Axel Rominger
Artificial intelligence for simplified patient-centered dosimetry in radiopharmaceutical therapies
Oct 14, 2025Abstract:KEY WORDS: Artificial Intelligence (AI), Theranostics, Dosimetry, Radiopharmaceutical Therapy (RPT), Patient-friendly dosimetry KEY POINTS - The rapid evolution of radiopharmaceutical therapy (RPT) highlights the growing need for personalized and patient-centered dosimetry. - Artificial Intelligence (AI) offers solutions to the key limitations in current dosimetry calculations. - The main advances on AI for simplified dosimetry toward patient-friendly RPT are reviewed. - Future directions on the role of AI in RPT dosimetry are discussed.
Fed-NDIF: A Noise-Embedded Federated Diffusion Model For Low-Count Whole-Body PET Denoising
Mar 20, 2025



Abstract:Low-count positron emission tomography (LCPET) imaging can reduce patients' exposure to radiation but often suffers from increased image noise and reduced lesion detectability, necessitating effective denoising techniques. Diffusion models have shown promise in LCPET denoising for recovering degraded image quality. However, training such models requires large and diverse datasets, which are challenging to obtain in the medical domain. To address data scarcity and privacy concerns, we combine diffusion models with federated learning -- a decentralized training approach where models are trained individually at different sites, and their parameters are aggregated on a central server over multiple iterations. The variation in scanner types and image noise levels within and across institutions poses additional challenges for federated learning in LCPET denoising. In this study, we propose a novel noise-embedded federated learning diffusion model (Fed-NDIF) to address these challenges, leveraging a multicenter dataset and varying count levels. Our approach incorporates liver normalized standard deviation (NSTD) noise embedding into a 2.5D diffusion model and utilizes the Federated Averaging (FedAvg) algorithm to aggregate locally trained models into a global model, which is subsequently fine-tuned on local datasets to optimize performance and obtain personalized models. Extensive validation on datasets from the University of Bern, Ruijin Hospital in Shanghai, and Yale-New Haven Hospital demonstrates the superior performance of our method in enhancing image quality and improving lesion quantification. The Fed-NDIF model shows significant improvements in PSNR, SSIM, and NMSE of the entire 3D volume, as well as enhanced lesion detectability and quantification, compared to local diffusion models and federated UNet-based models.
Developing a PET/CT Foundation Model for Cross-Modal Anatomical and Functional Imaging
Mar 04, 2025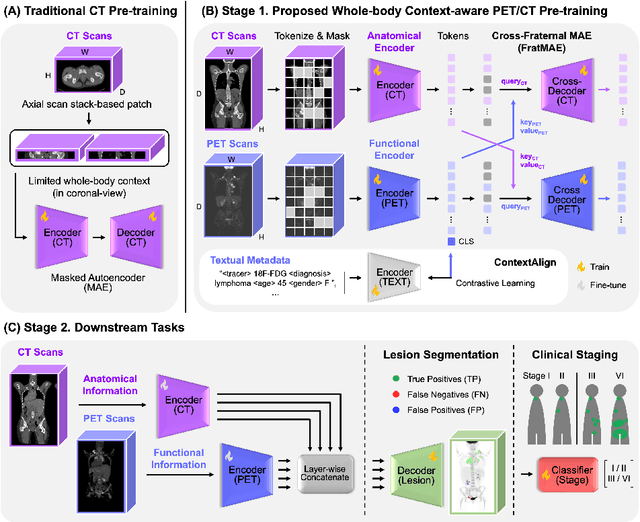
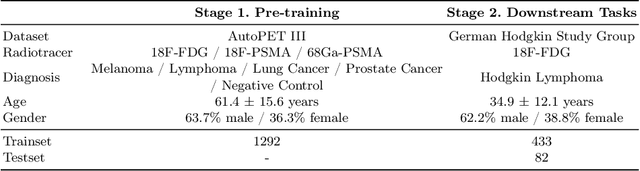
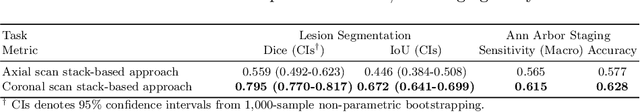


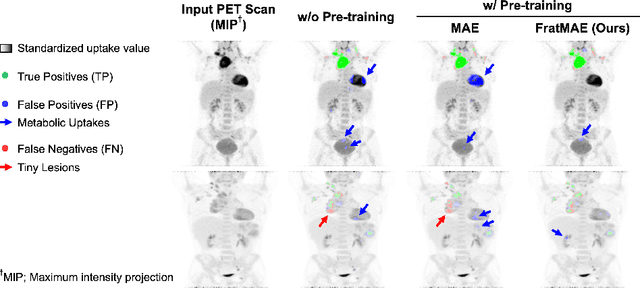
Abstract:In oncology, Positron Emission Tomography-Computed Tomography (PET/CT) is widely used in cancer diagnosis, staging, and treatment monitoring, as it combines anatomical details from CT with functional metabolic activity and molecular marker expression information from PET. However, existing artificial intelligence-driven PET/CT analyses rely predominantly on task-specific models trained from scratch or on limited datasets, limiting their generalizability and robustness. To address this, we propose a foundation model approach specifically designed for multimodal PET/CT imaging. We introduce the Cross-Fraternal Twin Masked Autoencoder (FratMAE), a novel framework that effectively integrates whole-body anatomical and functional or molecular information. FratMAE employs separate Vision Transformer (ViT) encoders for PET and CT scans, along with cross-attention decoders that enable synergistic interactions between modalities during masked autoencoder training. Additionally, it incorporates textual metadata to enhance PET representation learning. By pre-training on PET/CT datasets, FratMAE captures intricate cross-modal relationships and global uptake patterns, achieving superior performance on downstream tasks and demonstrating its potential as a generalizable foundation model.
LpQcM: Adaptable Lesion-Quantification-Consistent Modulation for Deep Learning Low-Count PET Image Denoising
Apr 27, 2024Abstract:Deep learning-based positron emission tomography (PET) image denoising offers the potential to reduce radiation exposure and scanning time by transforming low-count images into high-count equivalents. However, existing methods typically blur crucial details, leading to inaccurate lesion quantification. This paper proposes a lesion-perceived and quantification-consistent modulation (LpQcM) strategy for enhanced PET image denoising, via employing downstream lesion quantification analysis as auxiliary tools. The LpQcM is a plug-and-play design adaptable to a wide range of model architectures, modulating the sampling and optimization procedures of model training without adding any computational burden to the inference phase. Specifically, the LpQcM consists of two components, the lesion-perceived modulation (LpM) and the multiscale quantification-consistent modulation (QcM). The LpM enhances lesion contrast and visibility by allocating higher sampling weights and stricter loss criteria to lesion-present samples determined by an auxiliary segmentation network than lesion-absent ones. The QcM further emphasizes accuracy of quantification for both the mean and maximum standardized uptake value (SUVmean and SUVmax) across multiscale sub-regions throughout the entire image, thereby enhancing the overall image quality. Experiments conducted on large PET datasets from multiple centers and vendors, and varying noise levels demonstrated the LpQcM efficacy across various denoising frameworks. Compared to frameworks without LpQcM, the integration of LpQcM reduces the lesion SUVmean bias by 2.92% on average and increases the peak signal-to-noise ratio (PSNR) by 0.34 on average, for denoising images of extremely low-count levels below 10%.
FedFTN: Personalized Federated Learning with Deep Feature Transformation Network for Multi-institutional Low-count PET Denoising
Apr 02, 2023Abstract:Low-count PET is an efficient way to reduce radiation exposure and acquisition time, but the reconstructed images often suffer from low signal-to-noise ratio (SNR), thus affecting diagnosis and other downstream tasks. Recent advances in deep learning have shown great potential in improving low-count PET image quality, but acquiring a large, centralized, and diverse dataset from multiple institutions for training a robust model is difficult due to privacy and security concerns of patient data. Moreover, low-count PET data at different institutions may have different data distribution, thus requiring personalized models. While previous federated learning (FL) algorithms enable multi-institution collaborative training without the need of aggregating local data, addressing the large domain shift in the application of multi-institutional low-count PET denoising remains a challenge and is still highly under-explored. In this work, we propose FedFTN, a personalized federated learning strategy that addresses these challenges. FedFTN uses a local deep feature transformation network (FTN) to modulate the feature outputs of a globally shared denoising network, enabling personalized low-count PET denoising for each institution. During the federated learning process, only the denoising network's weights are communicated and aggregated, while the FTN remains at the local institutions for feature transformation. We evaluated our method using a large-scale dataset of multi-institutional low-count PET imaging data from three medical centers located across three continents, and showed that FedFTN provides high-quality low-count PET images, outperforming previous baseline FL reconstruction methods across all low-count levels at all three institutions.
Learning Optimal Deep Projection of $^{18}$F-FDG PET Imaging for Early Differential Diagnosis of Parkinsonian Syndromes
Oct 11, 2018


Abstract:Several diseases of parkinsonian syndromes present similar symptoms at early stage and no objective widely used diagnostic methods have been approved until now. Positron emission tomography (PET) with $^{18}$F-FDG was shown to be able to assess early neuronal dysfunction of synucleinopathies and tauopathies. Tensor factorization (TF) based approaches have been applied to identify characteristic metabolic patterns for differential diagnosis. However, these conventional dimension-reduction strategies assume linear or multi-linear relationships inside data, and are therefore insufficient to distinguish nonlinear metabolic differences between various parkinsonian syndromes. In this paper, we propose a Deep Projection Neural Network (DPNN) to identify characteristic metabolic pattern for early differential diagnosis of parkinsonian syndromes. We draw our inspiration from the existing TF methods. The network consists of a (i) compression part: which uses a deep network to learn optimal 2D projections of 3D scans, and a (ii) classification part: which maps the 2D projections to labels. The compression part can be pre-trained using surplus unlabelled datasets. Also, as the classification part operates on these 2D projections, it can be trained end-to-end effectively with limited labelled data, in contrast to 3D approaches. We show that DPNN is more effective in comparison to existing state-of-the-art and plausible baselines.
* 8 pages, 3 figures, conference, MICCAI DLMIA, 2018
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge