Large-Scale Multi-Center CT and MRI Segmentation of Pancreas with Deep Learning
Paper and Code
May 20, 2024
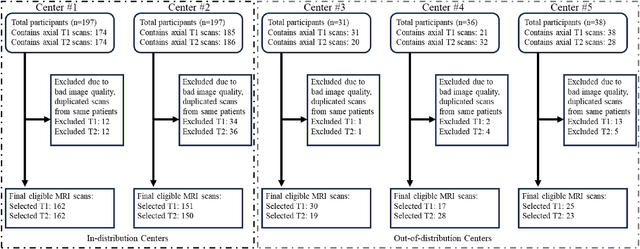
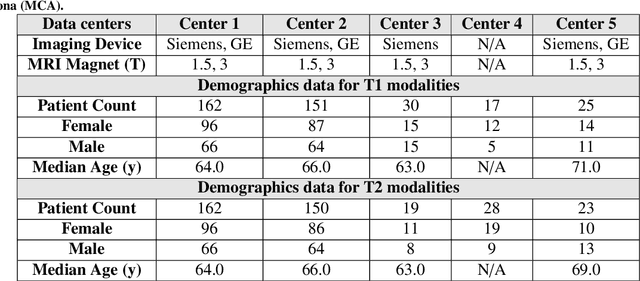
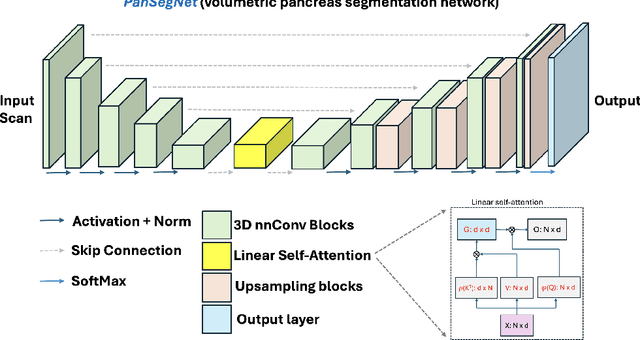
Automated volumetric segmentation of the pancreas on cross-sectional imaging is needed for diagnosis and follow-up of pancreatic diseases. While CT-based pancreatic segmentation is more established, MRI-based segmentation methods are understudied, largely due to a lack of publicly available datasets, benchmarking research efforts, and domain-specific deep learning methods. In this retrospective study, we collected a large dataset (767 scans from 499 participants) of T1-weighted (T1W) and T2-weighted (T2W) abdominal MRI series from five centers between March 2004 and November 2022. We also collected CT scans of 1,350 patients from publicly available sources for benchmarking purposes. We developed a new pancreas segmentation method, called PanSegNet, combining the strengths of nnUNet and a Transformer network with a new linear attention module enabling volumetric computation. We tested PanSegNet's accuracy in cross-modality (a total of 2,117 scans) and cross-center settings with Dice and Hausdorff distance (HD95) evaluation metrics. We used Cohen's kappa statistics for intra and inter-rater agreement evaluation and paired t-tests for volume and Dice comparisons, respectively. For segmentation accuracy, we achieved Dice coefficients of 88.3% (std: 7.2%, at case level) with CT, 85.0% (std: 7.9%) with T1W MRI, and 86.3% (std: 6.4%) with T2W MRI. There was a high correlation for pancreas volume prediction with R^2 of 0.91, 0.84, and 0.85 for CT, T1W, and T2W, respectively. We found moderate inter-observer (0.624 and 0.638 for T1W and T2W MRI, respectively) and high intra-observer agreement scores. All MRI data is made available at https://osf.io/kysnj/. Our source code is available at https://github.com/NUBagciLab/PaNSegNet.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge