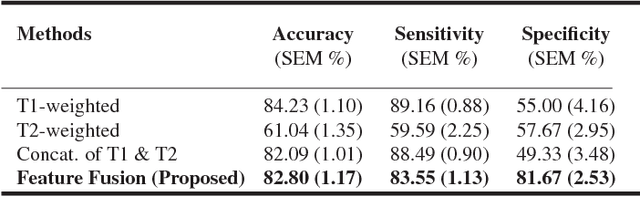
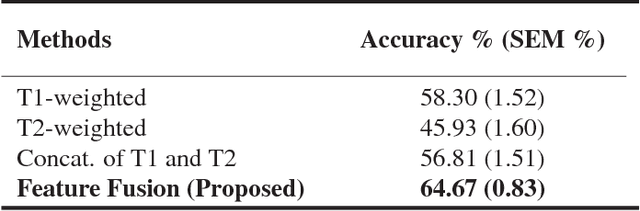
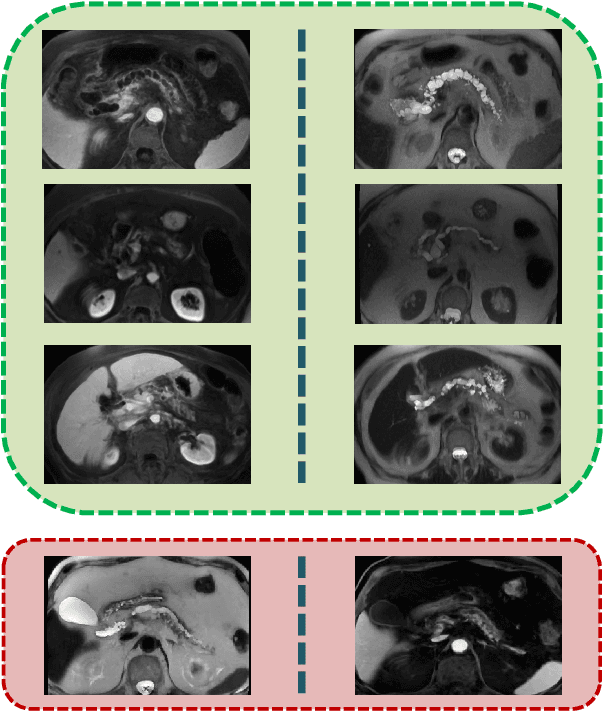
Abstract:Pancreatic cancer has the poorest prognosis among all cancer types. Intraductal Papillary Mucinous Neoplasms (IPMNs) are radiographically identifiable precursors to pancreatic cancer; hence, early detection and precise risk assessment of IPMN are vital. In this work, we propose a Convolutional Neural Network (CNN) based computer aided diagnosis (CAD) system to perform IPMN diagnosis and risk assessment by utilizing multi-modal MRI. In our proposed approach, we use minimum and maximum intensity projections to ease the annotation variations among different slices and type of MRIs. Then, we present a CNN to obtain deep feature representation corresponding to each MRI modality (T1-weighted and T2-weighted). At the final step, we employ canonical correlation analysis (CCA) to perform a fusion operation at the feature level, leading to discriminative canonical correlation features. Extracted features are used for classification. Our results indicate significant improvements over other potential approaches to solve this important problem. The proposed approach doesn't require explicit sample balancing in cases of imbalance between positive and negative examples. To the best of our knowledge, our study is the first to automatically diagnose IPMN using multi-modal MRI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge