Color Space-based HoVer-Net for Nuclei Instance Segmentation and Classification
Paper and Code
Mar 03, 2022
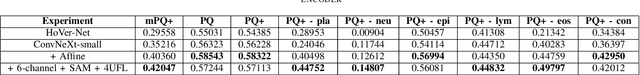
Nuclei segmentation and classification is the first and most crucial step that is utilized for many different microscopy medical analysis applications. However, it suffers from many issues such as the segmentation of small objects, imbalance, and fine-grained differences between types of nuclei. In this paper, multiple different contributions were done tackling these problems present. Firstly, the recently released "ConvNeXt" was used as the encoder for HoVer-Net model since it leverages the key components of transformers that make them perform well. Secondly, to enhance the visual differences between nuclei, a multi-channel color space-based approach is used to aid the model in extracting distinguishing features. Thirdly, Unified Focal loss (UFL) was used to tackle the background-foreground imbalance. Finally, Sharpness-Aware Minimization (SAM) was used to ensure generalizability of the model. Overall, we were able to outperform the current state-of-the-art (SOTA), HoVer-Net, on the preliminary test set of the CoNiC Challenge 2022 by 12.489% mPQ+.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge