Jana Lipkova
A Learnable Prior Improves Inverse Tumor Growth Modeling
Mar 07, 2024



Abstract:Biophysical modeling, particularly involving partial differential equations (PDEs), offers significant potential for tailoring disease treatment protocols to individual patients. However, the inverse problem-solving aspect of these models presents a substantial challenge, either due to the high computational requirements of model-based approaches or the limited robustness of deep learning (DL) methods. We propose a novel framework that leverages the unique strengths of both approaches in a synergistic manner. Our method incorporates a DL ensemble for initial parameter estimation, facilitating efficient downstream evolutionary sampling initialized with this DL-based prior. We showcase the effectiveness of integrating a rapid deep-learning algorithm with a high-precision evolution strategy in estimating brain tumor cell concentrations from magnetic resonance images. The DL-Prior plays a pivotal role, significantly constraining the effective sampling-parameter space. This reduction results in a fivefold convergence acceleration and a Dice-score of 95%
Learn-Morph-Infer: a new way of solving the inverse problem for brain tumor modeling
Nov 07, 2021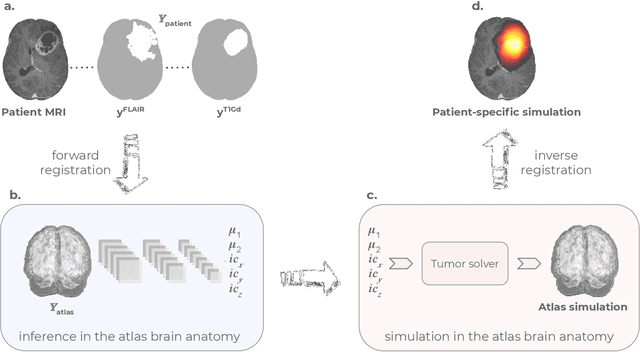
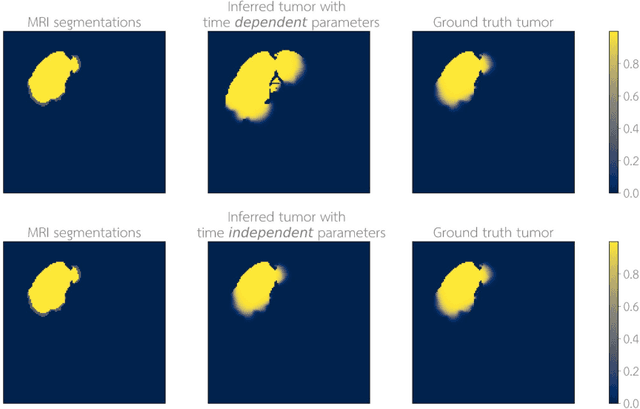
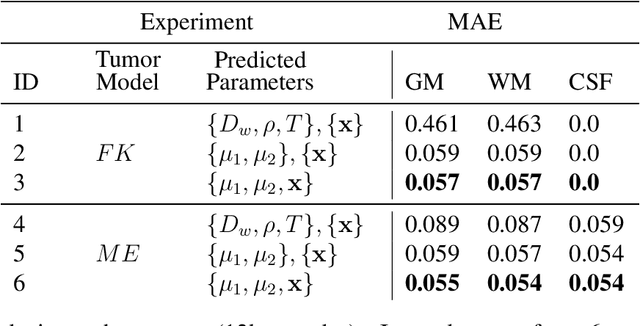
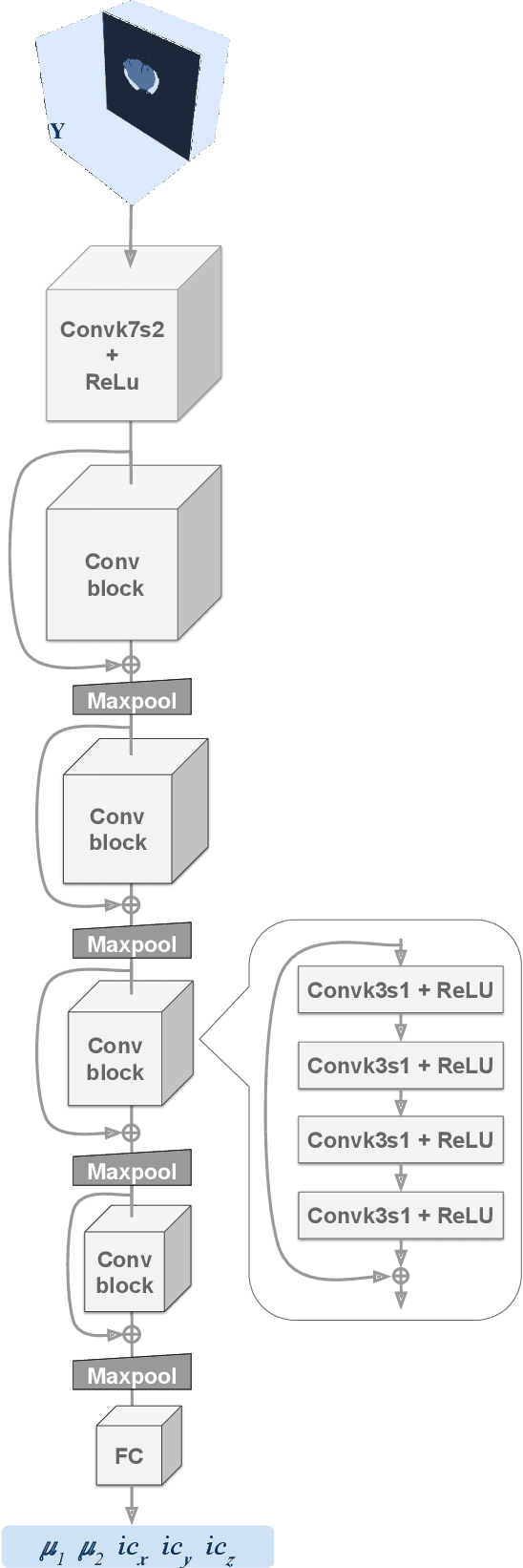

Abstract:Current treatment planning of patients diagnosed with brain tumor could significantly benefit by accessing the spatial distribution of tumor cell concentration. Existing diagnostic modalities, such as magnetic-resonance imaging (MRI), contrast sufficiently well areas of high cell density. However, they do not portray areas of low concentration, which can often serve as a source for the secondary appearance of the tumor after treatment. Numerical simulations of tumor growth could complement imaging information by providing estimates of full spatial distributions of tumor cells. Over recent years a corpus of literature on medical image-based tumor modeling was published. It includes different mathematical formalisms describing the forward tumor growth model. Alongside, various parametric inference schemes were developed to perform an efficient tumor model personalization, i.e. solving the inverse problem. However, the unifying drawback of all existing approaches is the time complexity of the model personalization that prohibits a potential integration of the modeling into clinical settings. In this work, we introduce a methodology for inferring patient-specific spatial distribution of brain tumor from T1Gd and FLAIR MRI medical scans. Coined as \textit{Learn-Morph-Infer} the method achieves real-time performance in the order of minutes on widely available hardware and the compute time is stable across tumor models of different complexity, such as reaction-diffusion and reaction-advection-diffusion models. We believe the proposed inverse solution approach not only bridges the way for clinical translation of brain tumor personalization but can also be adopted to other scientific and engineering domains.
Algorithm Fairness in AI for Medicine and Healthcare
Oct 01, 2021
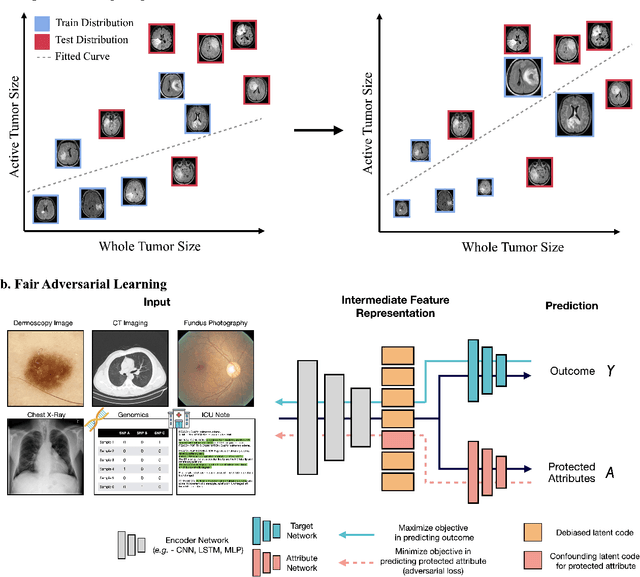

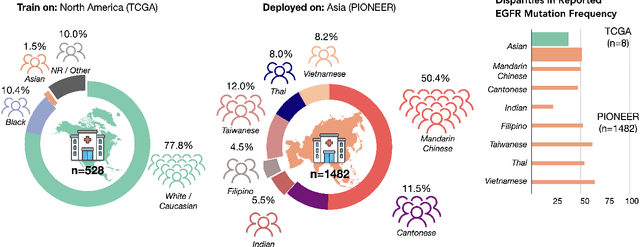
Abstract:In the current development and deployment of many artificial intelligence (AI) systems in healthcare, algorithm fairness is a challenging problem in delivering equitable care. Recent evaluation of AI models stratified across race sub-populations have revealed enormous inequalities in how patients are diagnosed, given treatments, and billed for healthcare costs. In this perspective article, we summarize the intersectional field of fairness in machine learning through the context of current issues in healthcare, outline how algorithmic biases (e.g. - image acquisition, genetic variation, intra-observer labeling variability) arise in current clinical workflows and their resulting healthcare disparities. Lastly, we also review emerging strategies for mitigating bias via decentralized learning, disentanglement, and model explainability.
Semi-Implicit Neural Solver for Time-dependent Partial Differential Equations
Sep 03, 2021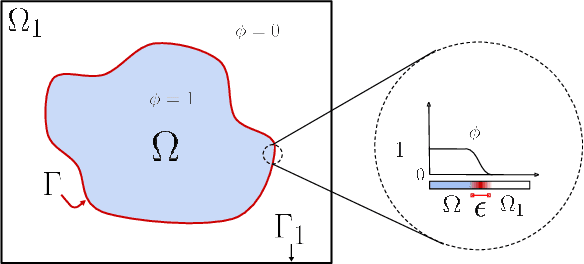
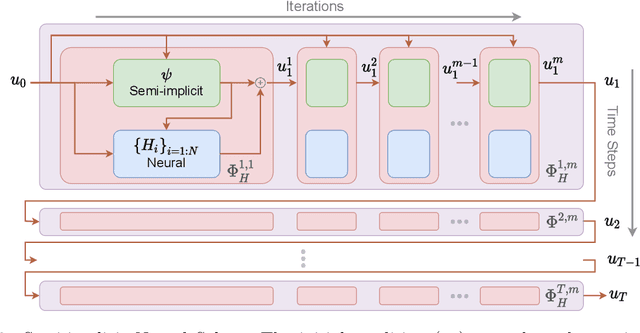
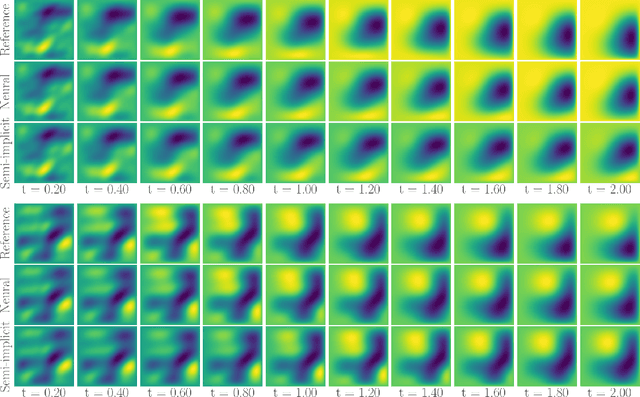
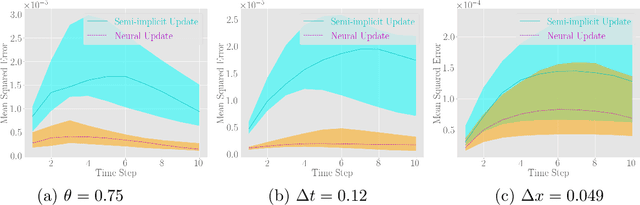
Abstract:Fast and accurate solutions of time-dependent partial differential equations (PDEs) are of pivotal interest to many research fields, including physics, engineering, and biology. Generally, implicit/semi-implicit schemes are preferred over explicit ones to improve stability and correctness. However, existing semi-implicit methods are usually iterative and employ a general-purpose solver, which may be sub-optimal for a specific class of PDEs. In this paper, we propose a neural solver to learn an optimal iterative scheme in a data-driven fashion for any class of PDEs. Specifically, we modify a single iteration of a semi-implicit solver using a deep neural network. We provide theoretical guarantees for the correctness and convergence of neural solvers analogous to conventional iterative solvers. In addition to the commonly used Dirichlet boundary condition, we adopt a diffuse domain approach to incorporate a diverse type of boundary conditions, e.g., Neumann. We show that the proposed neural solver can go beyond linear PDEs and applies to a class of non-linear PDEs, where the non-linear component is non-stiff. We demonstrate the efficacy of our method on 2D and 3D scenarios. To this end, we show how our model generalizes to parameter settings, which are different from training; and achieves faster convergence than semi-implicit schemes.
Pan-Cancer Integrative Histology-Genomic Analysis via Interpretable Multimodal Deep Learning
Aug 04, 2021
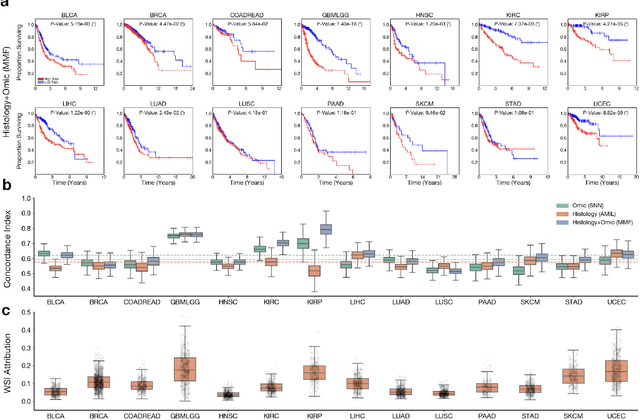
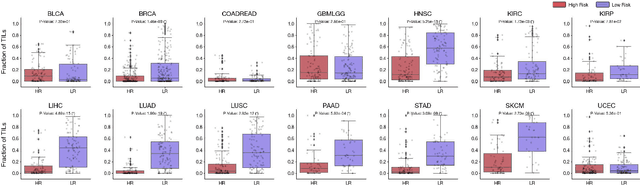
Abstract:The rapidly emerging field of deep learning-based computational pathology has demonstrated promise in developing objective prognostic models from histology whole slide images. However, most prognostic models are either based on histology or genomics alone and do not address how histology and genomics can be integrated to develop joint image-omic prognostic models. Additionally identifying explainable morphological and molecular descriptors from these models that govern such prognosis is of interest. We used multimodal deep learning to integrate gigapixel whole slide pathology images, RNA-seq abundance, copy number variation, and mutation data from 5,720 patients across 14 major cancer types. Our interpretable, weakly-supervised, multimodal deep learning algorithm is able to fuse these heterogeneous modalities for predicting outcomes and discover prognostic features from these modalities that corroborate with poor and favorable outcomes via multimodal interpretability. We compared our model with unimodal deep learning models trained on histology slides and molecular profiles alone, and demonstrate performance increase in risk stratification on 9 out of 14 cancers. In addition, we analyze morphologic and molecular markers responsible for prognostic predictions across all cancer types. All analyzed data, including morphological and molecular correlates of patient prognosis across the 14 cancer types at a disease and patient level are presented in an interactive open-access database (http://pancancer.mahmoodlab.org) to allow for further exploration and prognostic biomarker discovery. To validate that these model explanations are prognostic, we further analyzed high attention morphological regions in WSIs, which indicates that tumor-infiltrating lymphocyte presence corroborates with favorable cancer prognosis on 9 out of 14 cancer types studied.
Federated Learning for Computational Pathology on Gigapixel Whole Slide Images
Sep 23, 2020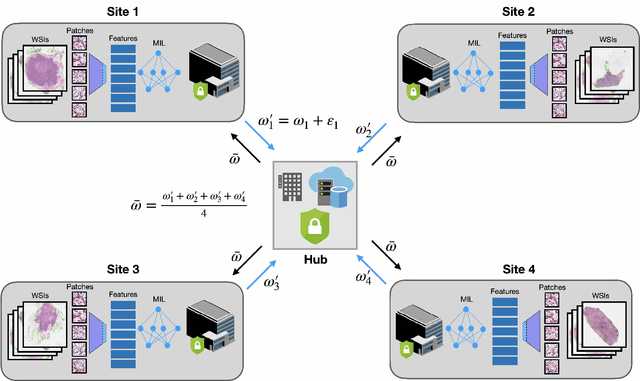
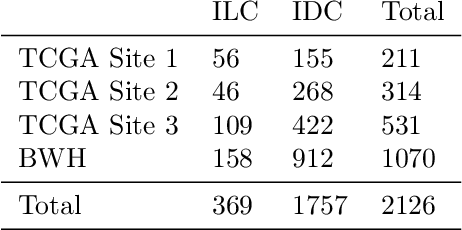
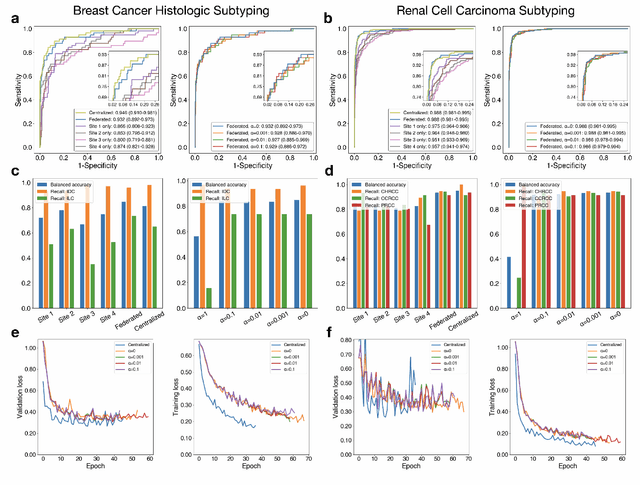
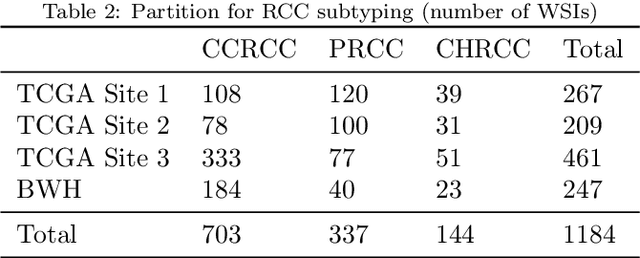
Abstract:Deep Learning-based computational pathology algorithms have demonstrated profound ability to excel in a wide array of tasks that range from characterization of well known morphological phenotypes to predicting non-human-identifiable features from histology such as molecular alterations. However, the development of robust, adaptable, and accurate deep learning-based models often rely on the collection and time-costly curation large high-quality annotated training data that should ideally come from diverse sources and patient populations to cater for the heterogeneity that exists in such datasets. Multi-centric and collaborative integration of medical data across multiple institutions can naturally help overcome this challenge and boost the model performance but is limited by privacy concerns amongst other difficulties that may arise in the complex data sharing process as models scale towards using hundreds of thousands of gigapixel whole slide images. In this paper, we introduce privacy-preserving federated learning for gigapixel whole slide images in computational pathology using weakly-supervised attention multiple instance learning and differential privacy. We evaluated our approach on two different diagnostic problems using thousands of histology whole slide images with only slide-level labels. Additionally, we present a weakly-supervised learning framework for survival prediction and patient stratification from whole slide images and demonstrate its effectiveness in a federated setting. Our results show that using federated learning, we can effectively develop accurate weakly supervised deep learning models from distributed data silos without direct data sharing and its associated complexities, while also preserving differential privacy using randomized noise generation.
Deep Learning-based Computational Pathology Predicts Origins for Cancers of Unknown Primary
Jun 29, 2020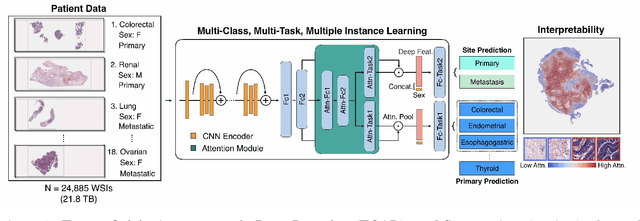
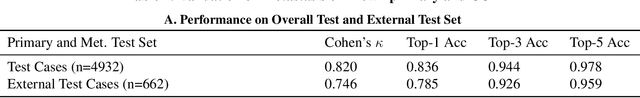
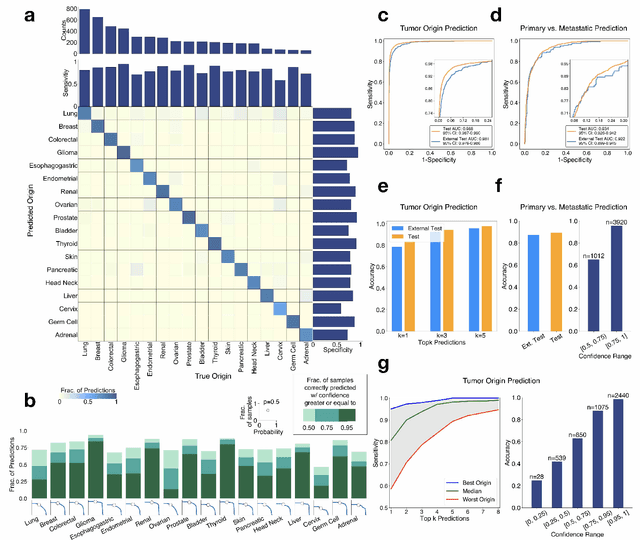
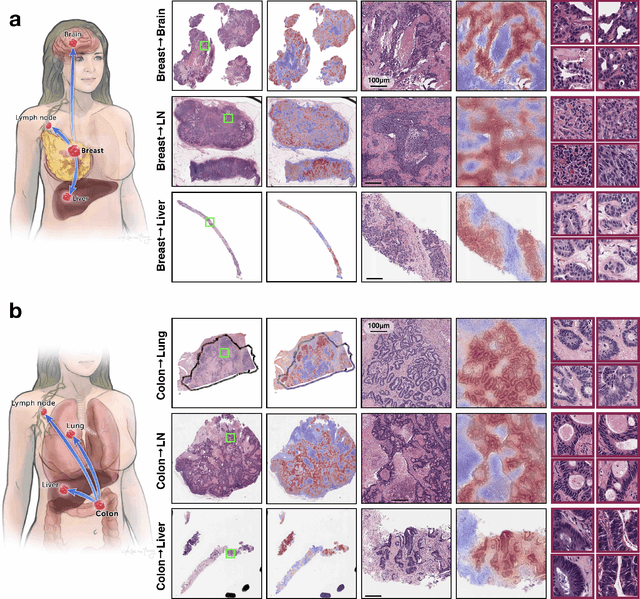
Abstract:Cancer of unknown primary (CUP) is an enigmatic group of diagnoses where the primary anatomical site of tumor origin cannot be determined. This poses a significant challenge since modern therapeutics such as chemotherapy regimen and immune checkpoint inhibitors are specific to the primary tumor. Recent work has focused on using genomics and transcriptomics for identification of tumor origins. However, genomic testing is not conducted for every patient and lacks clinical penetration in low resource settings. Herein, to overcome these challenges, we present a deep learning-based computational pathology algorithm-TOAD-that can provide a differential diagnosis for CUP using routinely acquired histology slides. We used 17,486 gigapixel whole slide images with known primaries spread over 18 common origins to train a multi-task deep model to simultaneously identify the tumor as primary or metastatic and predict its site of origin. We tested our model on an internal test set of 4,932 cases with known primaries and achieved a top-1 accuracy of 0.84, a top-3 accuracy of 0.94 while on our external test set of 662 cases from 202 different hospitals, it achieved a top-1 and top-3 accuracy of 0.79 and 0.93 respectively. We further curated a dataset of 717 CUP cases from 151 different medical centers and identified a subset of 290 cases for which a differential diagnosis was assigned. Our model predictions resulted in concordance for 50% of cases (\k{appa}=0.4 when adjusted for agreement by chance) and a top-3 agreement of 75%. Our proposed method can be used as an assistive tool to assign differential diagnosis to complicated metastatic and CUP cases and could be used in conjunction with or in lieu of immunohistochemical analysis and extensive diagnostic work-ups to reduce the occurrence of CUP.
Red-GAN: Attacking class imbalance via conditioned generation. Yet another medical imaging perspective
Apr 30, 2020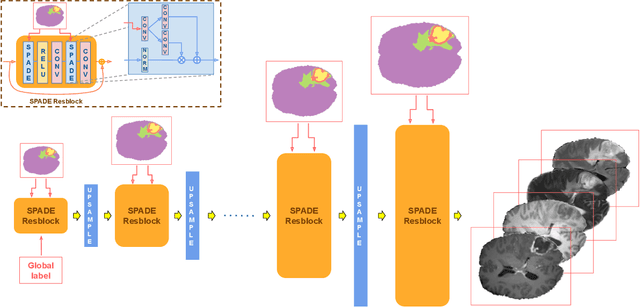

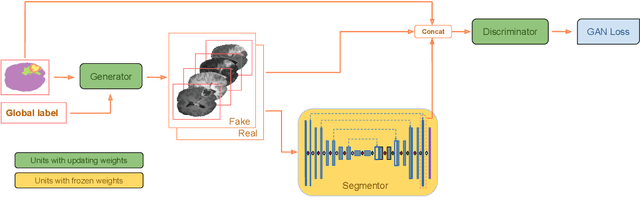
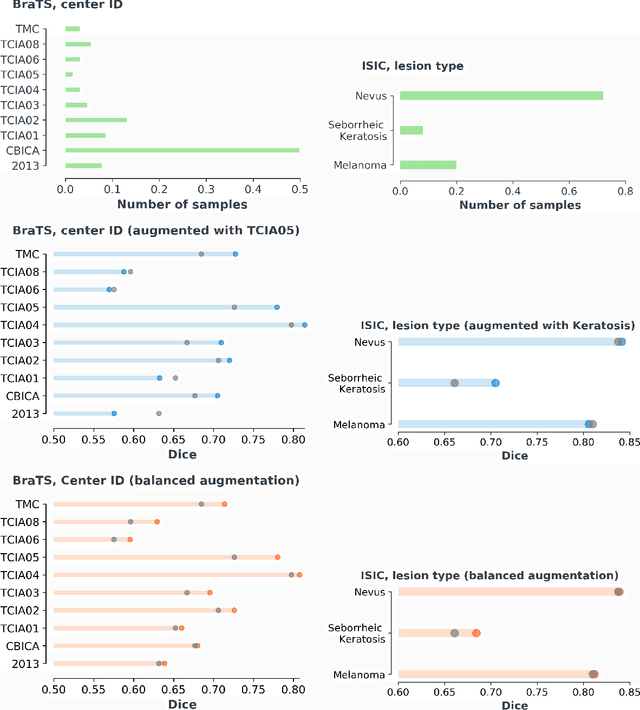
Abstract:Exploiting learning algorithms under scarce data regimes is a limitation and a reality of the medical imaging field. In an attempt to mitigate the problem, we propose a data augmentation protocol based on generative adversarial networks. We condition the networks at a pixel-level (segmentation mask) and at a global-level information (acquisition environment or lesion type). Such conditioning provides immediate access to the image-label pairs while controlling global class specific appearance of the synthesized images. To stimulate synthesis of the features relevant for the segmentation task, an additional passive player in a form of segmentor is introduced into the the adversarial game. We validate the approach on two medical datasets: BraTS, ISIC. By controlling the class distribution through injection of synthetic images into the training set we achieve control over the accuracy levels of the datasets' classes.
Implicit Neural Solver for Time-dependent Linear PDEs with Convergence Guarantee
Oct 09, 2019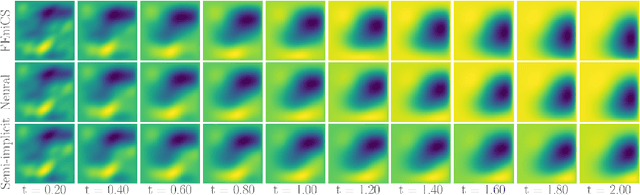

Abstract:Fast and accurate solution of time-dependent partial differential equations (PDEs) is of key interest in many research fields including physics, engineering, and biology. Generally, implicit schemes are preferred over the explicit ones for better stability and correctness. The existing implicit schemes are usually iterative and employ a general-purpose solver which may be sub-optimal for a specific class of PDEs. In this paper, we propose a neural solver to learn an optimal iterative scheme for a class of PDEs, in a data-driven fashion. We achieve this goal by modifying an iteration of an existing semi-implicit solver using a deep neural network. Further, we provide theoretical proof that our approach preserves the correctness and convergence guarantees provided by the existing iterative-solvers. We also demonstrate that our model generalizes to a different parameter setting than the one seen during training and achieves faster convergence compared to the semi-implicit schemes.
Neural parameters estimation for brain tumor growth modeling
Jul 01, 2019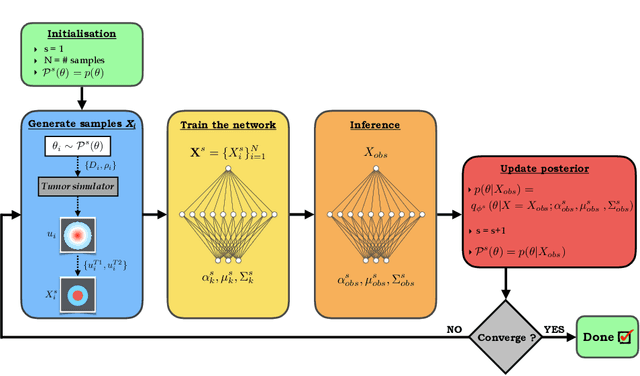
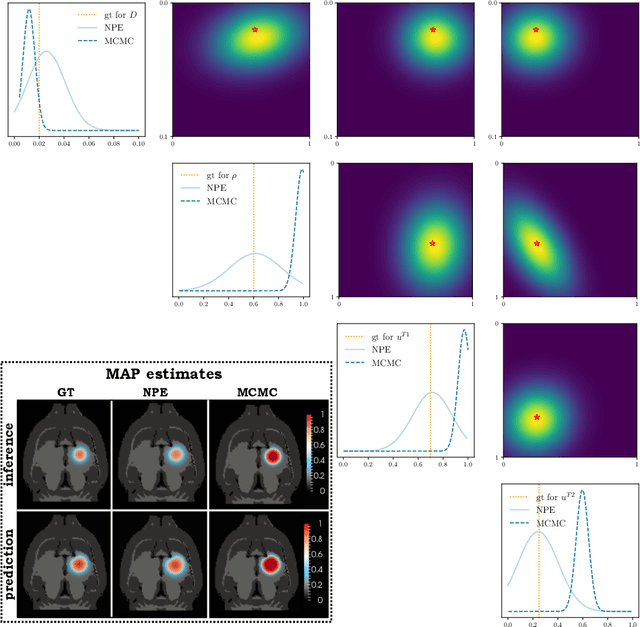

Abstract:Understanding the dynamics of brain tumor progression is essential for optimal treatment planning. Cast in a mathematical formulation, it is typically viewed as evaluation of a system of partial differential equations, wherein the physiological processes that govern the growth of the tumor are considered. To personalize the model, i.e. find a relevant set of parameters, with respect to the tumor dynamics of a particular patient, the model is informed from empirical data, e.g., medical images obtained from diagnostic modalities, such as magnetic-resonance imaging. Existing model-observation coupling schemes require a large number of forward integrations of the biophysical model and rely on simplifying assumption on the functional form, linking the output of the model with the image information. In this work, we propose a learning-based technique for the estimation of tumor growth model parameters from medical scans. The technique allows for explicit evaluation of the posterior distribution of the parameters by sequentially training a mixture-density network, relaxing the constraint on the functional form and reducing the number of samples necessary to propagate through the forward model for the estimation. We test the method on synthetic and real scans of rats injected with brain tumors to calibrate the model and to predict tumor progression.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge