Haichuan Zhang
CloneShield: A Framework for Universal Perturbation Against Zero-Shot Voice Cloning
May 25, 2025Abstract:Recent breakthroughs in text-to-speech (TTS) voice cloning have raised serious privacy concerns, allowing highly accurate vocal identity replication from just a few seconds of reference audio, while retaining the speaker's vocal authenticity. In this paper, we introduce CloneShield, a universal time-domain adversarial perturbation framework specifically designed to defend against zero-shot voice cloning. Our method provides protection that is robust across speakers and utterances, without requiring any prior knowledge of the synthesized text. We formulate perturbation generation as a multi-objective optimization problem, and propose Multi-Gradient Descent Algorithm (MGDA) to ensure the robust protection across diverse utterances. To preserve natural auditory perception for users, we decompose the adversarial perturbation via Mel-spectrogram representations and fine-tune it for each sample. This design ensures imperceptibility while maintaining strong degradation effects on zero-shot cloned outputs. Experiments on three state-of-the-art zero-shot TTS systems, five benchmark datasets and evaluations from 60 human listeners demonstrate that our method preserves near-original audio quality in protected inputs (PESQ = 3.90, SRS = 0.93) while substantially degrading both speaker similarity and speech quality in cloned samples (PESQ = 1.07, SRS = 0.08).
Deep, convergent, unrolled half-quadratic splitting for image deconvolution
Feb 25, 2024



Abstract:In recent years, algorithm unrolling has emerged as a powerful technique for designing interpretable neural networks based on iterative algorithms. Imaging inverse problems have particularly benefited from unrolling-based deep network design since many traditional model-based approaches rely on iterative optimization. Despite exciting progress, typical unrolling approaches heuristically design layer-specific convolution weights to improve performance. Crucially, convergence properties of the underlying iterative algorithm are lost once layer-specific parameters are learned from training data. We propose an unrolling technique that breaks the trade-off between retaining algorithm properties while simultaneously enhancing performance. We focus on image deblurring and unrolling the widely-applied Half-Quadratic Splitting (HQS) algorithm. We develop a new parametrization scheme which enforces layer-specific parameters to asymptotically approach certain fixed points. Through extensive experimental studies, we verify that our approach achieves competitive performance with state-of-the-art unrolled layer-specific learning and significantly improves over the traditional HQS algorithm. We further establish convergence of the proposed unrolled network as the number of layers approaches infinity, and characterize its convergence rate. Our experimental verification involves simulations that validate the analytical results as well as comparison with state-of-the-art non-blind deblurring techniques on benchmark datasets. The merits of the proposed convergent unrolled network are established over competing alternatives, especially in the regime of limited training.
PIMIP: An Open Source Platform for Pathology Information Management and Integration
Nov 09, 2021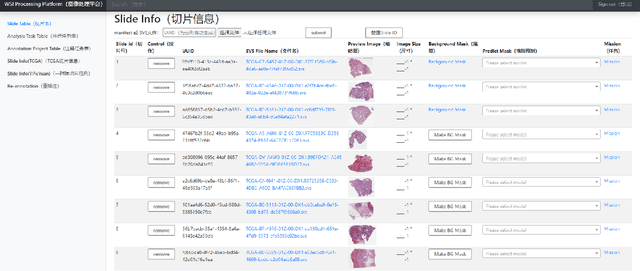
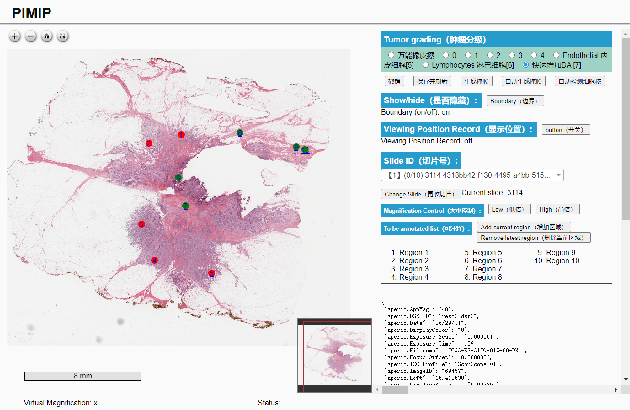
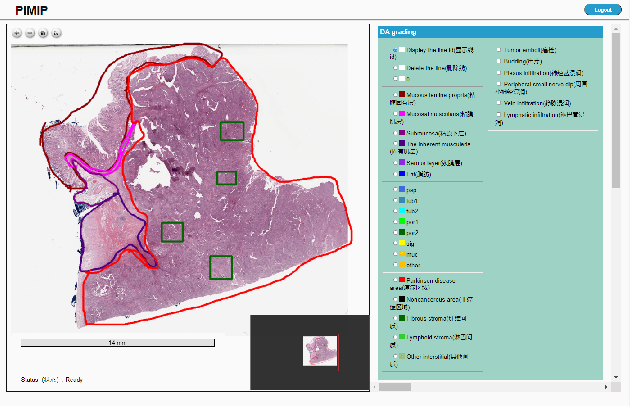
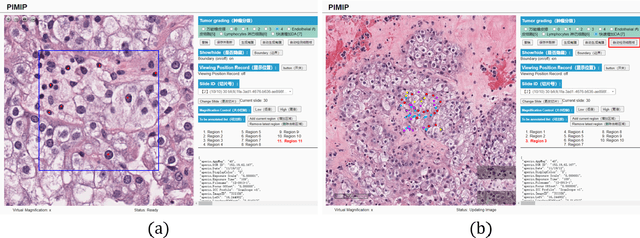
Abstract:Digital pathology plays a crucial role in the development of artificial intelligence in the medical field. The digital pathology platform can make the pathological resources digital and networked, and realize the permanent storage of visual data and the synchronous browsing processing without the limitation of time and space. It has been widely used in various fields of pathology. However, there is still a lack of an open and universal digital pathology platform to assist doctors in the management and analysis of digital pathological sections, as well as the management and structured description of relevant patient information. Most platforms cannot integrate image viewing, annotation and analysis, and text information management. To solve the above problems, we propose a comprehensive and extensible platform PIMIP. Our PIMIP has developed the image annotation functions based on the visualization of digital pathological sections. Our annotation functions support multi-user collaborative annotation and multi-device annotation, and realize the automation of some annotation tasks. In the annotation task, we invited a professional pathologist for guidance. We introduce a machine learning module for image analysis. The data we collected included public data from local hospitals and clinical examples. Our platform is more clinical and suitable for clinical use. In addition to image data, we also structured the management and display of text information. So our platform is comprehensive. The platform framework is built in a modular way to support users to add machine learning modules independently, which makes our platform extensible.
A Personalized Diagnostic Generation Framework Based on Multi-source Heterogeneous Data
Oct 26, 2021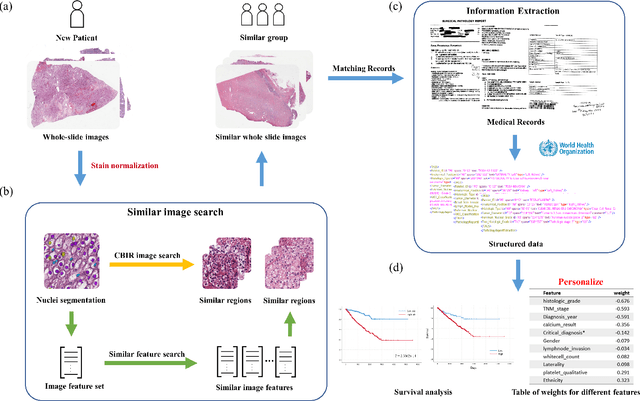
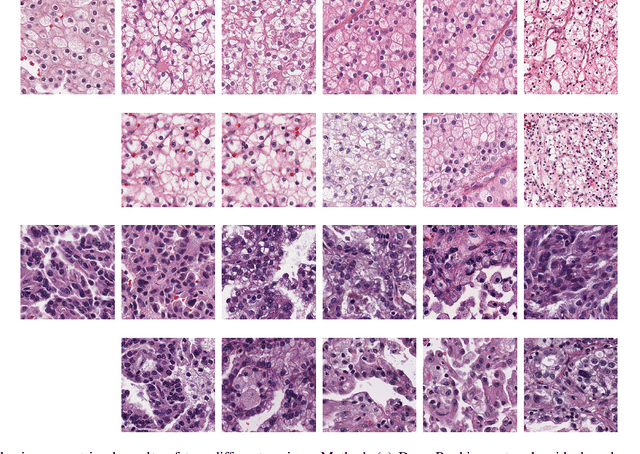
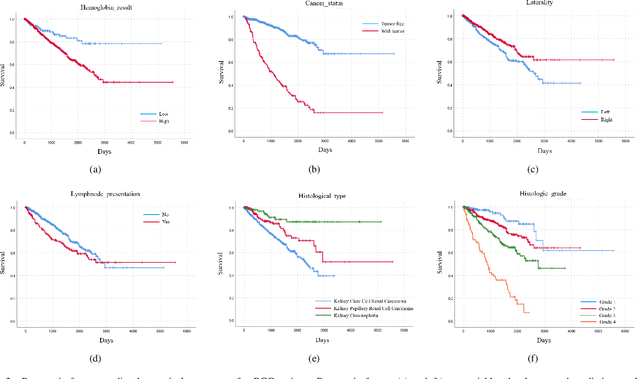
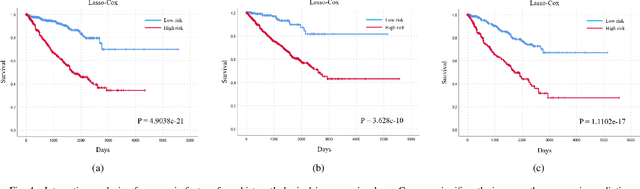
Abstract:Personalized diagnoses have not been possible due to sear amount of data pathologists have to bear during the day-to-day routine. This lead to the current generalized standards that are being continuously updated as new findings are reported. It is noticeable that these effective standards are developed based on a multi-source heterogeneous data, including whole-slide images and pathology and clinical reports. In this study, we propose a framework that combines pathological images and medical reports to generate a personalized diagnosis result for individual patient. We use nuclei-level image feature similarity and content-based deep learning method to search for a personalized group of population with similar pathological characteristics, extract structured prognostic information from descriptive pathology reports of the similar patient population, and assign importance of different prognostic factors to generate a personalized pathological diagnosis result. We use multi-source heterogeneous data from TCGA (The Cancer Genome Atlas) database. The result demonstrate that our framework matches the performance of pathologists in the diagnosis of renal cell carcinoma. This framework is designed to be generic, thus could be applied for other types of cancer. The weights could provide insights to the known prognostic factors and further guide more precise clinical treatment protocols.
A Precision Diagnostic Framework of Renal Cell Carcinoma on Whole-Slide Images using Deep Learning
Oct 26, 2021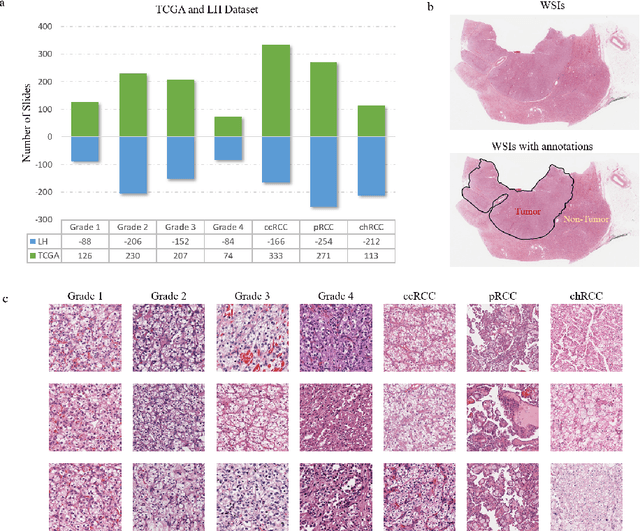
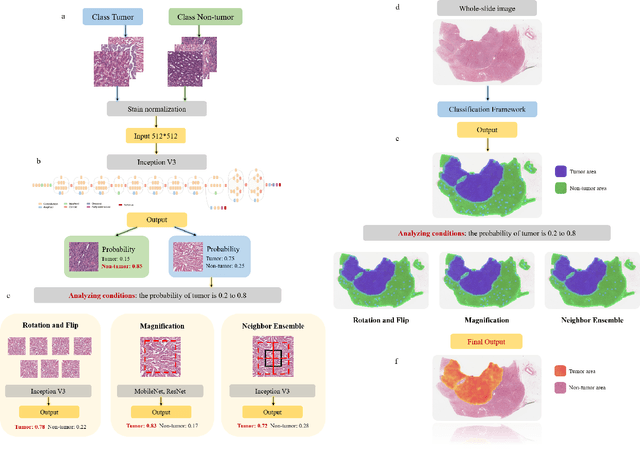
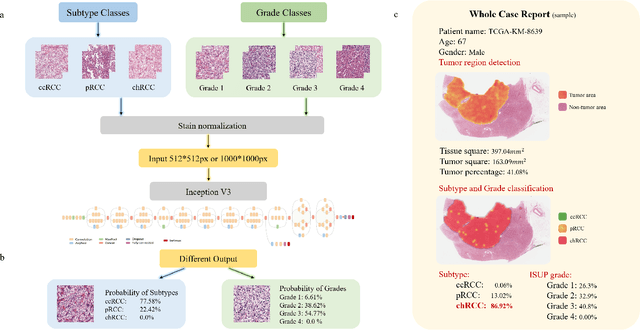
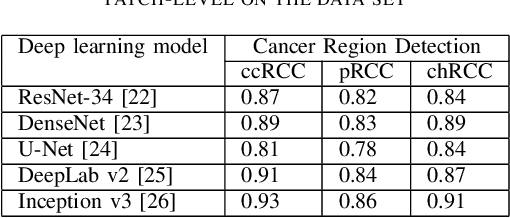
Abstract:Diagnostic pathology, which is the basis and gold standard of cancer diagnosis, provides essential information on the prognosis of the disease and vital evidence for clinical treatment. Tumor region detection, subtype and grade classification are the fundamental diagnostic indicators for renal cell carcinoma (RCC) in whole-slide images (WSIs). However, pathological diagnosis is subjective, differences in observation and diagnosis between pathologists is common in hospitals with inadequate diagnostic capacity. The main challenge for developing deep learning based RCC diagnostic system is the lack of large-scale datasets with precise annotations. In this work, we proposed a deep learning-based framework for analyzing histopathological images of patients with renal cell carcinoma, which has the potential to achieve pathologist-level accuracy in diagnosis. A deep convolutional neural network (InceptionV3) was trained on the high-quality annotated dataset of The Cancer Genome Atlas (TCGA) whole-slide histopathological image for accurate tumor area detection, classification of RCC subtypes, and ISUP grades classification of clear cell carcinoma subtypes. These results suggest that our framework can help pathologists in the detection of cancer region and classification of subtypes and grades, which could be applied to any cancer type, providing auxiliary diagnosis and promoting clinical consensus.
Nuclei Grading of Clear Cell Renal Cell Carcinoma in Histopathological Image by Composite High-Resolution Network
Jun 20, 2021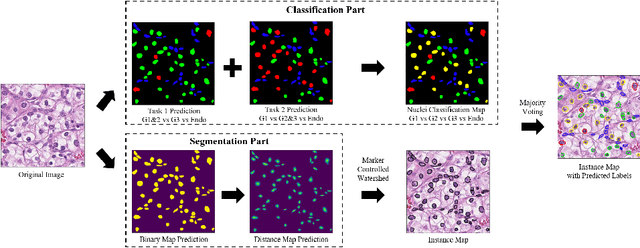
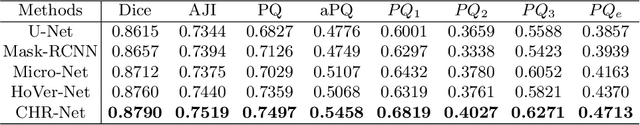
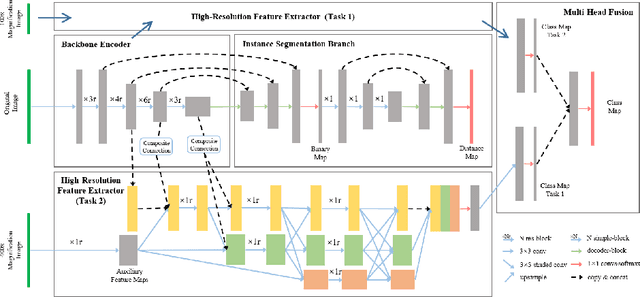
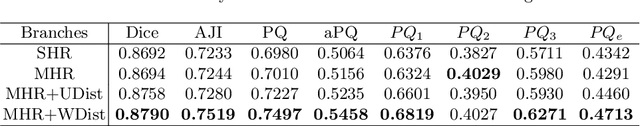
Abstract:The grade of clear cell renal cell carcinoma (ccRCC) is a critical prognostic factor, making ccRCC nuclei grading a crucial task in RCC pathology analysis. Computer-aided nuclei grading aims to improve pathologists' work efficiency while reducing their misdiagnosis rate by automatically identifying the grades of tumor nuclei within histopathological images. Such a task requires precisely segment and accurately classify the nuclei. However, most of the existing nuclei segmentation and classification methods can not handle the inter-class similarity property of nuclei grading, thus can not be directly applied to the ccRCC grading task. In this paper, we propose a Composite High-Resolution Network for ccRCC nuclei grading. Specifically, we propose a segmentation network called W-Net that can separate the clustered nuclei. Then, we recast the fine-grained classification of nuclei to two cross-category classification tasks, based on two high-resolution feature extractors (HRFEs) which are proposed for learning these two tasks. The two HRFEs share the same backbone encoder with W-Net by a composite connection so that meaningful features for the segmentation task can be inherited for the classification task. Last, a head-fusion block is applied to generate the predicted label of each nucleus. Furthermore, we introduce a dataset for ccRCC nuclei grading, containing 1000 image patches with 70945 annotated nuclei. We demonstrate that our proposed method achieves state-of-the-art performance compared to existing methods on this large ccRCC grading dataset.
OpenHI2 -- Open source histopathological image platform
Jan 15, 2020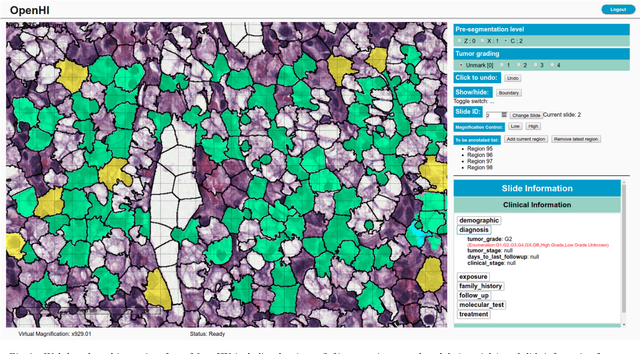
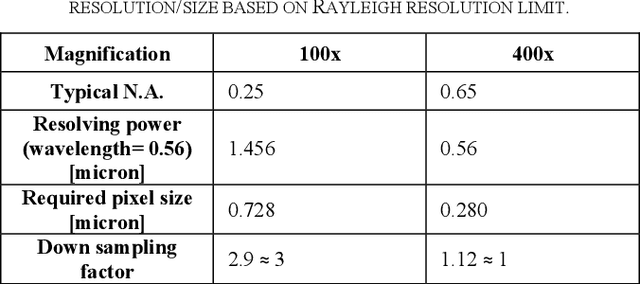
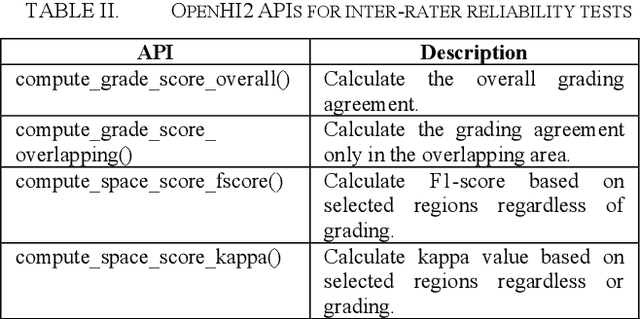
Abstract:Transition from conventional to digital pathology requires a new category of biomedical informatic infrastructure which could facilitate delicate pathological routine. Pathological diagnoses are sensitive to many external factors and is known to be subjective. Only systems that can meet strict requirements in pathology would be able to run along pathological routines and eventually digitized the study area, and the developed platform should comply with existing pathological routines and international standards. Currently, there are a number of available software tools which can perform histopathological tasks including virtual slide viewing, annotating, and basic image analysis, however, none of them can serve as a digital platform for pathology. Here we describe OpenHI2, an enhanced version Open Histopathological Image platform which is capable of supporting all basic pathological tasks and file formats; ready to be deployed in medical institutions on a standard server environment or cloud computing infrastructure. In this paper, we also describe the development decisions for the platform and propose solutions to overcome technical challenges so that OpenHI2 could be used as a platform for histopathological images. Further addition can be made to the platform since each component is modularized and fully documented. OpenHI2 is free, open-source, and available at https://gitlab.com/BioAI/OpenHI.
Effects of annotation granularity in deep learning models for histopathological images
Jan 14, 2020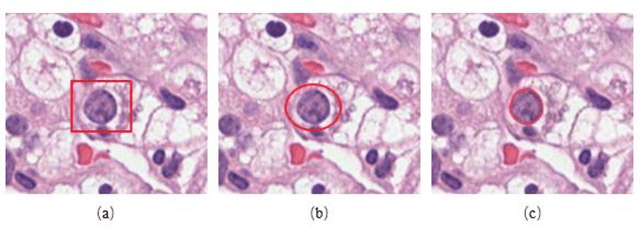
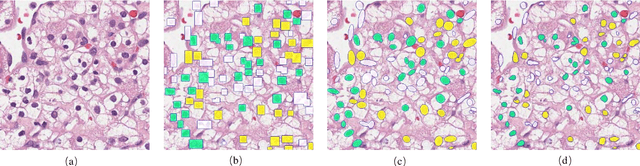
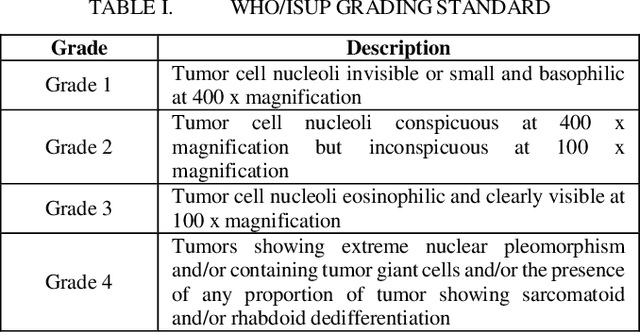

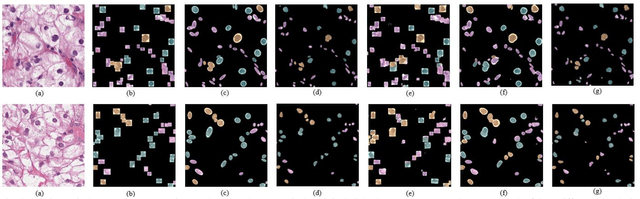

Abstract:Pathological is crucial to cancer diagnosis. Usually, Pathologists draw their conclusion based on observed cell and tissue structure on histology slides. Rapid development in machine learning, especially deep learning have established robust and accurate classifiers. They are being used to analyze histopathological slides and assist pathologists in diagnosis. Most machine learning systems rely heavily on annotated data sets to gain experiences and knowledge to correctly and accurately perform various tasks such as classification and segmentation. This work investigates different granularity of annotations in histopathological data set including image-wise, bounding box, ellipse-wise, and pixel-wise to verify the influence of annotation in pathological slide on deep learning models. We design corresponding experiments to test classification and segmentation performance of deep learning models based on annotations with different annotation granularity. In classification, state-of-the-art deep learning-based classifiers perform better when trained by pixel-wise annotation dataset. On average, precision, recall and F1-score improves by 7.87%, 8.83% and 7.85% respectively. Thus, it is suggested that finer granularity annotations are better utilized by deep learning algorithms in classification tasks. Similarly, semantic segmentation algorithms can achieve 8.33% better segmentation accuracy when trained by pixel-wise annotations. Our study shows not only that finer-grained annotation can improve the performance of deep learning models, but also help extracts more accurate phenotypic information from histopathological slides. Intelligence systems trained on granular annotations may help pathologists inspecting certain regions for better diagnosis. The compartmentalized prediction approach similar to this work may contribute to phenotype and genotype association studies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge