Simon A. A. Kohl
A Case for the Score: Identifying Image Anomalies using Variational Autoencoder Gradients
Nov 28, 2019
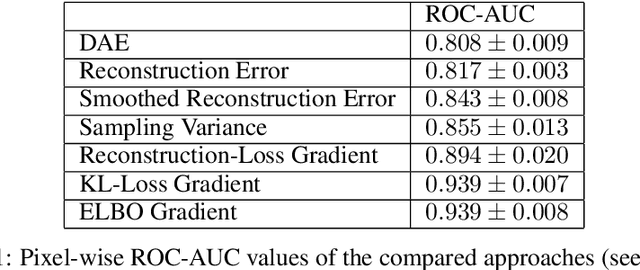


Abstract:Through training on unlabeled data, anomaly detection has the potential to impact computer-aided diagnosis by outlining suspicious regions. Previous work on deep-learning-based anomaly detection has primarily focused on the reconstruction error. We argue instead, that pixel-wise anomaly ratings derived from a Variational Autoencoder based score approximation yield a theoretically better grounded and more faithful estimate. In our experiments, Variational Autoencoder gradient-based rating outperforms other approaches on unsupervised pixel-wise tumor detection on the BraTS-2017 dataset with a ROC-AUC of 0.94.
Reg R-CNN: Lesion Detection and Grading under Noisy Labels
Aug 26, 2019



Abstract:For the task of concurrently detecting and categorizing objects, the medical imaging community commonly adopts methods developed on natural images. Current state-of-the-art object detectors are comprised of two stages: the first stage generates region proposals, the second stage subsequently categorizes them. Unlike in natural images, however, for anatomical structures of interest such as tumors, the appearance in the image (e.g., scale or intensity) links to a malignancy grade that lies on a continuous ordinal scale. While classification models discard this ordinal relation between grades by discretizing the continuous scale to an unordered bag of categories, regression models are trained with distance metrics, which preserve the relation. This advantage becomes all the more important in the setting of label confusions on ambiguous data sets, which is the usual case with medical images. To this end, we propose Reg R-CNN, which replaces the second-stage classification model of a current object detector with a regression model. We show the superiority of our approach on a public data set with 1026 patients and a series of toy experiments. Code will be available at github.com/MIC-DKFZ/RegRCNN.
Deep Probabilistic Modeling of Glioma Growth
Jul 09, 2019
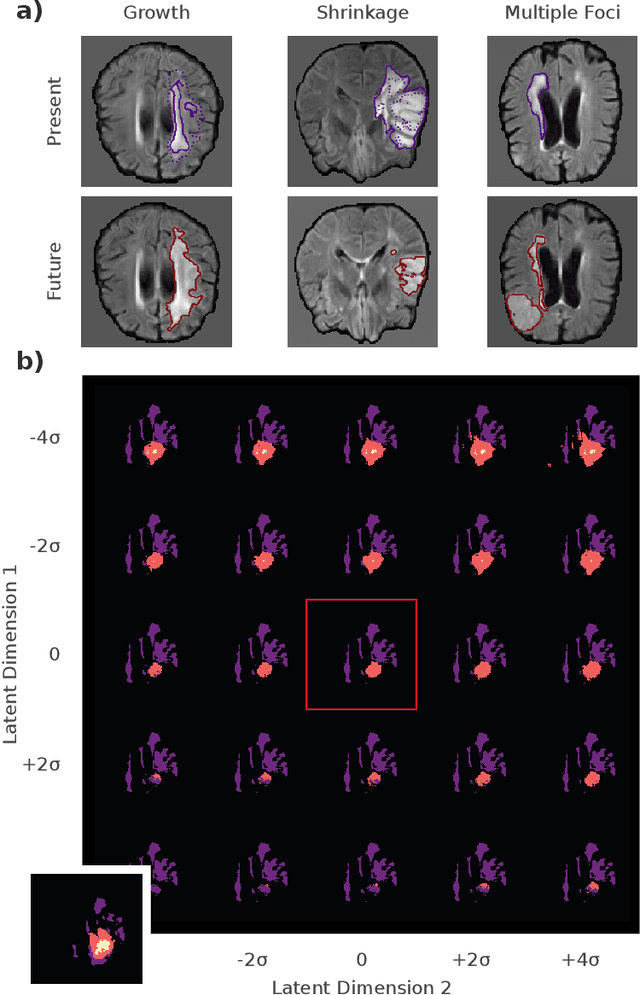
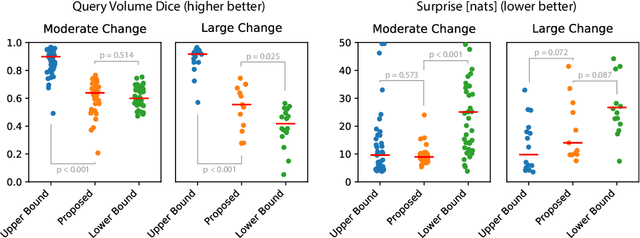
Abstract:Existing approaches to modeling the dynamics of brain tumor growth, specifically glioma, employ biologically inspired models of cell diffusion, using image data to estimate the associated parameters. In this work, we propose an alternative approach based on recent advances in probabilistic segmentation and representation learning that implicitly learns growth dynamics directly from data without an underlying explicit model. We present evidence that our approach is able to learn a distribution of plausible future tumor appearances conditioned on past observations of the same tumor.
A Hierarchical Probabilistic U-Net for Modeling Multi-Scale Ambiguities
May 30, 2019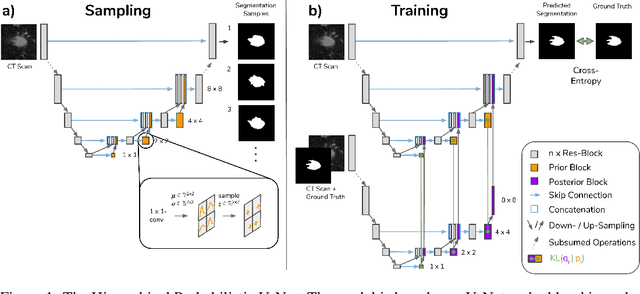
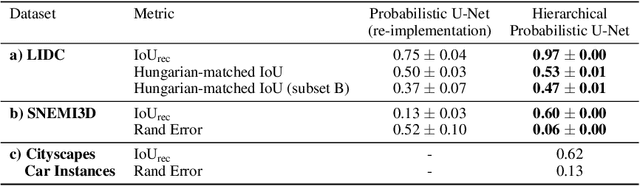
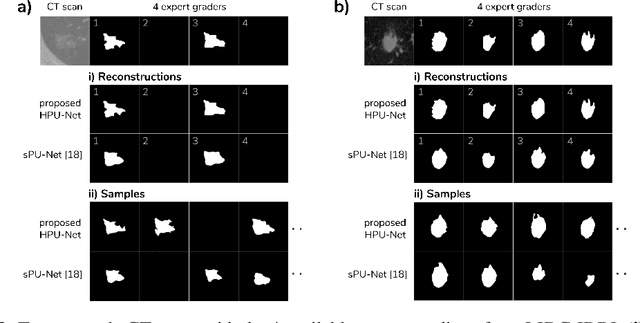
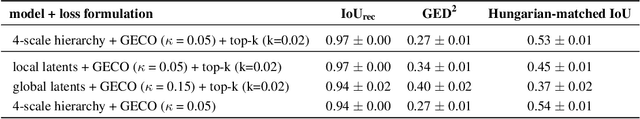
Abstract:Medical imaging only indirectly measures the molecular identity of the tissue within each voxel, which often produces only ambiguous image evidence for target measures of interest, like semantic segmentation. This diversity and the variations of plausible interpretations are often specific to given image regions and may thus manifest on various scales, spanning all the way from the pixel to the image level. In order to learn a flexible distribution that can account for multiple scales of variations, we propose the Hierarchical Probabilistic U-Net, a segmentation network with a conditional variational auto-encoder (cVAE) that uses a hierarchical latent space decomposition. We show that this model formulation enables sampling and reconstruction of segmenations with high fidelity, i.e. with finely resolved detail, while providing the flexibility to learn complex structured distributions across scales. We demonstrate these abilities on the task of segmenting ambiguous medical scans as well as on instance segmentation of neurobiological and natural images. Our model automatically separates independent factors across scales, an inductive bias that we deem beneficial in structured output prediction tasks beyond segmentation.
nnU-Net: Breaking the Spell on Successful Medical Image Segmentation
Apr 17, 2019
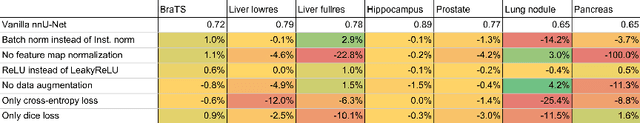
Abstract:Fueled by the diversity of datasets, semantic segmentation is a popular subfield in medical image analysis with a vast number of new methods being proposed each year. This ever-growing jungle of methodologies, however, becomes increasingly impenetrable. At the same time, many proposed methods fail to generalize beyond the experiments they were demonstrated on, thus hampering the process of developing a segmentation algorithm on a new dataset. Here we present nnU-Net ('no-new-Net'), a framework that automatically adapts itself to any given new dataset. While this process was completely human-driven so far, we make a first attempt to automate necessary adaptations such as preprocessing, the exact patch size, batch size, and inference settings based on the properties of a given dataset. Remarkably, nnU-Net strips away the architectural bells and whistles that are typically proposed in the literature and relies on just a simple U-Net architecture embedded in a robust training scheme. Out of the box, nnU-Net achieves state of the art performance on six well-established segmentation challenges. Source code is available at https://github.com/MIC-DKFZ/nnunet.
Context-encoding Variational Autoencoder for Unsupervised Anomaly Detection
Dec 14, 2018
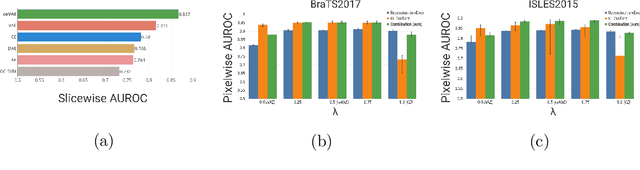
Abstract:Unsupervised learning can leverage large-scale data sources without the need for annotations. In this context, deep learning-based auto encoders have shown great potential in detecting anomalies in medical images. However, state-of-the-art anomaly scores are still based on the reconstruction error, which lacks in two essential parts: it ignores the model-internal representation employed for reconstruction, and it lacks formal assertions and comparability between samples. We address these shortcomings by proposing the Context-encoding Variational Autoencoder (ceVAE) which combines reconstruction- with density-based anomaly scoring. This improves the sample- as well as pixel-wise results. In our experiments on the BraTS-2017 and ISLES-2015 segmentation benchmarks, the ceVAE achieves unsupervised ROC-AUCs of 0.95 and 0.89, respectively, thus outperforming state-of-the-art methods by a considerable margin.
Retina U-Net: Embarrassingly Simple Exploitation of Segmentation Supervision for Medical Object Detection
Nov 21, 2018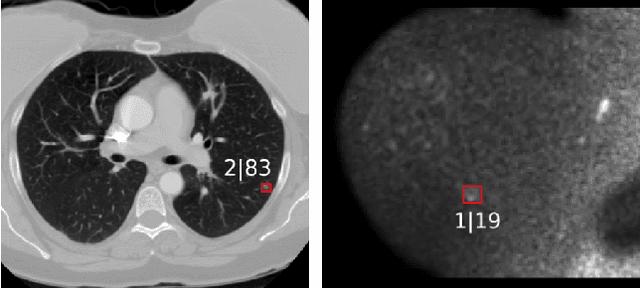
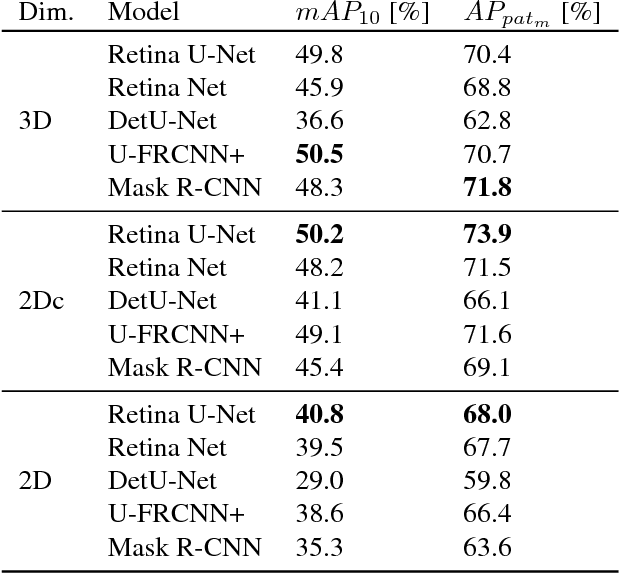
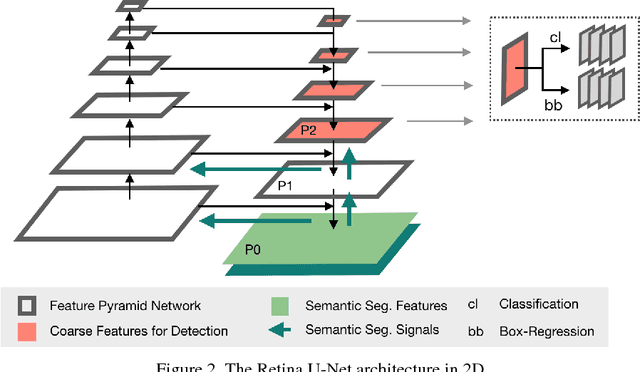
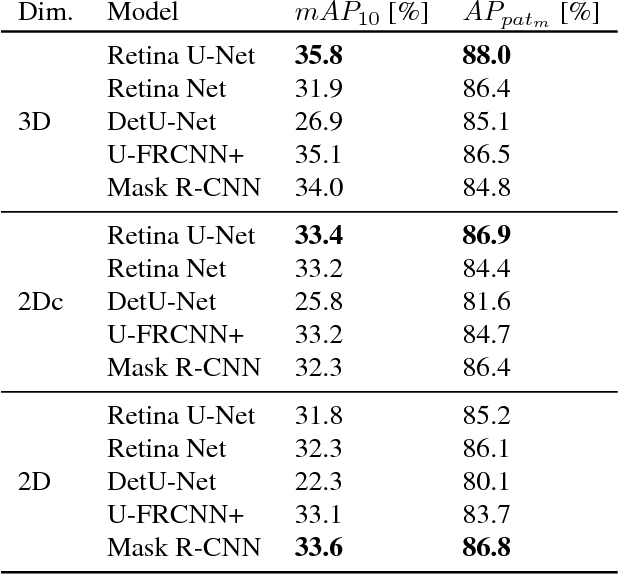
Abstract:The task of localizing and categorizing objects in medical images often remains formulated as a semantic segmentation problem. This approach, however, only indirectly solves the coarse localization task by predicting pixel-level scores, requiring ad-hoc heuristics when mapping back to object-level scores. State-of-the-art object detectors on the other hand, allow for individual object scoring in an end-to-end fashion, while ironically trading in the ability to exploit the full pixel-wise supervision signal. This can be particularly disadvantageous in the setting of medical image analysis, where data sets are notoriously small. In this paper, we propose Retina U-Net, a simple architecture, which naturally fuses the Retina Net one-stage detector with the U-Net architecture widely used for semantic segmentation in medical images. The proposed architecture recaptures discarded supervision signals by complementing object detection with an auxiliary task in the form of semantic segmentation without introducing the additional complexity of previously proposed two-stage detectors. We evaluate the importance of full segmentation supervision on two medical data sets, provide an in-depth analysis on a series of toy experiments and show how the corresponding performance gain grows in the limit of small data sets. Retina U-Net yields strong detection performance only reached by its more complex two-staged counterparts. Our framework including all methods implemented for operation on 2D and 3D images is available at github.com/pfjaeger/medicaldetectiontoolkit.
A Probabilistic U-Net for Segmentation of Ambiguous Images
Oct 29, 2018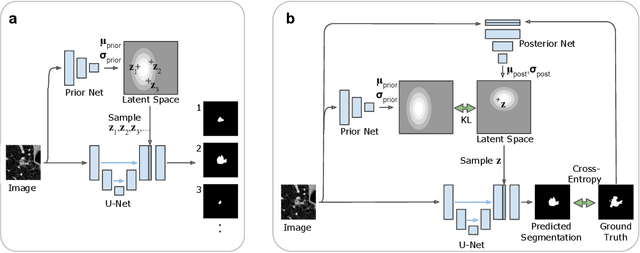

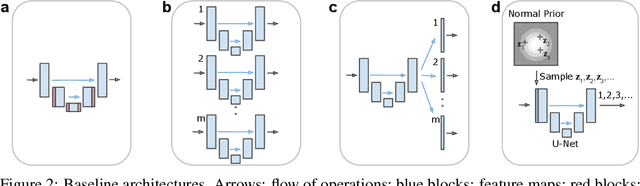

Abstract:Many real-world vision problems suffer from inherent ambiguities. In clinical applications for example, it might not be clear from a CT scan alone which particular region is cancer tissue. Therefore a group of graders typically produces a set of diverse but plausible segmentations. We consider the task of learning a distribution over segmentations given an input. To this end we propose a generative segmentation model based on a combination of a U-Net with a conditional variational autoencoder that is capable of efficiently producing an unlimited number of plausible hypotheses. We show on a lung abnormalities segmentation task and on a Cityscapes segmentation task that our model reproduces the possible segmentation variants as well as the frequencies with which they occur, doing so significantly better than published approaches. These models could have a high impact in real-world applications, such as being used as clinical decision-making algorithms accounting for multiple plausible semantic segmentation hypotheses to provide possible diagnoses and recommend further actions to resolve the present ambiguities.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge