Judy J. Wang
A General-Purpose Self-Supervised Model for Computational Pathology
Aug 29, 2023

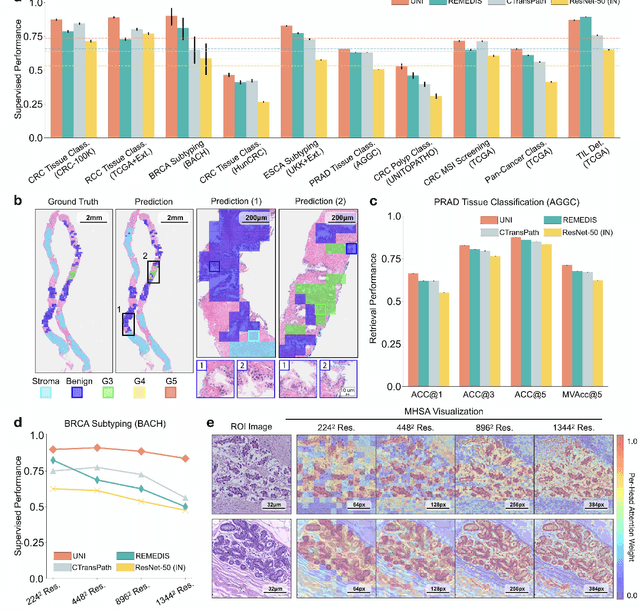

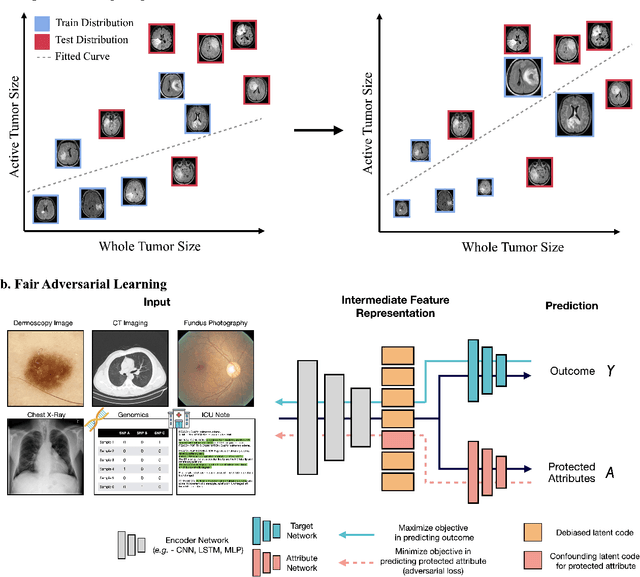
Abstract:Tissue phenotyping is a fundamental computational pathology (CPath) task in learning objective characterizations of histopathologic biomarkers in anatomic pathology. However, whole-slide imaging (WSI) poses a complex computer vision problem in which the large-scale image resolutions of WSIs and the enormous diversity of morphological phenotypes preclude large-scale data annotation. Current efforts have proposed using pretrained image encoders with either transfer learning from natural image datasets or self-supervised pretraining on publicly-available histopathology datasets, but have not been extensively developed and evaluated across diverse tissue types at scale. We introduce UNI, a general-purpose self-supervised model for pathology, pretrained using over 100 million tissue patches from over 100,000 diagnostic haematoxylin and eosin-stained WSIs across 20 major tissue types, and evaluated on 33 representative CPath clinical tasks in CPath of varying diagnostic difficulties. In addition to outperforming previous state-of-the-art models, we demonstrate new modeling capabilities in CPath such as resolution-agnostic tissue classification, slide classification using few-shot class prototypes, and disease subtyping generalization in classifying up to 108 cancer types in the OncoTree code classification system. UNI advances unsupervised representation learning at scale in CPath in terms of both pretraining data and downstream evaluation, enabling data-efficient AI models that can generalize and transfer to a gamut of diagnostically-challenging tasks and clinical workflows in anatomic pathology.
Algorithm Fairness in AI for Medicine and Healthcare
Oct 01, 2021

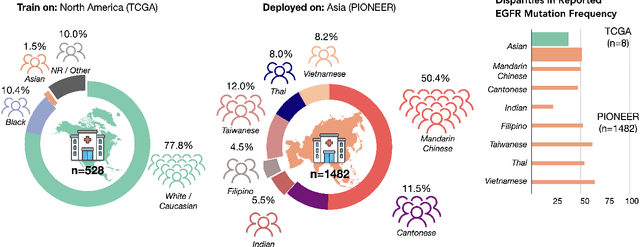
Abstract:In the current development and deployment of many artificial intelligence (AI) systems in healthcare, algorithm fairness is a challenging problem in delivering equitable care. Recent evaluation of AI models stratified across race sub-populations have revealed enormous inequalities in how patients are diagnosed, given treatments, and billed for healthcare costs. In this perspective article, we summarize the intersectional field of fairness in machine learning through the context of current issues in healthcare, outline how algorithmic biases (e.g. - image acquisition, genetic variation, intra-observer labeling variability) arise in current clinical workflows and their resulting healthcare disparities. Lastly, we also review emerging strategies for mitigating bias via decentralized learning, disentanglement, and model explainability.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge