Arkadiusz Sitek
Distilling Photon-Counting CT into Routine Chest CT through Clinically Validated Degradation Modeling
Apr 08, 2026Abstract:Photon-counting CT (PCCT) provides superior image quality with higher spatial resolution and lower noise compared to conventional energy-integrating CT (EICT), but its limited clinical availability restricts large-scale research and clinical deployment. To bridge this gap, we propose SUMI, a simulated degradation-to-enhancement method that learns to reverse realistic acquisition artifacts in low-quality EICT by leveraging high-quality PCCT as reference. Our central insight is to explicitly model realistic acquisition degradations, transforming PCCT into clinically plausible lower-quality counterparts and learning to invert this process. The simulated degradations were validated for clinical realism by board-certified radiologists, enabling faithful supervision without requiring paired acquisitions at scale. As outcomes of this technical contribution, we: (1) train a latent diffusion model on 1,046 PCCTs, using an autoencoder first pre-trained on both these PCCTs and 405,379 EICTs from 145 hospitals to extract general CT latent features that we release for reuse in other generative medical imaging tasks; (2) construct a large-scale dataset of over 17,316 publicly available EICTs enhanced to PCCT-like quality, with radiologist-validated voxel-wise annotations of airway trees, arteries, veins, lungs, and lobes; and (3) demonstrate substantial improvements: across external data, SUMI outperforms state-of-the-art image translation methods by 15% in SSIM and 20% in PSNR, improves radiologist-rated clinical utility in reader studies, and enhances downstream top-ranking lesion detection performance, increasing sensitivity by up to 15% and F1 score by up to 10%. Our results suggest that emerging imaging advances can be systematically distilled into routine EICT using limited high-quality scans as reference.
Early and Prediagnostic Detection of Pancreatic Cancer from Computed Tomography
Jan 29, 2026Abstract:Pancreatic ductal adenocarcinoma (PDAC), one of the deadliest solid malignancies, is often detected at a late and inoperable stage. Retrospective reviews of prediagnostic CT scans, when conducted by expert radiologists aware that the patient later developed PDAC, frequently reveal lesions that were previously overlooked. To help detecting these lesions earlier, we developed an automated system named ePAI (early Pancreatic cancer detection with Artificial Intelligence). It was trained on data from 1,598 patients from a single medical center. In the internal test involving 1,009 patients, ePAI achieved an area under the receiver operating characteristic curve (AUC) of 0.939-0.999, a sensitivity of 95.3%, and a specificity of 98.7% for detecting small PDAC less than 2 cm in diameter, precisely localizing PDAC as small as 2 mm. In an external test involving 7,158 patients across 6 centers, ePAI achieved an AUC of 0.918-0.945, a sensitivity of 91.5%, and a specificity of 88.0%, precisely localizing PDAC as small as 5 mm. Importantly, ePAI detected PDACs on prediagnostic CT scans obtained 3 to 36 months before clinical diagnosis that had originally been overlooked by radiologists. It successfully detected and localized PDACs in 75 of 159 patients, with a median lead time of 347 days before clinical diagnosis. Our multi-reader study showed that ePAI significantly outperformed 30 board-certified radiologists by 50.3% (P < 0.05) in sensitivity while maintaining a comparable specificity of 95.4% in detecting PDACs early and prediagnostic. These findings suggest its potential of ePAI as an assistive tool to improve early detection of pancreatic cancer.
GEPAR3D: Geometry Prior-Assisted Learning for 3D Tooth Segmentation
Jul 31, 2025Abstract:Tooth segmentation in Cone-Beam Computed Tomography (CBCT) remains challenging, especially for fine structures like root apices, which is critical for assessing root resorption in orthodontics. We introduce GEPAR3D, a novel approach that unifies instance detection and multi-class segmentation into a single step tailored to improve root segmentation. Our method integrates a Statistical Shape Model of dentition as a geometric prior, capturing anatomical context and morphological consistency without enforcing restrictive adjacency constraints. We leverage a deep watershed method, modeling each tooth as a continuous 3D energy basin encoding voxel distances to boundaries. This instance-aware representation ensures accurate segmentation of narrow, complex root apices. Trained on publicly available CBCT scans from a single center, our method is evaluated on external test sets from two in-house and two public medical centers. GEPAR3D achieves the highest overall segmentation performance, averaging a Dice Similarity Coefficient (DSC) of 95.0% (+2.8% over the second-best method) and increasing recall to 95.2% (+9.5%) across all test sets. Qualitative analyses demonstrated substantial improvements in root segmentation quality, indicating significant potential for more accurate root resorption assessment and enhanced clinical decision-making in orthodontics. We provide the implementation and dataset at https://github.com/tomek1911/GEPAR3D.
Mamba Goes HoME: Hierarchical Soft Mixture-of-Experts for 3D Medical Image Segmentation
Jul 08, 2025



Abstract:In recent years, artificial intelligence has significantly advanced medical image segmentation. However, challenges remain, including efficient 3D medical image processing across diverse modalities and handling data variability. In this work, we introduce Hierarchical Soft Mixture-of-Experts (HoME), a two-level token-routing layer for efficient long-context modeling, specifically designed for 3D medical image segmentation. Built on the Mamba state-space model (SSM) backbone, HoME enhances sequential modeling through sparse, adaptive expert routing. The first stage employs a Soft Mixture-of-Experts (SMoE) layer to partition input sequences into local groups, routing tokens to specialized per-group experts for localized feature extraction. The second stage aggregates these outputs via a global SMoE layer, enabling cross-group information fusion and global context refinement. This hierarchical design, combining local expert routing with global expert refinement improves generalizability and segmentation performance, surpassing state-of-the-art results across datasets from the three most commonly used 3D medical imaging modalities and data quality.
Foundation Model of Electronic Medical Records for Adaptive Risk Estimation
Feb 10, 2025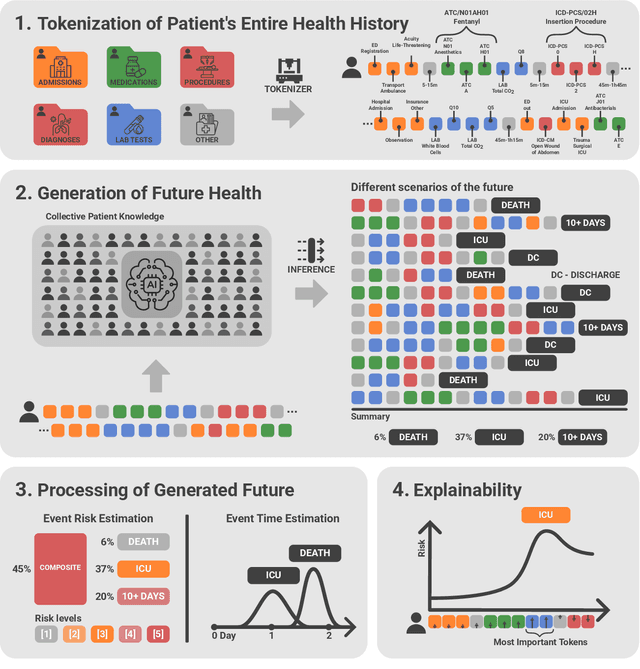
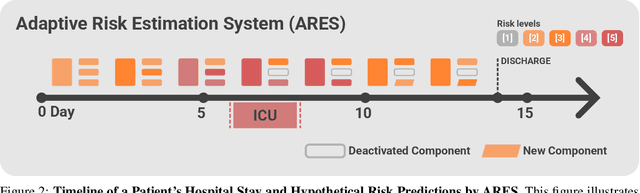
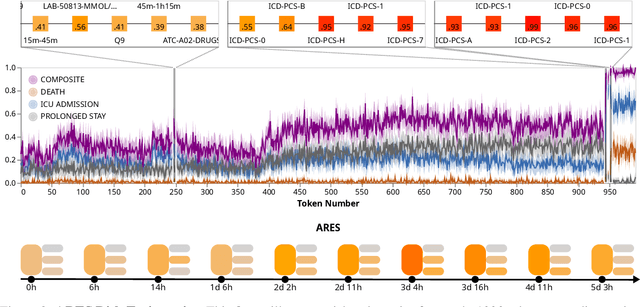
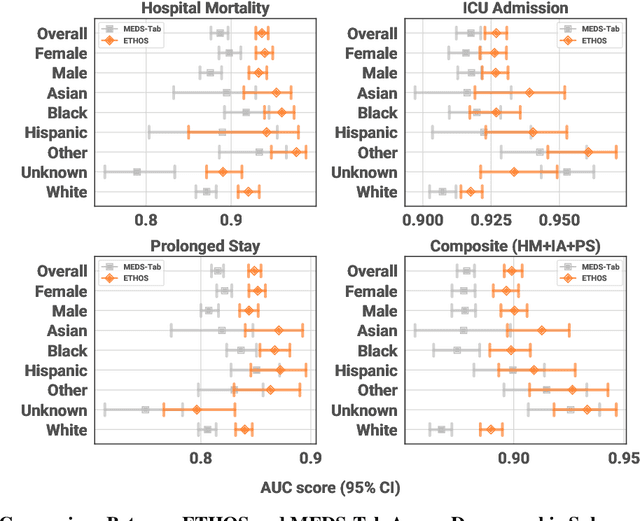
Abstract:We developed the Enhanced Transformer for Health Outcome Simulation (ETHOS), an AI model that tokenizes patient health timelines (PHTs) from EHRs. ETHOS predicts future PHTs using transformer-based architectures. The Adaptive Risk Estimation System (ARES) employs ETHOS to compute dynamic and personalized risk probabilities for clinician-defined critical events. ARES incorporates a personalized explainability module that identifies key clinical factors influencing risk estimates for individual patients. ARES was evaluated on the MIMIC-IV v2.2 dataset in emergency department (ED) settings, benchmarking its performance against traditional early warning systems and machine learning models. We processed 299,721 unique patients from MIMIC-IV into 285,622 PHTs, with 60% including hospital admissions. The dataset contained over 357 million tokens. ETHOS outperformed benchmark models in predicting hospital admissions, ICU admissions, and prolonged hospital stays, achieving superior AUC scores. ETHOS-based risk estimates demonstrated robustness across demographic subgroups with strong model reliability, confirmed via calibration curves. The personalized explainability module provides insights into patient-specific factors contributing to risk. ARES, powered by ETHOS, advances predictive healthcare AI by providing dynamic, real-time, and personalized risk estimation with patient-specific explainability to enhance clinician trust. Its adaptability and superior accuracy position it as a transformative tool for clinical decision-making, potentially improving patient outcomes and resource allocation in emergency and inpatient settings. We release the full code at github.com/ipolharvard/ethos-ares to facilitate future research.
TabMixer: Noninvasive Estimation of the Mean Pulmonary Artery Pressure via Imaging and Tabular Data Mixing
Sep 11, 2024Abstract:Right Heart Catheterization is a gold standard procedure for diagnosing Pulmonary Hypertension by measuring mean Pulmonary Artery Pressure (mPAP). It is invasive, costly, time-consuming and carries risks. In this paper, for the first time, we explore the estimation of mPAP from videos of noninvasive Cardiac Magnetic Resonance Imaging. To enhance the predictive capabilities of Deep Learning models used for this task, we introduce an additional modality in the form of demographic features and clinical measurements. Inspired by all-Multilayer Perceptron architectures, we present TabMixer, a novel module enabling the integration of imaging and tabular data through spatial, temporal and channel mixing. Specifically, we present the first approach that utilizes Multilayer Perceptrons to interchange tabular information with imaging features in vision models. We test TabMixer for mPAP estimation and show that it enhances the performance of Convolutional Neural Networks, 3D-MLP and Vision Transformers while being competitive with previous modules for imaging and tabular data. Our approach has the potential to improve clinical processes involving both modalities, particularly in noninvasive mPAP estimation, thus, significantly enhancing the quality of life for individuals affected by Pulmonary Hypertension. We provide a source code for using TabMixer at https://github.com/SanoScience/TabMixer.
Zero Shot Health Trajectory Prediction Using Transformer
Jul 30, 2024Abstract:Integrating modern machine learning and clinical decision-making has great promise for mitigating healthcare's increasing cost and complexity. We introduce the Enhanced Transformer for Health Outcome Simulation (ETHOS), a novel application of the transformer deep-learning architecture for analyzing high-dimensional, heterogeneous, and episodic health data. ETHOS is trained using Patient Health Timelines (PHTs)-detailed, tokenized records of health events-to predict future health trajectories, leveraging a zero-shot learning approach. ETHOS represents a significant advancement in foundation model development for healthcare analytics, eliminating the need for labeled data and model fine-tuning. Its ability to simulate various treatment pathways and consider patient-specific factors positions ETHOS as a tool for care optimization and addressing biases in healthcare delivery. Future developments will expand ETHOS' capabilities to incorporate a wider range of data types and data sources. Our work demonstrates a pathway toward accelerated AI development and deployment in healthcare.
Let Me DeCode You: Decoder Conditioning with Tabular Data
Jul 12, 2024



Abstract:Training deep neural networks for 3D segmentation tasks can be challenging, often requiring efficient and effective strategies to improve model performance. In this study, we introduce a novel approach, DeCode, that utilizes label-derived features for model conditioning to support the decoder in the reconstruction process dynamically, aiming to enhance the efficiency of the training process. DeCode focuses on improving 3D segmentation performance through the incorporation of conditioning embedding with learned numerical representation of 3D-label shape features. Specifically, we develop an approach, where conditioning is applied during the training phase to guide the network toward robust segmentation. When labels are not available during inference, our model infers the necessary conditioning embedding directly from the input data, thanks to a feed-forward network learned during the training phase. This approach is tested using synthetic data and cone-beam computed tomography (CBCT) images of teeth. For CBCT, three datasets are used: one publicly available and two in-house. Our results show that DeCode significantly outperforms traditional, unconditioned models in terms of generalization to unseen data, achieving higher accuracy at a reduced computational cost. This work represents the first of its kind to explore conditioning strategies in 3D data segmentation, offering a novel and more efficient method for leveraging annotated data. Our code, pre-trained models are publicly available at https://github.com/SanoScience/DeCode .
Multi-Center Fetal Brain Tissue Annotation (FeTA) Challenge 2022 Results
Feb 08, 2024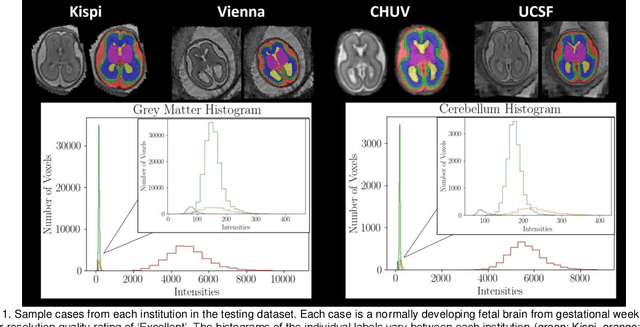
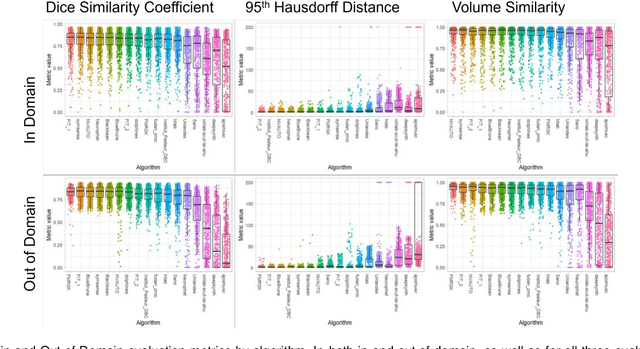
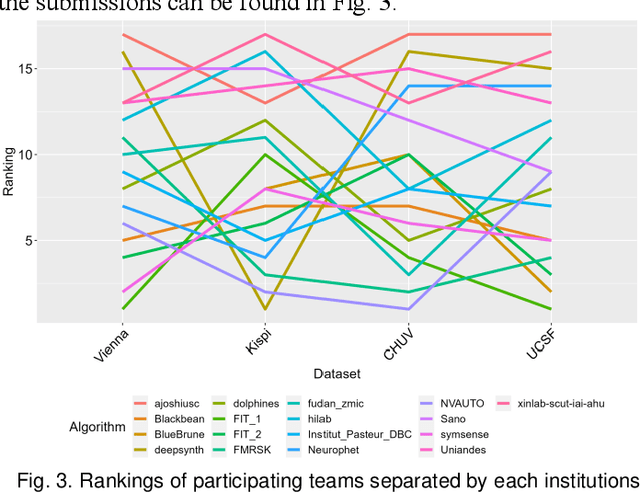
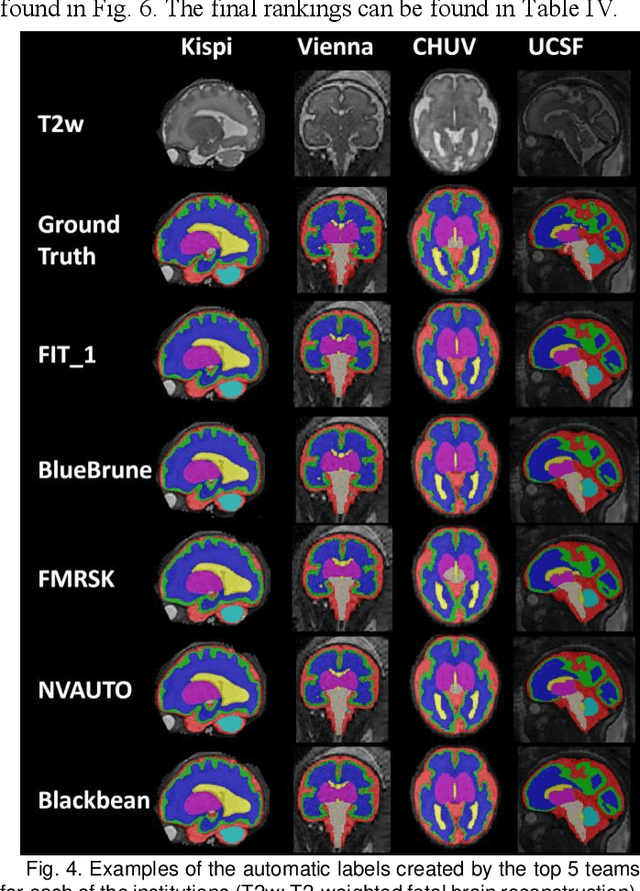
Abstract:Segmentation is a critical step in analyzing the developing human fetal brain. There have been vast improvements in automatic segmentation methods in the past several years, and the Fetal Brain Tissue Annotation (FeTA) Challenge 2021 helped to establish an excellent standard of fetal brain segmentation. However, FeTA 2021 was a single center study, and the generalizability of algorithms across different imaging centers remains unsolved, limiting real-world clinical applicability. The multi-center FeTA Challenge 2022 focuses on advancing the generalizability of fetal brain segmentation algorithms for magnetic resonance imaging (MRI). In FeTA 2022, the training dataset contained images and corresponding manually annotated multi-class labels from two imaging centers, and the testing data contained images from these two imaging centers as well as two additional unseen centers. The data from different centers varied in many aspects, including scanners used, imaging parameters, and fetal brain super-resolution algorithms applied. 16 teams participated in the challenge, and 17 algorithms were evaluated. Here, a detailed overview and analysis of the challenge results are provided, focusing on the generalizability of the submissions. Both in- and out of domain, the white matter and ventricles were segmented with the highest accuracy, while the most challenging structure remains the cerebral cortex due to anatomical complexity. The FeTA Challenge 2022 was able to successfully evaluate and advance generalizability of multi-class fetal brain tissue segmentation algorithms for MRI and it continues to benchmark new algorithms. The resulting new methods contribute to improving the analysis of brain development in utero.
MISS: Multiclass Interpretable Scoring Systems
Jan 10, 2024Abstract:In this work, we present a novel, machine-learning approach for constructing Multiclass Interpretable Scoring Systems (MISS) - a fully data-driven methodology for generating single, sparse, and user-friendly scoring systems for multiclass classification problems. Scoring systems are commonly utilized as decision support models in healthcare, criminal justice, and other domains where interpretability of predictions and ease of use are crucial. Prior methods for data-driven scoring, such as SLIM (Supersparse Linear Integer Model), were limited to binary classification tasks and extensions to multiclass domains were primarily accomplished via one-versus-all-type techniques. The scores produced by our method can be easily transformed into class probabilities via the softmax function. We demonstrate techniques for dimensionality reduction and heuristics that enhance the training efficiency and decrease the optimality gap, a measure that can certify the optimality of the model. Our approach has been extensively evaluated on datasets from various domains, and the results indicate that it is competitive with other machine learning models in terms of classification performance metrics and provides well-calibrated class probabilities.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge