Muhammad Ridzuan
Efficient Parameter Adaptation for Multi-Modal Medical Image Segmentation and Prognosis
Apr 18, 2025
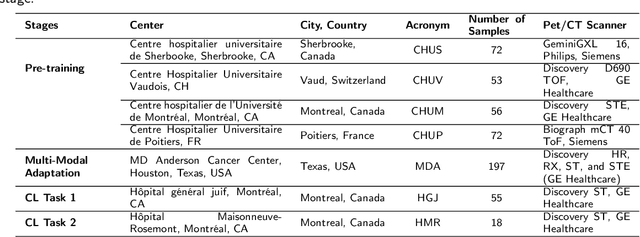
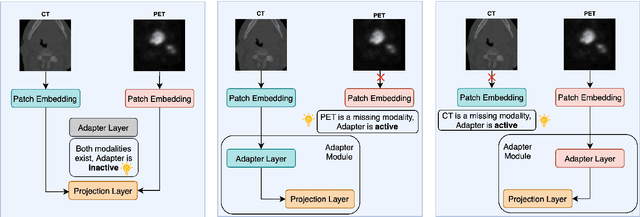
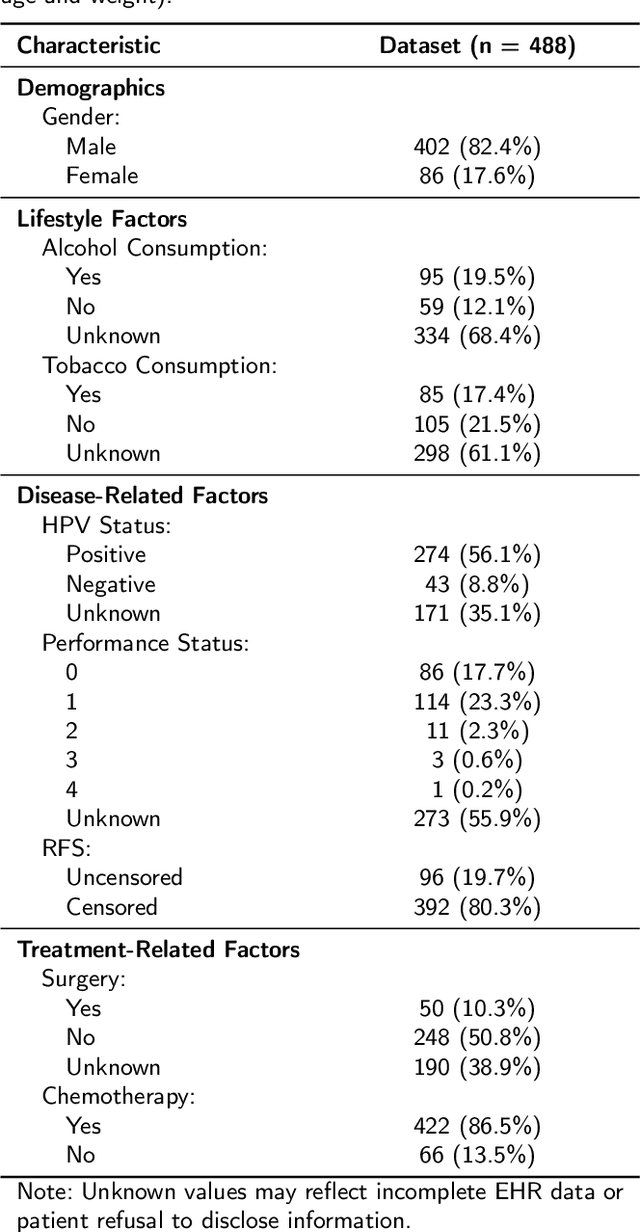
Abstract:Cancer detection and prognosis relies heavily on medical imaging, particularly CT and PET scans. Deep Neural Networks (DNNs) have shown promise in tumor segmentation by fusing information from these modalities. However, a critical bottleneck exists: the dependency on CT-PET data concurrently for training and inference, posing a challenge due to the limited availability of PET scans. Hence, there is a clear need for a flexible and efficient framework that can be trained with the widely available CT scans and can be still adapted for PET scans when they become available. In this work, we propose a parameter-efficient multi-modal adaptation (PEMMA) framework for lightweight upgrading of a transformer-based segmentation model trained only on CT scans such that it can be efficiently adapted for use with PET scans when they become available. This framework is further extended to perform prognosis task maintaining the same efficient cross-modal fine-tuning approach. The proposed approach is tested with two well-known segementation backbones, namely UNETR and Swin UNETR. Our approach offers two main advantages. Firstly, we leverage the inherent modularity of the transformer architecture and perform low-rank adaptation (LoRA) as well as decomposed low-rank adaptation (DoRA) of the attention weights to achieve parameter-efficient adaptation. Secondly, by minimizing cross-modal entanglement, PEMMA allows updates using only one modality without causing catastrophic forgetting in the other. Our method achieves comparable performance to early fusion, but with only 8% of the trainable parameters, and demonstrates a significant +28% Dice score improvement on PET scans when trained with a single modality. Furthermore, in prognosis, our method improves the concordance index by +10% when adapting a CT-pretrained model to include PET scans, and by +23% when adapting for both PET and EHR data.
All Languages Matter: Evaluating LMMs on Culturally Diverse 100 Languages
Nov 25, 2024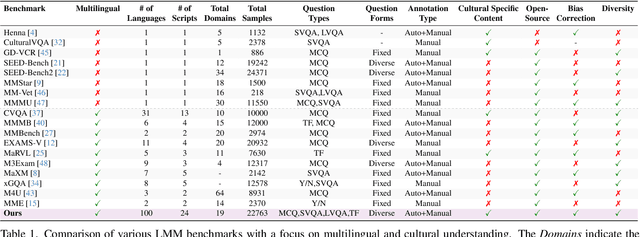
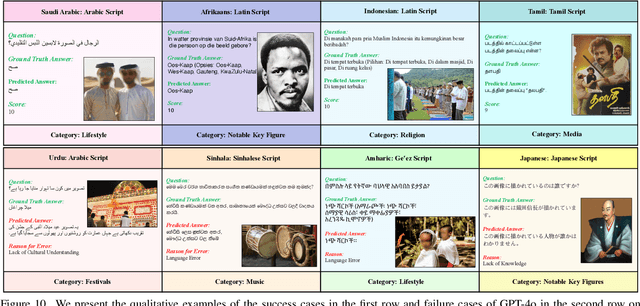
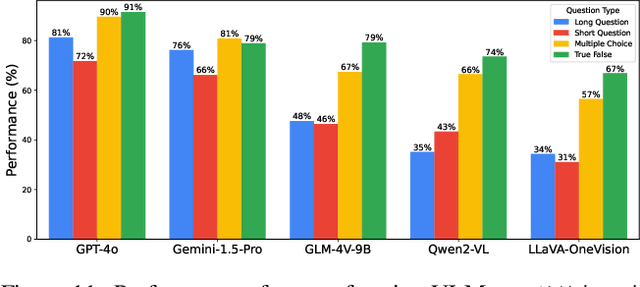
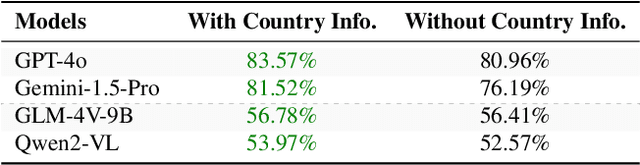
Abstract:Existing Large Multimodal Models (LMMs) generally focus on only a few regions and languages. As LMMs continue to improve, it is increasingly important to ensure they understand cultural contexts, respect local sensitivities, and support low-resource languages, all while effectively integrating corresponding visual cues. In pursuit of culturally diverse global multimodal models, our proposed All Languages Matter Benchmark (ALM-bench) represents the largest and most comprehensive effort to date for evaluating LMMs across 100 languages. ALM-bench challenges existing models by testing their ability to understand and reason about culturally diverse images paired with text in various languages, including many low-resource languages traditionally underrepresented in LMM research. The benchmark offers a robust and nuanced evaluation framework featuring various question formats, including true/false, multiple choice, and open-ended questions, which are further divided into short and long-answer categories. ALM-bench design ensures a comprehensive assessment of a model's ability to handle varied levels of difficulty in visual and linguistic reasoning. To capture the rich tapestry of global cultures, ALM-bench carefully curates content from 13 distinct cultural aspects, ranging from traditions and rituals to famous personalities and celebrations. Through this, ALM-bench not only provides a rigorous testing ground for state-of-the-art open and closed-source LMMs but also highlights the importance of cultural and linguistic inclusivity, encouraging the development of models that can serve diverse global populations effectively. Our benchmark is publicly available.
SurvCORN: Survival Analysis with Conditional Ordinal Ranking Neural Network
Sep 30, 2024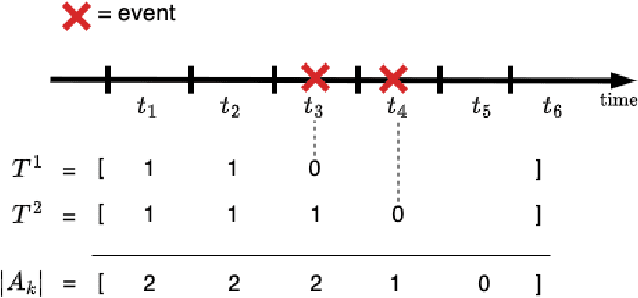
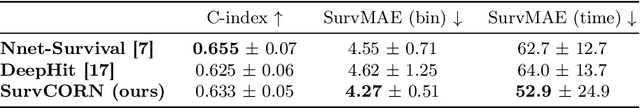
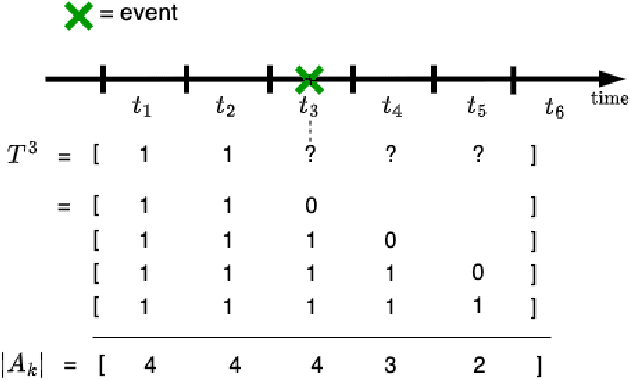
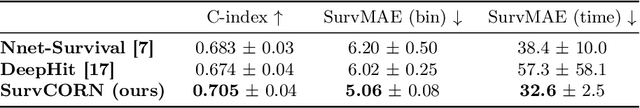
Abstract:Survival analysis plays a crucial role in estimating the likelihood of future events for patients by modeling time-to-event data, particularly in healthcare settings where predictions about outcomes such as death and disease recurrence are essential. However, this analysis poses challenges due to the presence of censored data, where time-to-event information is missing for certain data points. Yet, censored data can offer valuable insights, provided we appropriately incorporate the censoring time during modeling. In this paper, we propose SurvCORN, a novel method utilizing conditional ordinal ranking networks to predict survival curves directly. Additionally, we introduce SurvMAE, a metric designed to evaluate the accuracy of model predictions in estimating time-to-event outcomes. Through empirical evaluation on two real-world cancer datasets, we demonstrate SurvCORN's ability to maintain accurate ordering between patient outcomes while improving individual time-to-event predictions. Our contributions extend recent advancements in ordinal regression to survival analysis, offering valuable insights into accurate prognosis in healthcare settings.
HuLP: Human-in-the-Loop for Prognosis
Mar 19, 2024Abstract:This paper introduces HuLP, a Human-in-the-Loop for Prognosis model designed to enhance the reliability and interpretability of prognostic models in clinical contexts, especially when faced with the complexities of missing covariates and outcomes. HuLP offers an innovative approach that enables human expert intervention, empowering clinicians to interact with and correct models' predictions, thus fostering collaboration between humans and AI models to produce more accurate prognosis. Additionally, HuLP addresses the challenges of missing data by utilizing neural networks and providing a tailored methodology that effectively handles missing data. Traditional methods often struggle to capture the nuanced variations within patient populations, leading to compromised prognostic predictions. HuLP imputes missing covariates based on imaging features, aligning more closely with clinician workflows and enhancing reliability. We conduct our experiments on two real-world, publicly available medical datasets to demonstrate the superiority of HuLP.
SurvRNC: Learning Ordered Representations for Survival Prediction using Rank-N-Contrast
Mar 15, 2024



Abstract:Predicting the likelihood of survival is of paramount importance for individuals diagnosed with cancer as it provides invaluable information regarding prognosis at an early stage. This knowledge enables the formulation of effective treatment plans that lead to improved patient outcomes. In the past few years, deep learning models have provided a feasible solution for assessing medical images, electronic health records, and genomic data to estimate cancer risk scores. However, these models often fall short of their potential because they struggle to learn regression-aware feature representations. In this study, we propose Survival Rank-N Contrast (SurvRNC) method, which introduces a loss function as a regularizer to obtain an ordered representation based on the survival times. This function can handle censored data and can be incorporated into any survival model to ensure that the learned representation is ordinal. The model was extensively evaluated on a HEad \& NeCK TumOR (HECKTOR) segmentation and the outcome-prediction task dataset. We demonstrate that using the SurvRNC method for training can achieve higher performance on different deep survival models. Additionally, it outperforms state-of-the-art methods by 3.6% on the concordance index. The code is publicly available on https://github.com/numanai/SurvRNC
DGM-DR: Domain Generalization with Mutual Information Regularized Diabetic Retinopathy Classification
Sep 18, 2023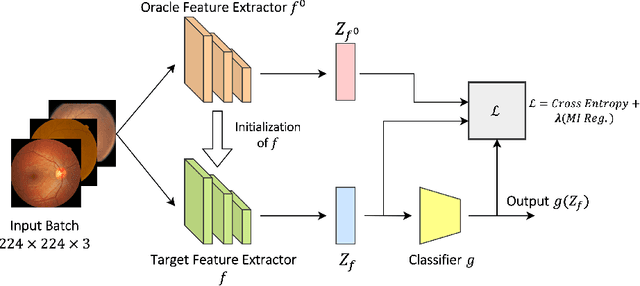
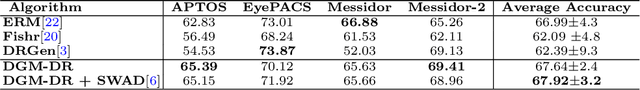
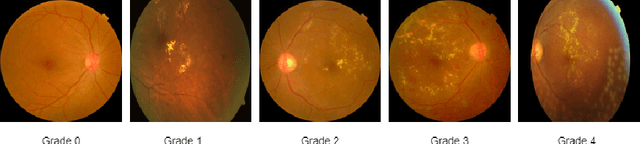
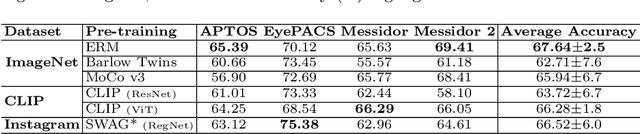
Abstract:The domain shift between training and testing data presents a significant challenge for training generalizable deep learning models. As a consequence, the performance of models trained with the independent and identically distributed (i.i.d) assumption deteriorates when deployed in the real world. This problem is exacerbated in the medical imaging context due to variations in data acquisition across clinical centers, medical apparatus, and patients. Domain generalization (DG) aims to address this problem by learning a model that generalizes well to any unseen target domain. Many domain generalization techniques were unsuccessful in learning domain-invariant representations due to the large domain shift. Furthermore, multiple tasks in medical imaging are not yet extensively studied in existing literature when it comes to DG point of view. In this paper, we introduce a DG method that re-establishes the model objective function as a maximization of mutual information with a large pretrained model to the medical imaging field. We re-visit the problem of DG in Diabetic Retinopathy (DR) classification to establish a clear benchmark with a correct model selection strategy and to achieve robust domain-invariant representation for an improved generalization. Moreover, we conduct extensive experiments on public datasets to show that our proposed method consistently outperforms the previous state-of-the-art by a margin of 5.25% in average accuracy and a lower standard deviation. Source code available at https://github.com/BioMedIA-MBZUAI/DGM-DR
Prompt-based Tuning of Transformer Models for Multi-Center Medical Image Segmentation
May 30, 2023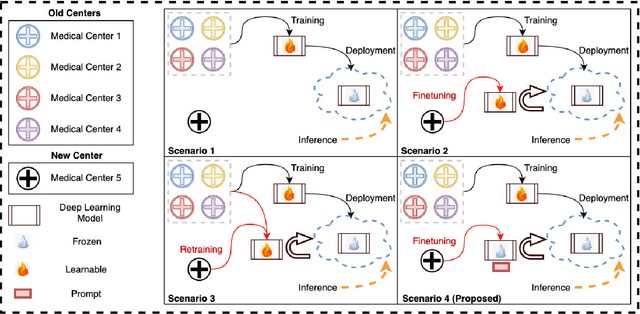
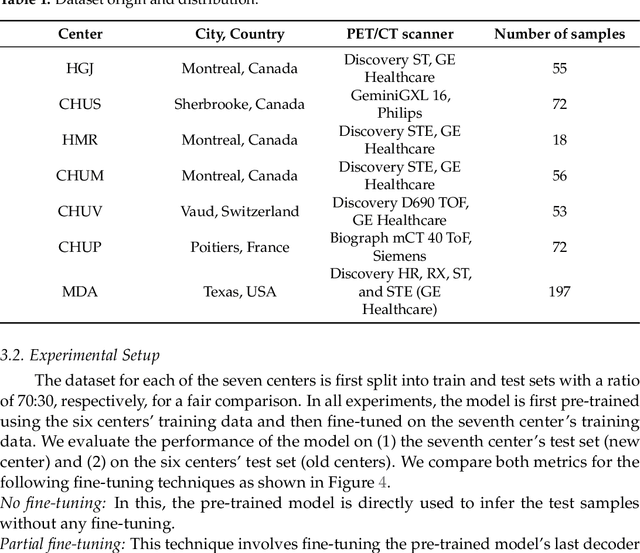
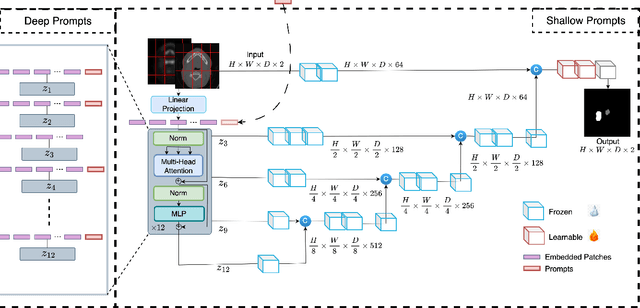
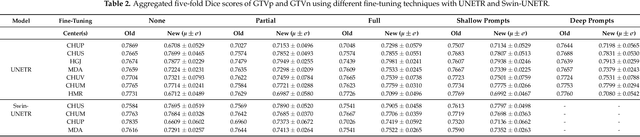
Abstract:Medical image segmentation is a vital healthcare endeavor requiring precise and efficient models for appropriate diagnosis and treatment. Vision transformer-based segmentation models have shown great performance in accomplishing this task. However, to build a powerful backbone, the self-attention block of ViT requires large-scale pre-training data. The present method of modifying pre-trained models entails updating all or some of the backbone parameters. This paper proposes a novel fine-tuning strategy for adapting a pretrained transformer-based segmentation model on data from a new medical center. This method introduces a small number of learnable parameters, termed prompts, into the input space (less than 1\% of model parameters) while keeping the rest of the model parameters frozen. Extensive studies employing data from new unseen medical centers show that prompts-based fine-tuning of medical segmentation models provides excellent performance on the new center data with a negligible drop on the old centers. Additionally, our strategy delivers great accuracy with minimum re-training on new center data, significantly decreasing the computational and time costs of fine-tuning pre-trained models.
MGMT promoter methylation status prediction using MRI scans? An extensive experimental evaluation of deep learning models
Apr 03, 2023Abstract:The number of studies on deep learning for medical diagnosis is expanding, and these systems are often claimed to outperform clinicians. However, only a few systems have shown medical efficacy. From this perspective, we examine a wide range of deep learning algorithms for the assessment of glioblastoma - a common brain tumor in older adults that is lethal. Surgery, chemotherapy, and radiation are the standard treatments for glioblastoma patients. The methylation status of the MGMT promoter, a specific genetic sequence found in the tumor, affects chemotherapy's effectiveness. MGMT promoter methylation improves chemotherapy response and survival in several cancers. MGMT promoter methylation is determined by a tumor tissue biopsy, which is then genetically tested. This lengthy and invasive procedure increases the risk of infection and other complications. Thus, researchers have used deep learning models to examine the tumor from brain MRI scans to determine the MGMT promoter's methylation state. We employ deep learning models and one of the largest public MRI datasets of 585 participants to predict the methylation status of the MGMT promoter in glioblastoma tumors using MRI scans. We test these models using Grad-CAM, occlusion sensitivity, feature visualizations, and training loss landscapes. Our results show no correlation between these two, indicating that external cohort data should be used to verify these models' performance to assure the accuracy and reliability of deep learning systems in cancer diagnosis.
CoNIC Challenge: Pushing the Frontiers of Nuclear Detection, Segmentation, Classification and Counting
Mar 14, 2023



Abstract:Nuclear detection, segmentation and morphometric profiling are essential in helping us further understand the relationship between histology and patient outcome. To drive innovation in this area, we setup a community-wide challenge using the largest available dataset of its kind to assess nuclear segmentation and cellular composition. Our challenge, named CoNIC, stimulated the development of reproducible algorithms for cellular recognition with real-time result inspection on public leaderboards. We conducted an extensive post-challenge analysis based on the top-performing models using 1,658 whole-slide images of colon tissue. With around 700 million detected nuclei per model, associated features were used for dysplasia grading and survival analysis, where we demonstrated that the challenge's improvement over the previous state-of-the-art led to significant boosts in downstream performance. Our findings also suggest that eosinophils and neutrophils play an important role in the tumour microevironment. We release challenge models and WSI-level results to foster the development of further methods for biomarker discovery.
Color Space-based HoVer-Net for Nuclei Instance Segmentation and Classification
Mar 03, 2022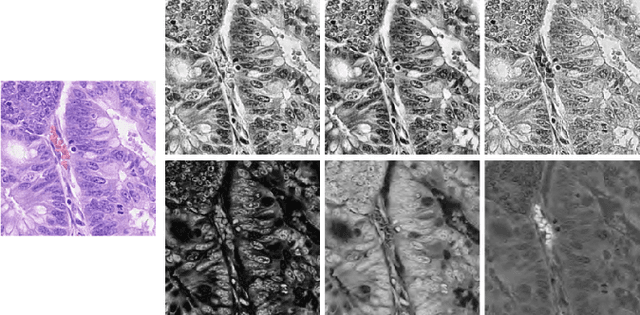
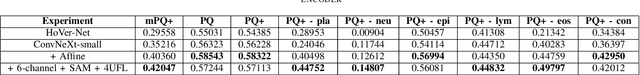
Abstract:Nuclei segmentation and classification is the first and most crucial step that is utilized for many different microscopy medical analysis applications. However, it suffers from many issues such as the segmentation of small objects, imbalance, and fine-grained differences between types of nuclei. In this paper, multiple different contributions were done tackling these problems present. Firstly, the recently released "ConvNeXt" was used as the encoder for HoVer-Net model since it leverages the key components of transformers that make them perform well. Secondly, to enhance the visual differences between nuclei, a multi-channel color space-based approach is used to aid the model in extracting distinguishing features. Thirdly, Unified Focal loss (UFL) was used to tackle the background-foreground imbalance. Finally, Sharpness-Aware Minimization (SAM) was used to ensure generalizability of the model. Overall, we were able to outperform the current state-of-the-art (SOTA), HoVer-Net, on the preliminary test set of the CoNiC Challenge 2022 by 12.489% mPQ+.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge