Jannis Born
Quantum Doubly Stochastic Transformers
Apr 22, 2025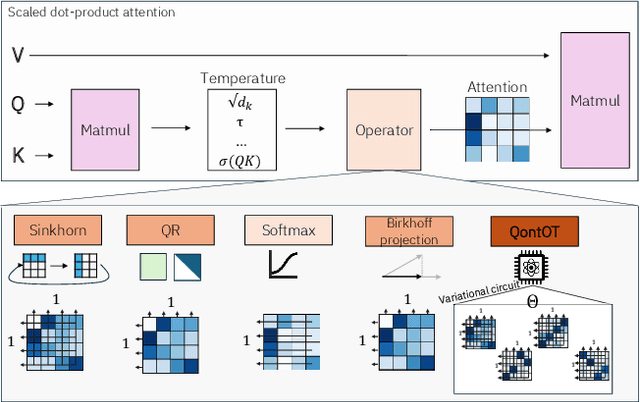
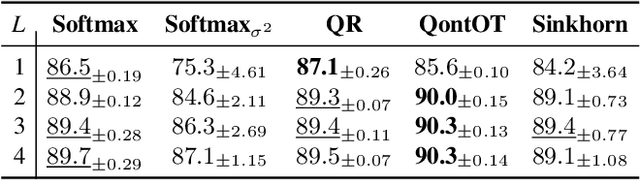

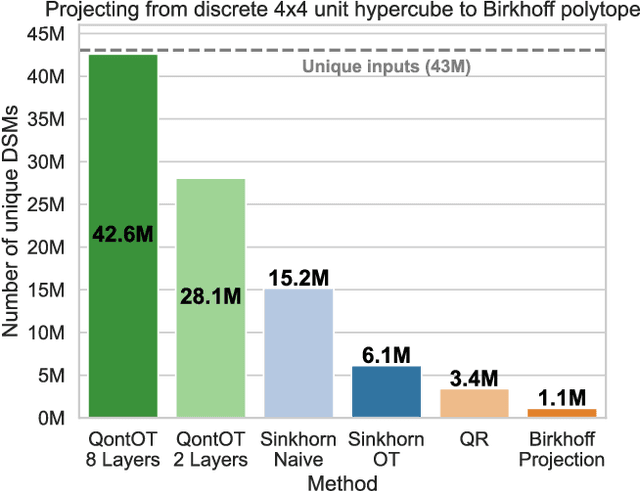

Abstract:At the core of the Transformer, the Softmax normalizes the attention matrix to be right stochastic. Previous research has shown that this often destabilizes training and that enforcing the attention matrix to be doubly stochastic (through Sinkhorn's algorithm) consistently improves performance across different tasks, domains and Transformer flavors. However, Sinkhorn's algorithm is iterative, approximative, non-parametric and thus inflexible w.r.t. the obtained doubly stochastic matrix (DSM). Recently, it has been proven that DSMs can be obtained with a parametric quantum circuit, yielding a novel quantum inductive bias for DSMs with no known classical analogue. Motivated by this, we demonstrate the feasibility of a hybrid classical-quantum doubly stochastic Transformer (QDSFormer) that replaces the Softmax in the self-attention layer with a variational quantum circuit. We study the expressive power of the circuit and find that it yields more diverse DSMs that better preserve information than classical operators. Across multiple small-scale object recognition tasks, we find that our QDSFormer consistently surpasses both a standard Vision Transformer and other doubly stochastic Transformers. Beyond the established Sinkformer, this comparison includes a novel quantum-inspired doubly stochastic Transformer (based on QR decomposition) that can be of independent interest. The QDSFormer also shows improved training stability and lower performance variation suggesting that it may mitigate the notoriously unstable training of ViTs on small-scale data.
Towards generalizable single-cell perturbation modeling via the Conditional Monge Gap
Apr 11, 2025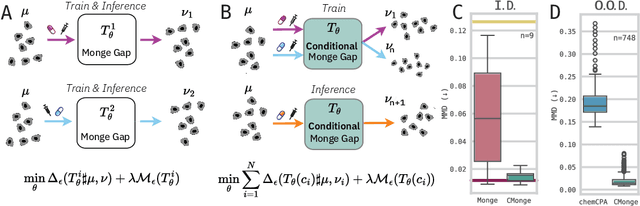
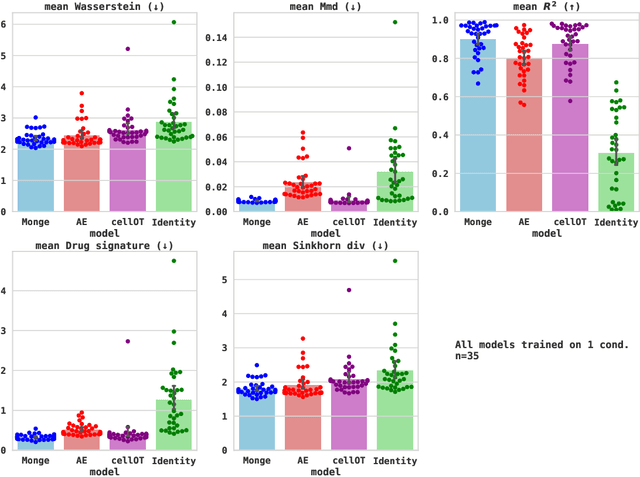

Abstract:Learning the response of single-cells to various treatments offers great potential to enable targeted therapies. In this context, neural optimal transport (OT) has emerged as a principled methodological framework because it inherently accommodates the challenges of unpaired data induced by cell destruction during data acquisition. However, most existing OT approaches are incapable of conditioning on different treatment contexts (e.g., time, drug treatment, drug dosage, or cell type) and we still lack methods that unanimously show promising generalization performance to unseen treatments. Here, we propose the Conditional Monge Gap which learns OT maps conditionally on arbitrary covariates. We demonstrate its value in predicting single-cell perturbation responses conditional to one or multiple drugs, a drug dosage, or combinations thereof. We find that our conditional models achieve results comparable and sometimes even superior to the condition-specific state-of-the-art on scRNA-seq as well as multiplexed protein imaging data. Notably, by aggregating data across conditions we perform cross-task learning which unlocks remarkable generalization abilities to unseen drugs or drug dosages, widely outperforming other conditional models in capturing heterogeneity (i.e., higher moments) in the perturbed population. Finally, by scaling to hundreds of conditions and testing on unseen drugs, we narrow the gap between structure-based and effect-based drug representations, suggesting a promising path to the successful prediction of perturbation effects for unseen treatments.
We Need Improved Data Curation and Attribution in AI for Scientific Discovery
Apr 03, 2025



Abstract:As the interplay between human-generated and synthetic data evolves, new challenges arise in scientific discovery concerning the integrity of the data and the stability of the models. In this work, we examine the role of synthetic data as opposed to that of real experimental data for scientific research. Our analyses indicate that nearly three-quarters of experimental datasets available on open-access platforms have relatively low adoption rates, opening new opportunities to enhance their discoverability and usability by automated methods. Additionally, we observe an increasing difficulty in distinguishing synthetic from real experimental data. We propose supplementing ongoing efforts in automating synthetic data detection by increasing the focus on watermarking real experimental data, thereby strengthening data traceability and integrity. Our estimates suggest that watermarking even less than half of the real world data generated annually could help sustain model robustness, while promoting a balanced integration of synthetic and human-generated content.
Multiscale Byte Language Models -- A Hierarchical Architecture for Causal Million-Length Sequence Modeling
Feb 20, 2025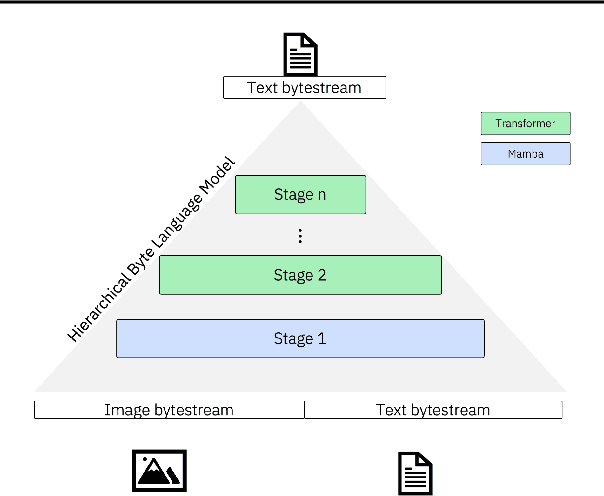
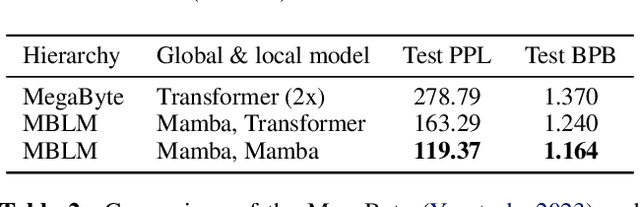
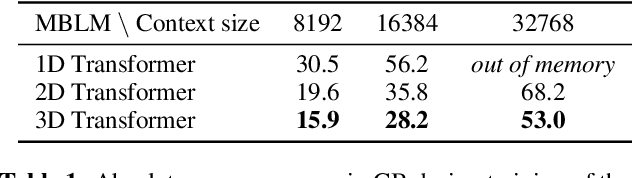
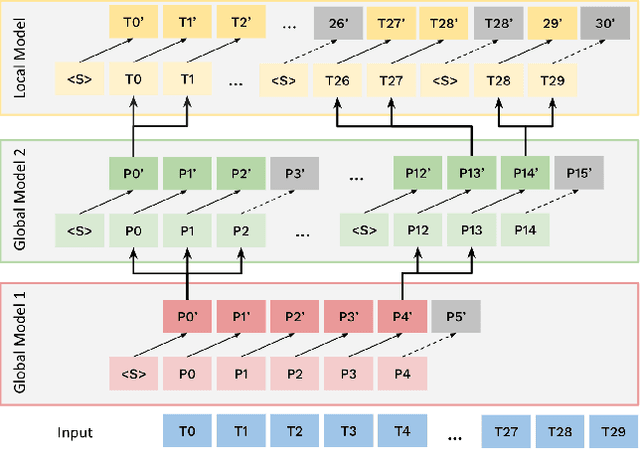

Abstract:Bytes form the basis of the digital world and thus are a promising building block for multimodal foundation models. Recently, Byte Language Models (BLMs) have emerged to overcome tokenization, yet the excessive length of bytestreams requires new architectural paradigms. Therefore, we present the Multiscale Byte Language Model (MBLM), a model-agnostic hierarchical decoder stack that allows training with context windows of $5$M bytes on single GPU in full model precision. We thoroughly examine MBLM's performance with Transformer and Mamba blocks on both unimodal and multimodal tasks. Our experiments demonstrate that hybrid architectures are efficient in handling extremely long byte sequences during training while achieving near-linear generational efficiency. To the best of our knowledge, we present the first evaluation of BLMs on visual Q\&A tasks and find that, despite serializing images and the absence of an encoder, a MBLM with pure next token prediction can match custom CNN-LSTM architectures with designated classification heads. We show that MBLMs exhibit strong adaptability in integrating diverse data representations, including pixel and image filestream bytes, underlining their potential toward omnimodal foundation models. Source code is publicly available at: https://github.com/ai4sd/multiscale-byte-lm
Regress, Don't Guess -- A Regression-like Loss on Number Tokens for Language Models
Nov 04, 2024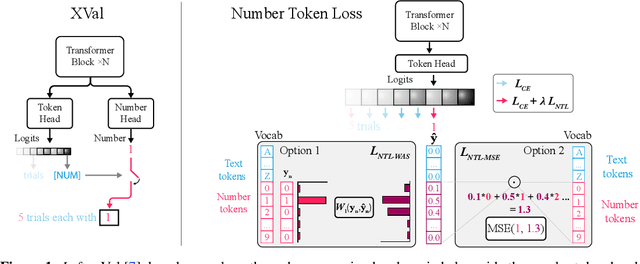
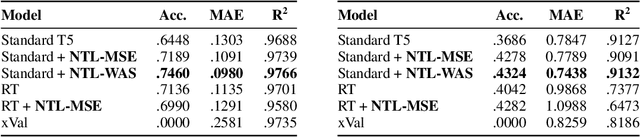
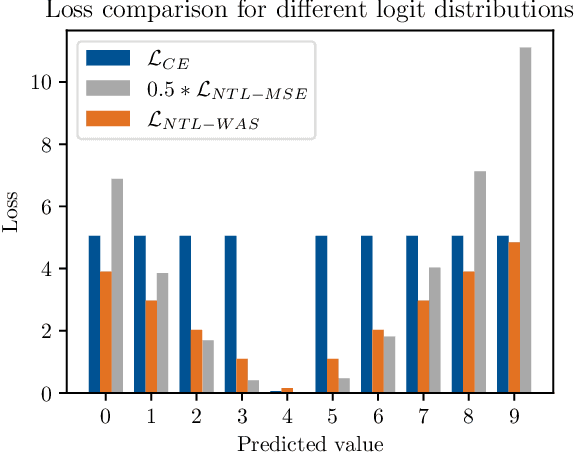
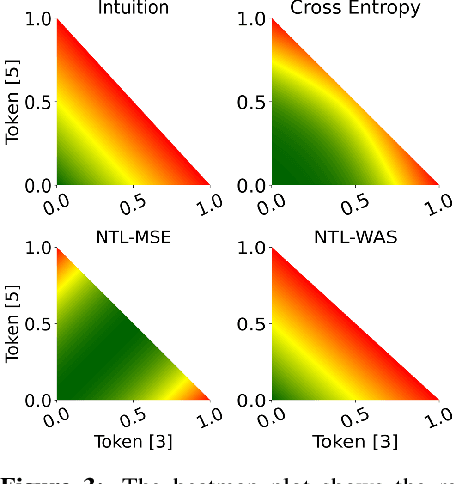
Abstract:While language models have exceptional capabilities at text generation, they lack a natural inductive bias for emitting numbers and thus struggle in tasks involving reasoning over quantities, especially arithmetics. This has particular relevance in scientific datasets where combinations of text and numerical data are abundant. One fundamental limitation is the nature of the CE loss, which assumes a nominal (categorical) scale and thus cannot convey proximity between generated number tokens. As a remedy, we here present two versions of a number token loss. The first is based on an $L_p$ loss between the ground truth token value and the weighted sum of the predicted class probabilities. The second loss minimizes the Wasserstein-1 distance between the distribution of the predicted output probabilities and the ground truth distribution. These regression-like losses can easily be added to any language model and extend the CE objective during training. We compare the proposed schemes on a mathematics dataset against existing tokenization, encoding, and decoding schemes for improving number representation in language models. Our results reveal a significant improvement in numerical accuracy when equipping a standard T5 model with the proposed loss schemes.
Quantum Theory and Application of Contextual Optimal Transport
Feb 22, 2024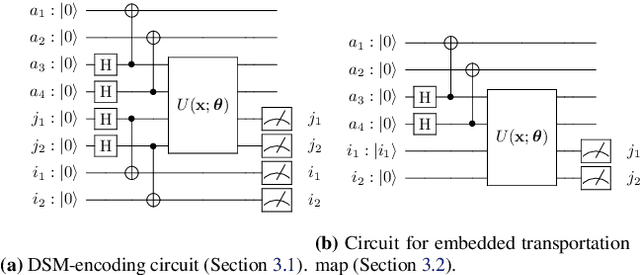
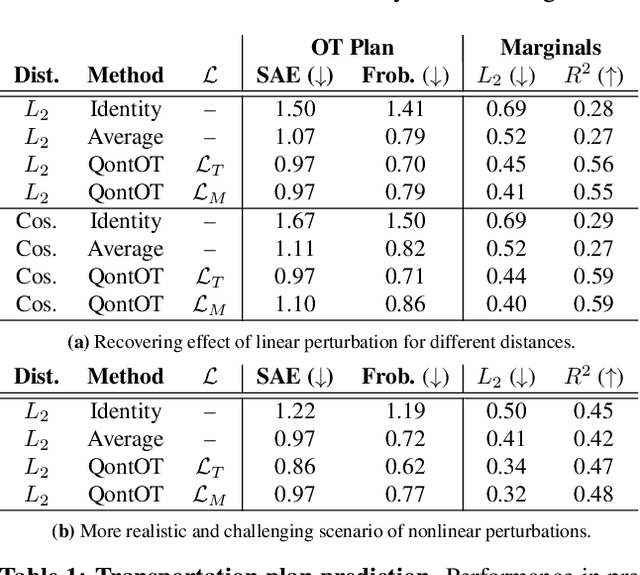
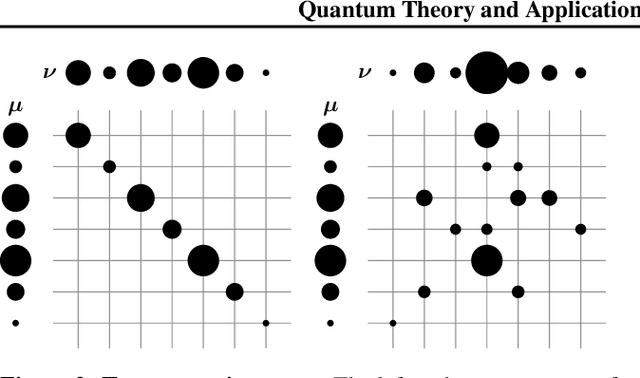
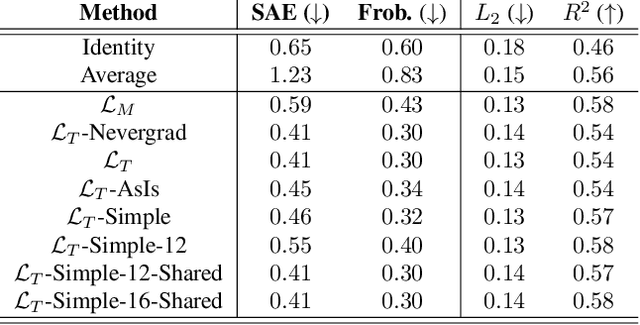
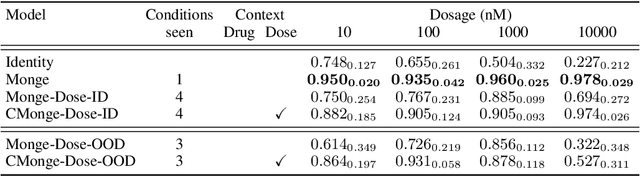
Abstract:Optimal Transport (OT) has fueled machine learning (ML) applications across many domains. In cases where paired data measurements ($\mu$, $\nu$) are coupled to a context variable $p_i$ , one may aspire to learn a global transportation map that can be parameterized through a potentially unseen con-text. Existing approaches utilize Neural OT and largely rely on Brenier's theorem. Here, we propose a first-of-its-kind quantum computing formulation for amortized optimization of contextualized transportation plans. We exploit a direct link between doubly stochastic matrices and unitary operators thus finding a natural connection between OT and quantum computation. We verify our method on synthetic and real data, by predicting variations in cell type distributions parameterized through drug dosage as context. Our comparisons to several baselines reveal that our method can capture dose-induced variations in cell distributions, even to some extent when dosages are extrapolated and sometimes with performance similar to the best classical models. In summary, this is a first step toward learning to predict contextualized transportation plans through quantum.
Language models in molecular discovery
Sep 28, 2023Abstract:The success of language models, especially transformer-based architectures, has trickled into other domains giving rise to "scientific language models" that operate on small molecules, proteins or polymers. In chemistry, language models contribute to accelerating the molecule discovery cycle as evidenced by promising recent findings in early-stage drug discovery. Here, we review the role of language models in molecular discovery, underlining their strength in de novo drug design, property prediction and reaction chemistry. We highlight valuable open-source software assets thus lowering the entry barrier to the field of scientific language modeling. Last, we sketch a vision for future molecular design that combines a chatbot interface with access to computational chemistry tools. Our contribution serves as a valuable resource for researchers, chemists, and AI enthusiasts interested in understanding how language models can and will be used to accelerate chemical discovery.
Unifying Molecular and Textual Representations via Multi-task Language Modelling
Jan 29, 2023Abstract:The recent advances in neural language models have also been successfully applied to the field of chemistry, offering generative solutions for classical problems in molecular design and synthesis planning. These new methods have the potential to optimize laboratory operations and fuel a new era of data-driven automation in scientific discovery. However, specialized models are still typically required for each task, leading to the need for problem-specific fine-tuning and neglecting task interrelations. The main obstacle in this field is the lack of a unified representation between natural language and chemical representations, complicating and limiting human-machine interaction. Here, we propose a multi-domain, multi-task language model to solve a wide range of tasks in both the chemical and natural language domains. By leveraging multi-task learning, our model can handle chemical and natural language concurrently, without requiring expensive pre-training on single domains or task-specific models. Interestingly, sharing weights across domains remarkably improves our model when benchmarked against state-of-the-art baselines on single-domain and cross-domain tasks. In particular, sharing information across domains and tasks gives rise to large improvements in cross-domain tasks, the magnitude of which increase with scale, as measured by more than a dozen of relevant metrics. Our work suggests that such models can robustly and efficiently accelerate discovery in physical sciences by superseding problem-specific fine-tuning and enhancing human-model interactions.
Domain-agnostic and Multi-level Evaluation of Generative Models
Jan 20, 2023



Abstract:While the capabilities of generative models heavily improved in different domains (images, text, graphs, molecules, etc.), their evaluation metrics largely remain based on simplified quantities or manual inspection with limited practicality. To this end, we propose a framework for Multi-level Performance Evaluation of Generative mOdels (MPEGO), which could be employed across different domains. MPEGO aims to quantify generation performance hierarchically, starting from a sub-feature-based low-level evaluation to a global features-based high-level evaluation. MPEGO offers great customizability as the employed features are entirely user-driven and can thus be highly domain/problem-specific while being arbitrarily complex (e.g., outcomes of experimental procedures). We validate MPEGO using multiple generative models across several datasets from the material discovery domain. An ablation study is conducted to study the plausibility of intermediate steps in MPEGO. Results demonstrate that MPEGO provides a flexible, user-driven, and multi-level evaluation framework, with practical insights on the generation quality. The framework, source code, and experiments will be available at https://github.com/GT4SD/mpego.
GT4SD: Generative Toolkit for Scientific Discovery
Jul 08, 2022
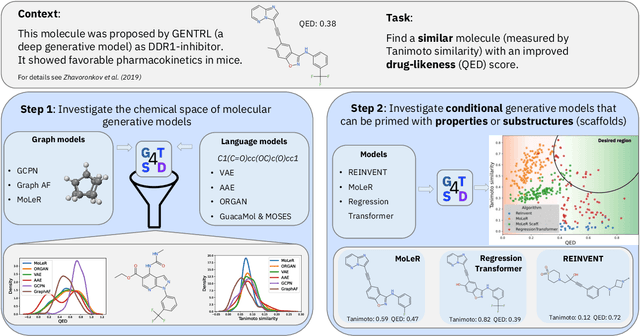
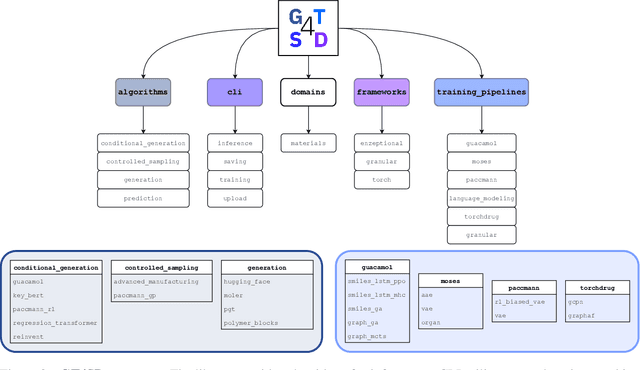
Abstract:With the growing availability of data within various scientific domains, generative models hold enormous potential to accelerate scientific discovery at every step of the scientific method. Perhaps their most valuable application lies in the speeding up of what has traditionally been the slowest and most challenging step of coming up with a hypothesis. Powerful representations are now being learned from large volumes of data to generate novel hypotheses, which is making a big impact on scientific discovery applications ranging from material design to drug discovery. The GT4SD (https://github.com/GT4SD/gt4sd-core) is an extensible open-source library that enables scientists, developers and researchers to train and use state-of-the-art generative models for hypothesis generation in scientific discovery. GT4SD supports a variety of uses of generative models across material science and drug discovery, including molecule discovery and design based on properties related to target proteins, omic profiles, scaffold distances, binding energies and more.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge