Yihui Wang
MambaMIL+: Modeling Long-Term Contextual Patterns for Gigapixel Whole Slide Image
Dec 19, 2025
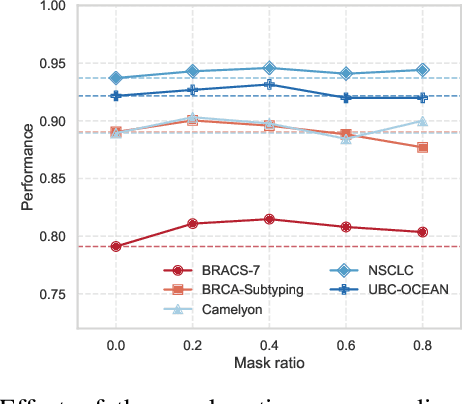
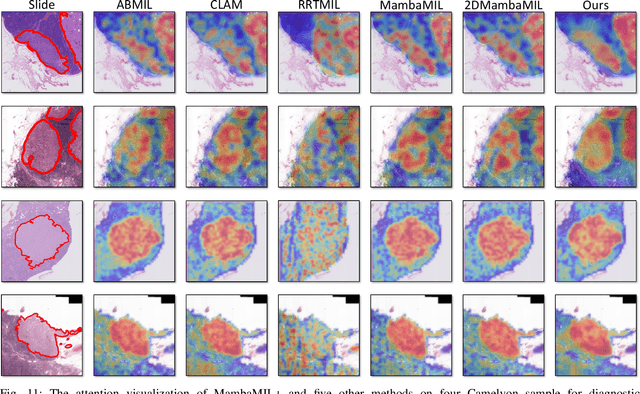
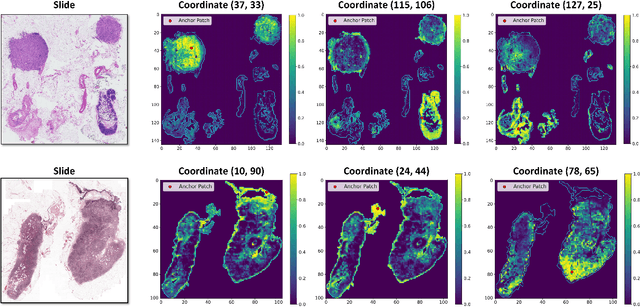
Abstract:Whole-slide images (WSIs) are an important data modality in computational pathology, yet their gigapixel resolution and lack of fine-grained annotations challenge conventional deep learning models. Multiple instance learning (MIL) offers a solution by treating each WSI as a bag of patch-level instances, but effectively modeling ultra-long sequences with rich spatial context remains difficult. Recently, Mamba has emerged as a promising alternative for long sequence learning, scaling linearly to thousands of tokens. However, despite its efficiency, it still suffers from limited spatial context modeling and memory decay, constraining its effectiveness to WSI analysis. To address these limitations, we propose MambaMIL+, a new MIL framework that explicitly integrates spatial context while maintaining long-range dependency modeling without memory forgetting. Specifically, MambaMIL+ introduces 1) overlapping scanning, which restructures the patch sequence to embed spatial continuity and instance correlations; 2) a selective stripe position encoder (S2PE) that encodes positional information while mitigating the biases of fixed scanning orders; and 3) a contextual token selection (CTS) mechanism, which leverages supervisory knowledge to dynamically enlarge the contextual memory for stable long-range modeling. Extensive experiments on 20 benchmarks across diagnostic classification, molecular prediction, and survival analysis demonstrate that MambaMIL+ consistently achieves state-of-the-art performance under three feature extractors (ResNet-50, PLIP, and CONCH), highlighting its effectiveness and robustness for large-scale computational pathology
LLM-driven Knowledge Enhancement for Multimodal Cancer Survival Prediction
Dec 16, 2025Abstract:Current multimodal survival prediction methods typically rely on pathology images (WSIs) and genomic data, both of which are high-dimensional and redundant, making it difficult to extract discriminative features from them and align different modalities. Moreover, using a simple survival follow-up label is insufficient to supervise such a complex task. To address these challenges, we propose KEMM, an LLM-driven Knowledge-Enhanced Multimodal Model for cancer survival prediction, which integrates expert reports and prognostic background knowledge. 1) Expert reports, provided by pathologists on a case-by-case basis and refined by large language model (LLM), offer succinct and clinically focused diagnostic statements. This information may typically suggest different survival outcomes. 2) Prognostic background knowledge (PBK), generated concisely by LLM, provides valuable prognostic background knowledge on different cancer types, which also enhances survival prediction. To leverage these knowledge, we introduce the knowledge-enhanced cross-modal (KECM) attention module. KECM can effectively guide the network to focus on discriminative and survival-relevant features from highly redundant modalities. Extensive experiments on five datasets demonstrate that KEMM achieves state-of-the-art performance. The code will be released upon acceptance.
A Clinical-grade Universal Foundation Model for Intraoperative Pathology
Oct 06, 2025Abstract:Intraoperative pathology is pivotal to precision surgery, yet its clinical impact is constrained by diagnostic complexity and the limited availability of high-quality frozen-section data. While computational pathology has made significant strides, the lack of large-scale, prospective validation has impeded its routine adoption in surgical workflows. Here, we introduce CRISP, a clinical-grade foundation model developed on over 100,000 frozen sections from eight medical centers, specifically designed to provide Clinical-grade Robust Intraoperative Support for Pathology (CRISP). CRISP was comprehensively evaluated on more than 15,000 intraoperative slides across nearly 100 retrospective diagnostic tasks, including benign-malignant discrimination, key intraoperative decision-making, and pan-cancer detection, etc. The model demonstrated robust generalization across diverse institutions, tumor types, and anatomical sites-including previously unseen sites and rare cancers. In a prospective cohort of over 2,000 patients, CRISP sustained high diagnostic accuracy under real-world conditions, directly informing surgical decisions in 92.6% of cases. Human-AI collaboration further reduced diagnostic workload by 35%, avoided 105 ancillary tests and enhanced detection of micrometastases with 87.5% accuracy. Together, these findings position CRISP as a clinical-grade paradigm for AI-driven intraoperative pathology, bridging computational advances with surgical precision and accelerating the translation of artificial intelligence into routine clinical practice.
GenAR: Next-Scale Autoregressive Generation for Spatial Gene Expression Prediction
Oct 05, 2025



Abstract:Spatial Transcriptomics (ST) offers spatially resolved gene expression but remains costly. Predicting expression directly from widely available Hematoxylin and Eosin (H&E) stained images presents a cost-effective alternative. However, most computational approaches (i) predict each gene independently, overlooking co-expression structure, and (ii) cast the task as continuous regression despite expression being discrete counts. This mismatch can yield biologically implausible outputs and complicate downstream analyses. We introduce GenAR, a multi-scale autoregressive framework that refines predictions from coarse to fine. GenAR clusters genes into hierarchical groups to expose cross-gene dependencies, models expression as codebook-free discrete token generation to directly predict raw counts, and conditions decoding on fused histological and spatial embeddings. From an information-theoretic perspective, the discrete formulation avoids log-induced biases and the coarse-to-fine factorization aligns with a principled conditional decomposition. Extensive experimental results on four Spatial Transcriptomics datasets across different tissue types demonstrate that GenAR achieves state-of-the-art performance, offering potential implications for precision medicine and cost-effective molecular profiling. Code is publicly available at https://github.com/oyjr/genar.
A Versatile Pathology Co-pilot via Reasoning Enhanced Multimodal Large Language Model
Jul 23, 2025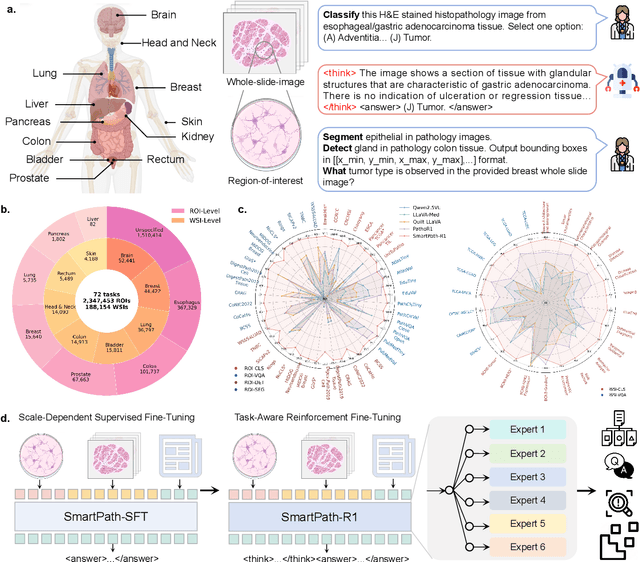
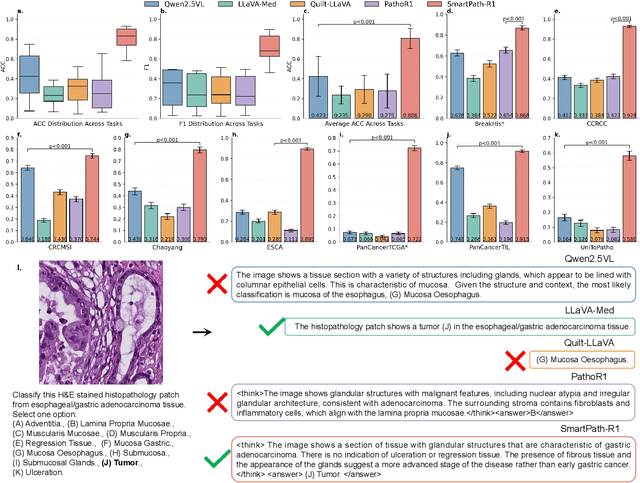
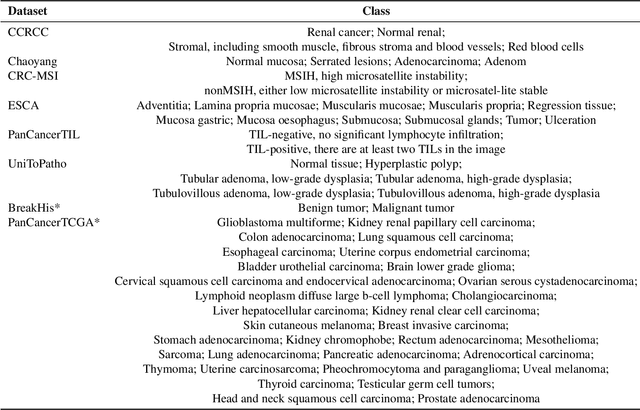
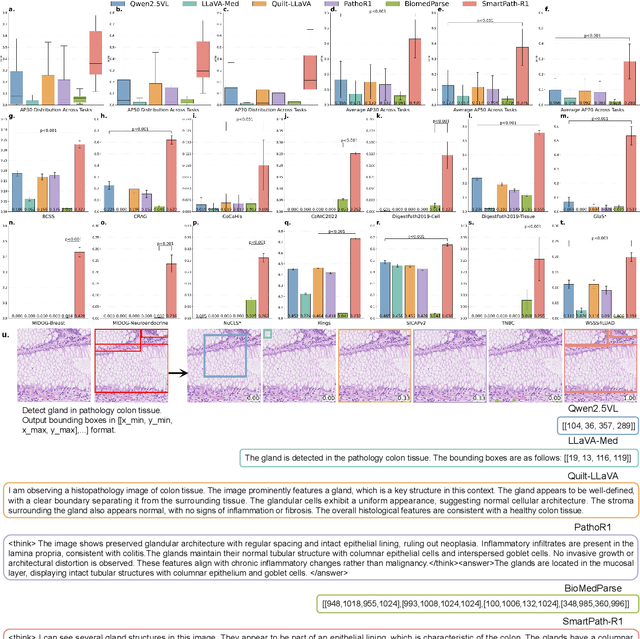
Abstract:Multimodal large language models (MLLMs) have emerged as powerful tools for computational pathology, offering unprecedented opportunities to integrate pathological images with language context for comprehensive diagnostic analysis. These models hold particular promise for automating complex tasks that traditionally require expert interpretation of pathologists. However, current MLLM approaches in pathology demonstrate significantly constrained reasoning capabilities, primarily due to their reliance on expensive chain-of-thought annotations. Additionally, existing methods remain limited to simplex application of visual question answering (VQA) at region-of-interest (ROI) level, failing to address the full spectrum of diagnostic needs such as ROI classification, detection, segmentation, whole-slide-image (WSI) classification and VQA in clinical practice. In this study, we present SmartPath-R1, a versatile MLLM capable of simultaneously addressing both ROI-level and WSI-level tasks while demonstrating robust pathological reasoning capability. Our framework combines scale-dependent supervised fine-tuning and task-aware reinforcement fine-tuning, which circumvents the requirement for chain-of-thought supervision by leveraging the intrinsic knowledge within MLLM. Furthermore, SmartPath-R1 integrates multiscale and multitask analysis through a mixture-of-experts mechanism, enabling dynamic processing for diverse tasks. We curate a large-scale dataset comprising 2.3M ROI samples and 188K WSI samples for training and evaluation. Extensive experiments across 72 tasks validate the effectiveness and superiority of the proposed approach. This work represents a significant step toward developing versatile, reasoning-enhanced AI systems for precision pathology.
Comparative validation of surgical phase recognition, instrument keypoint estimation, and instrument instance segmentation in endoscopy: Results of the PhaKIR 2024 challenge
Jul 22, 2025Abstract:Reliable recognition and localization of surgical instruments in endoscopic video recordings are foundational for a wide range of applications in computer- and robot-assisted minimally invasive surgery (RAMIS), including surgical training, skill assessment, and autonomous assistance. However, robust performance under real-world conditions remains a significant challenge. Incorporating surgical context - such as the current procedural phase - has emerged as a promising strategy to improve robustness and interpretability. To address these challenges, we organized the Surgical Procedure Phase, Keypoint, and Instrument Recognition (PhaKIR) sub-challenge as part of the Endoscopic Vision (EndoVis) challenge at MICCAI 2024. We introduced a novel, multi-center dataset comprising thirteen full-length laparoscopic cholecystectomy videos collected from three distinct medical institutions, with unified annotations for three interrelated tasks: surgical phase recognition, instrument keypoint estimation, and instrument instance segmentation. Unlike existing datasets, ours enables joint investigation of instrument localization and procedural context within the same data while supporting the integration of temporal information across entire procedures. We report results and findings in accordance with the BIAS guidelines for biomedical image analysis challenges. The PhaKIR sub-challenge advances the field by providing a unique benchmark for developing temporally aware, context-driven methods in RAMIS and offers a high-quality resource to support future research in surgical scene understanding.
Segment Anything in Pathology Images with Natural Language
Jun 26, 2025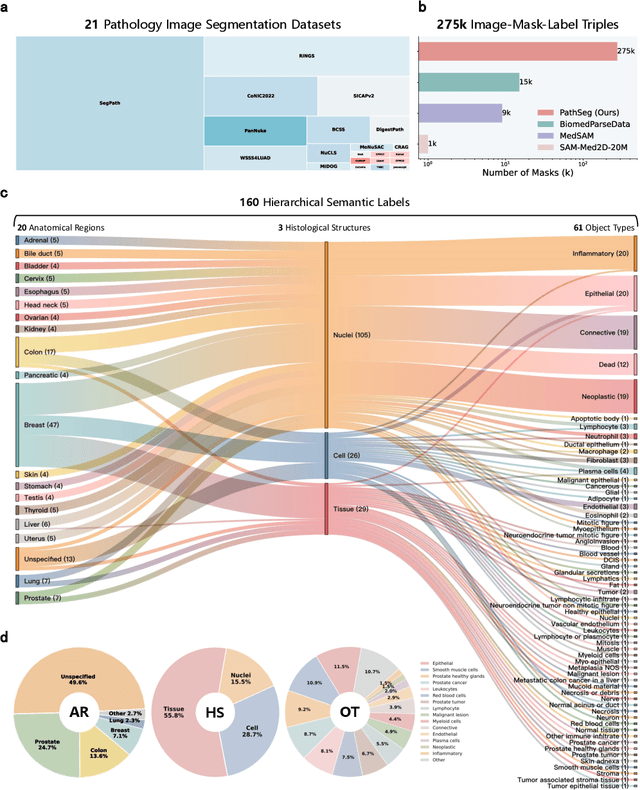
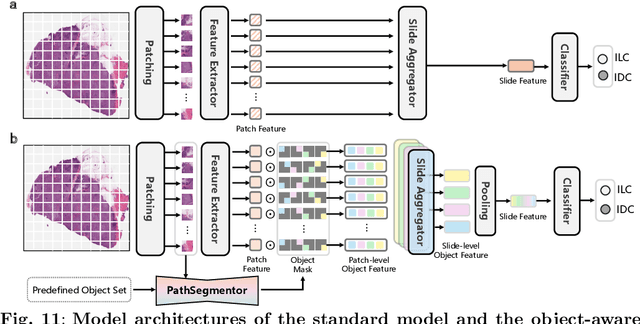
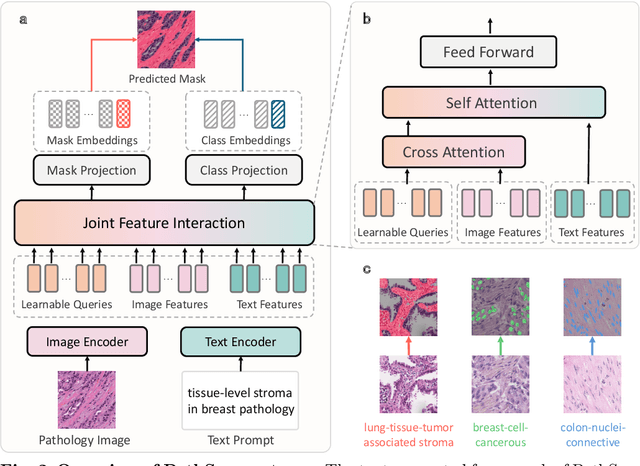
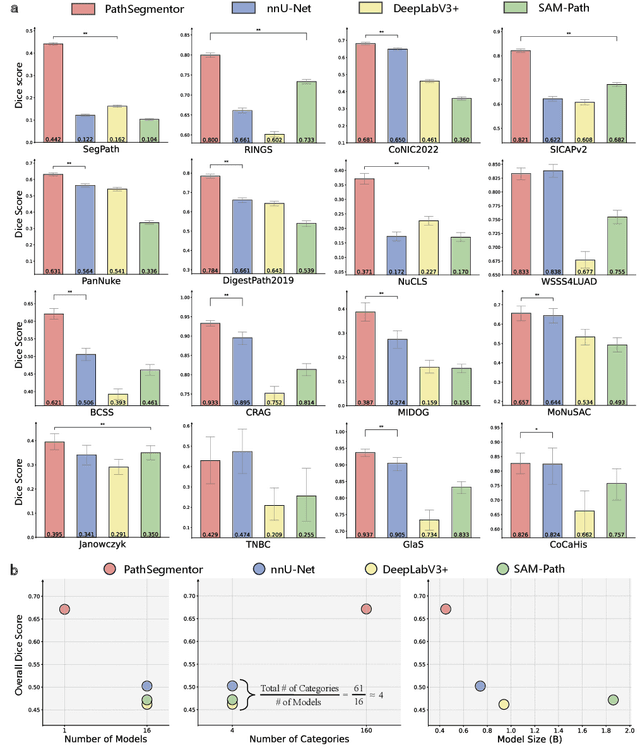
Abstract:Pathology image segmentation is crucial in computational pathology for analyzing histological features relevant to cancer diagnosis and prognosis. However, current methods face major challenges in clinical applications due to limited annotated data and restricted category definitions. To address these limitations, we propose PathSegmentor, the first text-prompted segmentation foundation model designed specifically for pathology images. We also introduce PathSeg , the largest and most comprehensive dataset for pathology segmentation, built from 17 public sources and containing 275k image-mask-label triples across 160 diverse categories. With PathSegmentor, users can perform semantic segmentation using natural language prompts, eliminating the need for laborious spatial inputs such as points or boxes. Extensive experiments demonstrate that PathSegmentor outperforms specialized models with higher accuracy and broader applicability, while maintaining a compact architecture. It significantly surpasses existing spatial- and text-prompted models by 0.145 and 0.429 in overall Dice scores, respectively, showing strong robustness in segmenting complex structures and generalizing to external datasets. Moreover, PathSegmentor's outputs enhance the interpretability of diagnostic models through feature importance estimation and imaging biomarker discovery, offering pathologists evidence-based support for clinical decision-making. This work advances the development of explainable AI in precision oncology.
Genome-Anchored Foundation Model Embeddings Improve Molecular Prediction from Histology Images
Jun 24, 2025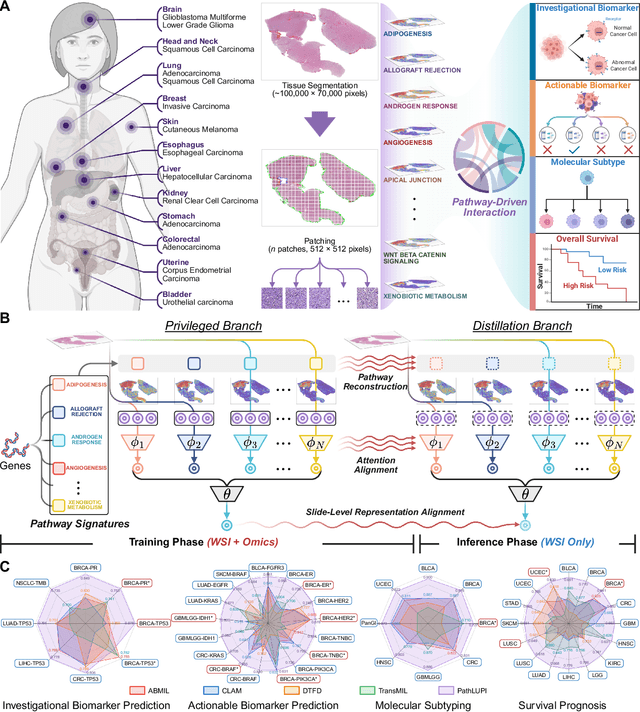
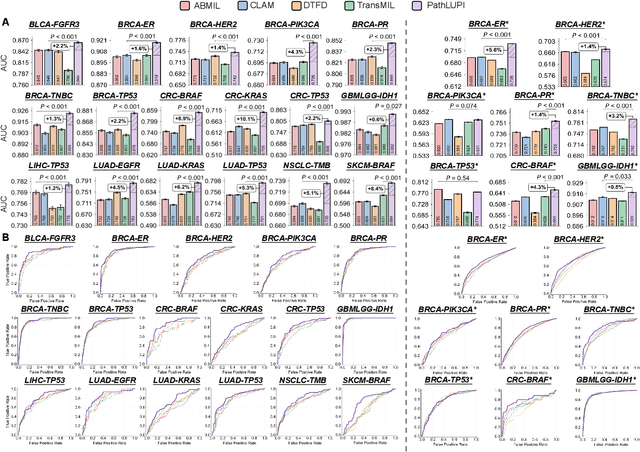
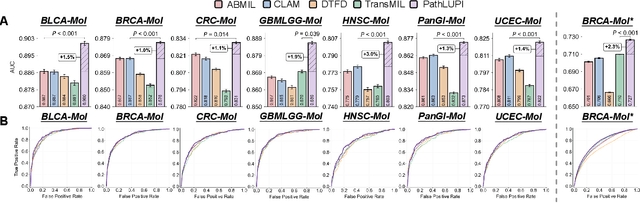
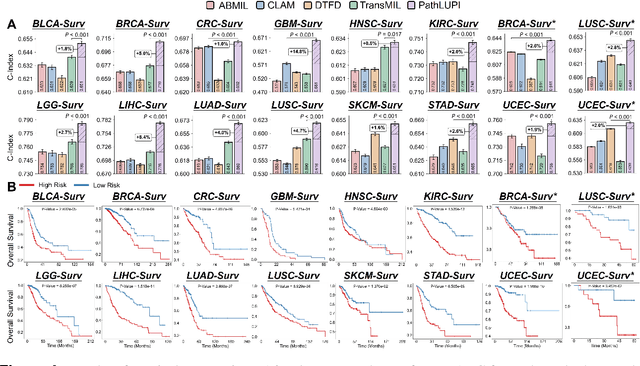
Abstract:Precision oncology requires accurate molecular insights, yet obtaining these directly from genomics is costly and time-consuming for broad clinical use. Predicting complex molecular features and patient prognosis directly from routine whole-slide images (WSI) remains a major challenge for current deep learning methods. Here we introduce PathLUPI, which uses transcriptomic privileged information during training to extract genome-anchored histological embeddings, enabling effective molecular prediction using only WSIs at inference. Through extensive evaluation across 49 molecular oncology tasks using 11,257 cases among 20 cohorts, PathLUPI demonstrated superior performance compared to conventional methods trained solely on WSIs. Crucially, it achieves AUC $\geq$ 0.80 in 14 of the biomarker prediction and molecular subtyping tasks and C-index $\geq$ 0.70 in survival cohorts of 5 major cancer types. Moreover, PathLUPI embeddings reveal distinct cellular morphological signatures associated with specific genotypes and related biological pathways within WSIs. By effectively encoding molecular context to refine WSI representations, PathLUPI overcomes a key limitation of existing models and offers a novel strategy to bridge molecular insights with routine pathology workflows for wider clinical application.
PathBench: A comprehensive comparison benchmark for pathology foundation models towards precision oncology
May 26, 2025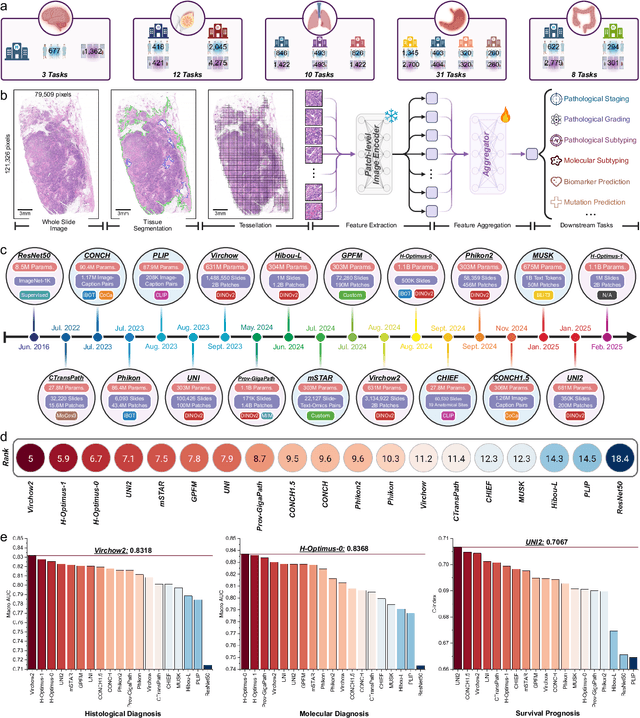



Abstract:The emergence of pathology foundation models has revolutionized computational histopathology, enabling highly accurate, generalized whole-slide image analysis for improved cancer diagnosis, and prognosis assessment. While these models show remarkable potential across cancer diagnostics and prognostics, their clinical translation faces critical challenges including variability in optimal model across cancer types, potential data leakage in evaluation, and lack of standardized benchmarks. Without rigorous, unbiased evaluation, even the most advanced PFMs risk remaining confined to research settings, delaying their life-saving applications. Existing benchmarking efforts remain limited by narrow cancer-type focus, potential pretraining data overlaps, or incomplete task coverage. We present PathBench, the first comprehensive benchmark addressing these gaps through: multi-center in-hourse datasets spanning common cancers with rigorous leakage prevention, evaluation across the full clinical spectrum from diagnosis to prognosis, and an automated leaderboard system for continuous model assessment. Our framework incorporates large-scale data, enabling objective comparison of PFMs while reflecting real-world clinical complexity. All evaluation data comes from private medical providers, with strict exclusion of any pretraining usage to avoid data leakage risks. We have collected 15,888 WSIs from 8,549 patients across 10 hospitals, encompassing over 64 diagnosis and prognosis tasks. Currently, our evaluation of 19 PFMs shows that Virchow2 and H-Optimus-1 are the most effective models overall. This work provides researchers with a robust platform for model development and offers clinicians actionable insights into PFM performance across diverse clinical scenarios, ultimately accelerating the translation of these transformative technologies into routine pathology practice.
Discovering Pathology Rationale and Token Allocation for Efficient Multimodal Pathology Reasoning
May 21, 2025Abstract:Multimodal pathological image understanding has garnered widespread interest due to its potential to improve diagnostic accuracy and enable personalized treatment through integrated visual and textual data. However, existing methods exhibit limited reasoning capabilities, which hamper their ability to handle complex diagnostic scenarios. Additionally, the enormous size of pathological images leads to severe computational burdens, further restricting their practical deployment. To address these limitations, we introduce a novel bilateral reinforcement learning framework comprising two synergistic branches. One reinforcement branch enhances the reasoning capability by enabling the model to learn task-specific decision processes, i.e., pathology rationales, directly from labels without explicit reasoning supervision. While the other branch dynamically allocates a tailored number of tokens to different images based on both their visual content and task context, thereby optimizing computational efficiency. We apply our method to various pathological tasks such as visual question answering, cancer subtyping, and lesion detection. Extensive experiments show an average +41.7 absolute performance improvement with 70.3% lower inference costs over the base models, achieving both reasoning accuracy and computational efficiency.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge