Lanhong Yao
Large-Scale Multi-Center CT and MRI Segmentation of Pancreas with Deep Learning
May 20, 2024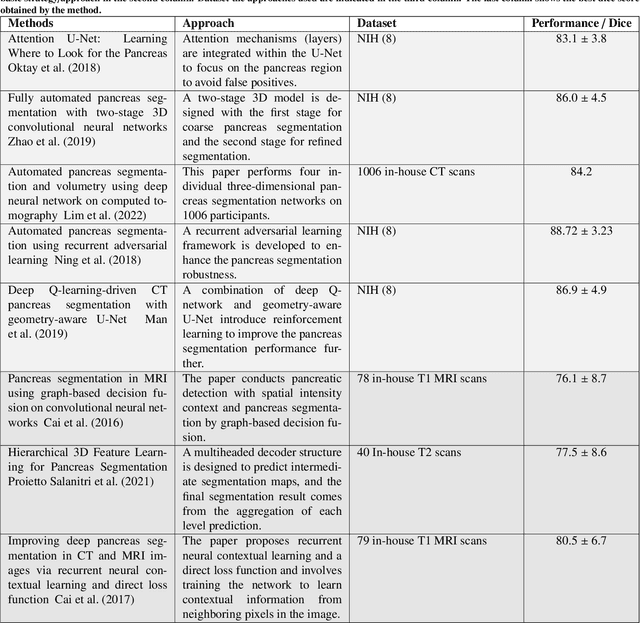
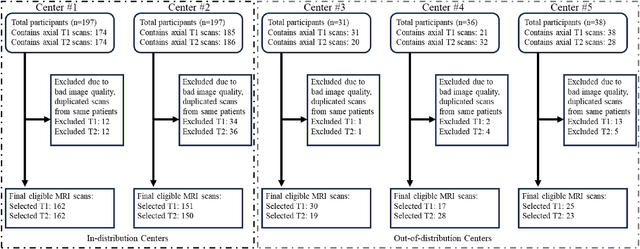
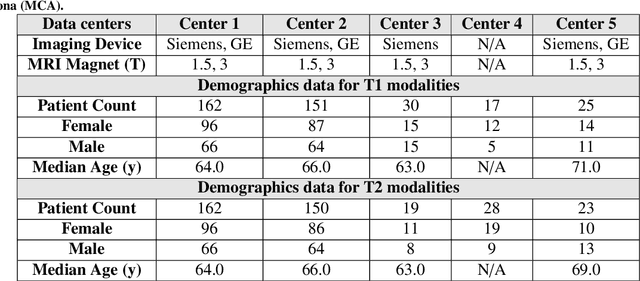
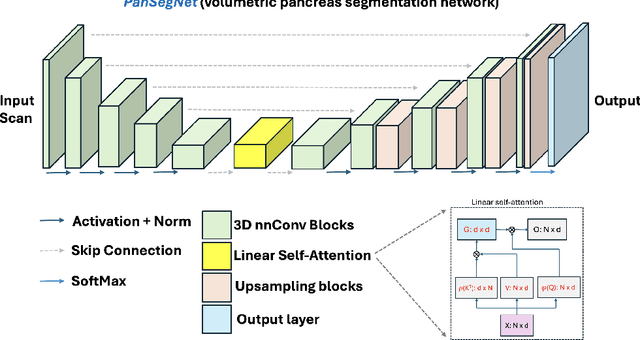
Abstract:Automated volumetric segmentation of the pancreas on cross-sectional imaging is needed for diagnosis and follow-up of pancreatic diseases. While CT-based pancreatic segmentation is more established, MRI-based segmentation methods are understudied, largely due to a lack of publicly available datasets, benchmarking research efforts, and domain-specific deep learning methods. In this retrospective study, we collected a large dataset (767 scans from 499 participants) of T1-weighted (T1W) and T2-weighted (T2W) abdominal MRI series from five centers between March 2004 and November 2022. We also collected CT scans of 1,350 patients from publicly available sources for benchmarking purposes. We developed a new pancreas segmentation method, called PanSegNet, combining the strengths of nnUNet and a Transformer network with a new linear attention module enabling volumetric computation. We tested PanSegNet's accuracy in cross-modality (a total of 2,117 scans) and cross-center settings with Dice and Hausdorff distance (HD95) evaluation metrics. We used Cohen's kappa statistics for intra and inter-rater agreement evaluation and paired t-tests for volume and Dice comparisons, respectively. For segmentation accuracy, we achieved Dice coefficients of 88.3% (std: 7.2%, at case level) with CT, 85.0% (std: 7.9%) with T1W MRI, and 86.3% (std: 6.4%) with T2W MRI. There was a high correlation for pancreas volume prediction with R^2 of 0.91, 0.84, and 0.85 for CT, T1W, and T2W, respectively. We found moderate inter-observer (0.624 and 0.638 for T1W and T2W MRI, respectively) and high intra-observer agreement scores. All MRI data is made available at https://osf.io/kysnj/. Our source code is available at https://github.com/NUBagciLab/PaNSegNet.
EMIT-Diff: Enhancing Medical Image Segmentation via Text-Guided Diffusion Model
Oct 19, 2023Abstract:Large-scale, big-variant, and high-quality data are crucial for developing robust and successful deep-learning models for medical applications since they potentially enable better generalization performance and avoid overfitting. However, the scarcity of high-quality labeled data always presents significant challenges. This paper proposes a novel approach to address this challenge by developing controllable diffusion models for medical image synthesis, called EMIT-Diff. We leverage recent diffusion probabilistic models to generate realistic and diverse synthetic medical image data that preserve the essential characteristics of the original medical images by incorporating edge information of objects to guide the synthesis process. In our approach, we ensure that the synthesized samples adhere to medically relevant constraints and preserve the underlying structure of imaging data. Due to the random sampling process by the diffusion model, we can generate an arbitrary number of synthetic images with diverse appearances. To validate the effectiveness of our proposed method, we conduct an extensive set of medical image segmentation experiments on multiple datasets, including Ultrasound breast (+13.87%), CT spleen (+0.38%), and MRI prostate (+7.78%), achieving significant improvements over the baseline segmentation methods. For the first time, to our best knowledge, the promising results demonstrate the effectiveness of our EMIT-Diff for medical image segmentation tasks and show the feasibility of introducing a first-ever text-guided diffusion model for general medical image segmentation tasks. With carefully designed ablation experiments, we investigate the influence of various data augmentation ratios, hyper-parameter settings, patch size for generating random merging mask settings, and combined influence with different network architectures.
Radiomics Boosts Deep Learning Model for IPMN Classification
Sep 11, 2023Abstract:Intraductal Papillary Mucinous Neoplasm (IPMN) cysts are pre-malignant pancreas lesions, and they can progress into pancreatic cancer. Therefore, detecting and stratifying their risk level is of ultimate importance for effective treatment planning and disease control. However, this is a highly challenging task because of the diverse and irregular shape, texture, and size of the IPMN cysts as well as the pancreas. In this study, we propose a novel computer-aided diagnosis pipeline for IPMN risk classification from multi-contrast MRI scans. Our proposed analysis framework includes an efficient volumetric self-adapting segmentation strategy for pancreas delineation, followed by a newly designed deep learning-based classification scheme with a radiomics-based predictive approach. We test our proposed decision-fusion model in multi-center data sets of 246 multi-contrast MRI scans and obtain superior performance to the state of the art (SOTA) in this field. Our ablation studies demonstrate the significance of both radiomics and deep learning modules for achieving the new SOTA performance compared to international guidelines and published studies (81.9\% vs 61.3\% in accuracy). Our findings have important implications for clinical decision-making. In a series of rigorous experiments on multi-center data sets (246 MRI scans from five centers), we achieved unprecedented performance (81.9\% accuracy).
Ensemble Learning with Residual Transformer for Brain Tumor Segmentation
Jul 31, 2023Abstract:Brain tumor segmentation is an active research area due to the difficulty in delineating highly complex shaped and textured tumors as well as the failure of the commonly used U-Net architectures. The combination of different neural architectures is among the mainstream research recently, particularly the combination of U-Net with Transformers because of their innate attention mechanism and pixel-wise labeling. Different from previous efforts, this paper proposes a novel network architecture that integrates Transformers into a self-adaptive U-Net to draw out 3D volumetric contexts with reasonable computational costs. We further add a residual connection to prevent degradation in information flow and explore ensemble methods, as the evaluated models have edges on different cases and sub-regions. On the BraTS 2021 dataset (3D), our model achieves 87.6% mean Dice score and outperforms the state-of-the-art methods, demonstrating the potential for combining multiple architectures to optimize brain tumor segmentation.
Domain Generalization with Adversarial Intensity Attack for Medical Image Segmentation
Apr 05, 2023Abstract:Most statistical learning algorithms rely on an over-simplified assumption, that is, the train and test data are independent and identically distributed. In real-world scenarios, however, it is common for models to encounter data from new and different domains to which they were not exposed to during training. This is often the case in medical imaging applications due to differences in acquisition devices, imaging protocols, and patient characteristics. To address this problem, domain generalization (DG) is a promising direction as it enables models to handle data from previously unseen domains by learning domain-invariant features robust to variations across different domains. To this end, we introduce a novel DG method called Adversarial Intensity Attack (AdverIN), which leverages adversarial training to generate training data with an infinite number of styles and increase data diversity while preserving essential content information. We conduct extensive evaluation experiments on various multi-domain segmentation datasets, including 2D retinal fundus optic disc/cup and 3D prostate MRI. Our results demonstrate that AdverIN significantly improves the generalization ability of the segmentation models, achieving significant improvement on these challenging datasets. Code is available upon publication.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge