Roba Al Majzoub
How Good is my Histopathology Vision-Language Foundation Model? A Holistic Benchmark
Mar 17, 2025



Abstract:Recently, histopathology vision-language foundation models (VLMs) have gained popularity due to their enhanced performance and generalizability across different downstream tasks. However, most existing histopathology benchmarks are either unimodal or limited in terms of diversity of clinical tasks, organs, and acquisition instruments, as well as their partial availability to the public due to patient data privacy. As a consequence, there is a lack of comprehensive evaluation of existing histopathology VLMs on a unified benchmark setting that better reflects a wide range of clinical scenarios. To address this gap, we introduce HistoVL, a fully open-source comprehensive benchmark comprising images acquired using up to 11 various acquisition tools that are paired with specifically crafted captions by incorporating class names and diverse pathology descriptions. Our Histo-VL includes 26 organs, 31 cancer types, and a wide variety of tissue obtained from 14 heterogeneous patient cohorts, totaling more than 5 million patches obtained from over 41K WSIs viewed under various magnification levels. We systematically evaluate existing histopathology VLMs on Histo-VL to simulate diverse tasks performed by experts in real-world clinical scenarios. Our analysis reveals interesting findings, including large sensitivity of most existing histopathology VLMs to textual changes with a drop in balanced accuracy of up to 25% in tasks such as Metastasis detection, low robustness to adversarial attacks, as well as improper calibration of models evident through high ECE values and low model prediction confidence, all of which can affect their clinical implementation.
Distilling Local Texture Features for Colorectal Tissue Classification in Low Data Regimes
Jan 02, 2024Abstract:Multi-class colorectal tissue classification is a challenging problem that is typically addressed in a setting, where it is assumed that ample amounts of training data is available. However, manual annotation of fine-grained colorectal tissue samples of multiple classes, especially the rare ones like stromal tumor and anal cancer is laborious and expensive. To address this, we propose a knowledge distillation-based approach, named KD-CTCNet, that effectively captures local texture information from few tissue samples, through a distillation loss, to improve the standard CNN features. The resulting enriched feature representation achieves improved classification performance specifically in low data regimes. Extensive experiments on two public datasets of colorectal tissues reveal the merits of the proposed contributions, with a consistent gain achieved over different approaches across low data settings. The code and models are publicly available on GitHub.
Prompt-based Tuning of Transformer Models for Multi-Center Medical Image Segmentation
May 30, 2023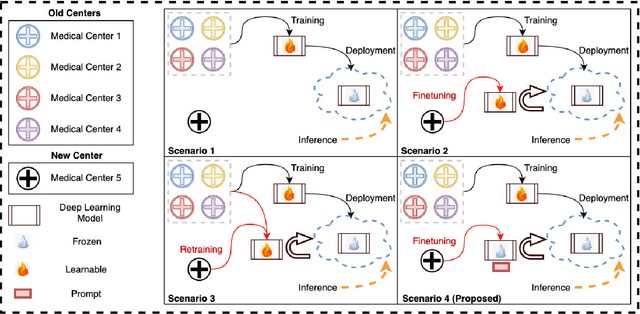
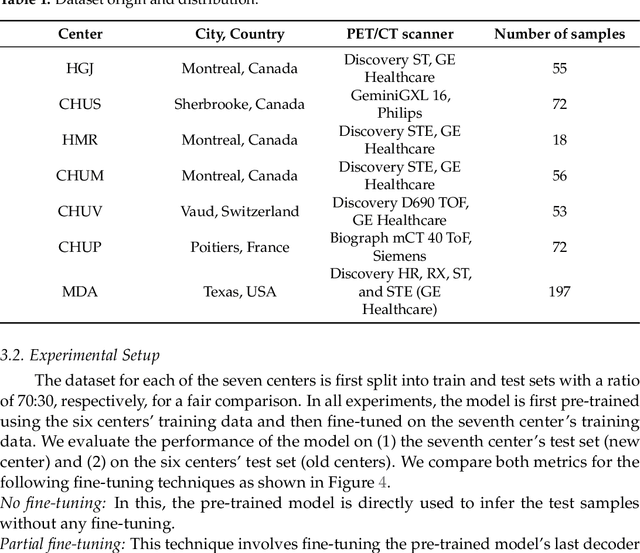
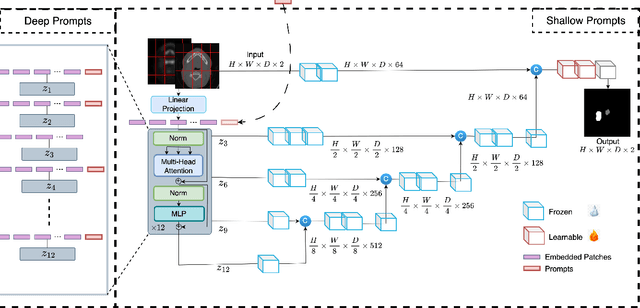
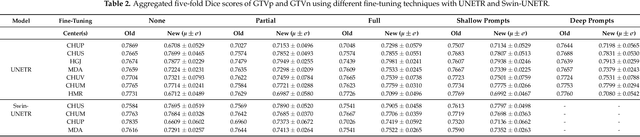
Abstract:Medical image segmentation is a vital healthcare endeavor requiring precise and efficient models for appropriate diagnosis and treatment. Vision transformer-based segmentation models have shown great performance in accomplishing this task. However, to build a powerful backbone, the self-attention block of ViT requires large-scale pre-training data. The present method of modifying pre-trained models entails updating all or some of the backbone parameters. This paper proposes a novel fine-tuning strategy for adapting a pretrained transformer-based segmentation model on data from a new medical center. This method introduces a small number of learnable parameters, termed prompts, into the input space (less than 1\% of model parameters) while keeping the rest of the model parameters frozen. Extensive studies employing data from new unseen medical centers show that prompts-based fine-tuning of medical segmentation models provides excellent performance on the new center data with a negligible drop on the old centers. Additionally, our strategy delivers great accuracy with minimum re-training on new center data, significantly decreasing the computational and time costs of fine-tuning pre-trained models.
TMSS: An End-to-End Transformer-based Multimodal Network for Segmentation and Survival Prediction
Sep 12, 2022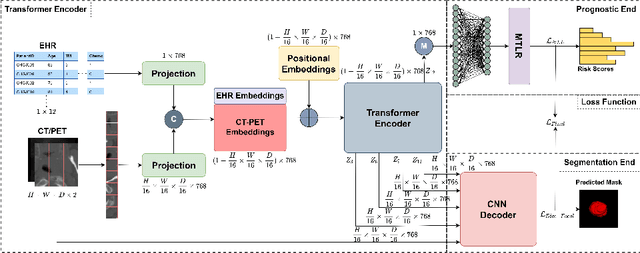

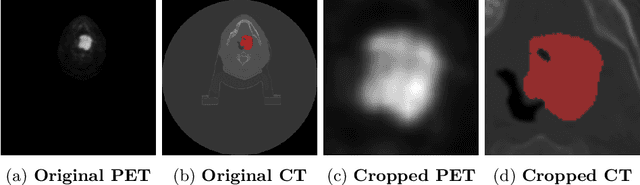
Abstract:When oncologists estimate cancer patient survival, they rely on multimodal data. Even though some multimodal deep learning methods have been proposed in the literature, the majority rely on having two or more independent networks that share knowledge at a later stage in the overall model. On the other hand, oncologists do not do this in their analysis but rather fuse the information in their brain from multiple sources such as medical images and patient history. This work proposes a deep learning method that mimics oncologists' analytical behavior when quantifying cancer and estimating patient survival. We propose TMSS, an end-to-end Transformer based Multimodal network for Segmentation and Survival prediction that leverages the superiority of transformers that lies in their abilities to handle different modalities. The model was trained and validated for segmentation and prognosis tasks on the training dataset from the HEad & NeCK TumOR segmentation and the outcome prediction in PET/CT images challenge (HECKTOR). We show that the proposed prognostic model significantly outperforms state-of-the-art methods with a concordance index of 0.763+/-0.14 while achieving a comparable dice score of 0.772+/-0.030 to a standalone segmentation model. The code is publicly available.
An Ensemble Approach for Patient Prognosis of Head and Neck Tumor Using Multimodal Data
Feb 25, 2022



Abstract:Accurate prognosis of a tumor can help doctors provide a proper course of treatment and, therefore, save the lives of many. Traditional machine learning algorithms have been eminently useful in crafting prognostic models in the last few decades. Recently, deep learning algorithms have shown significant improvement when developing diagnosis and prognosis solutions to different healthcare problems. However, most of these solutions rely solely on either imaging or clinical data. Utilizing patient tabular data such as demographics and patient medical history alongside imaging data in a multimodal approach to solve a prognosis task has started to gain more interest recently and has the potential to create more accurate solutions. The main issue when using clinical and imaging data to train a deep learning model is to decide on how to combine the information from these sources. We propose a multimodal network that ensembles deep multi-task logistic regression (MTLR), Cox proportional hazard (CoxPH) and CNN models to predict prognostic outcomes for patients with head and neck tumors using patients' clinical and imaging (CT and PET) data. Features from CT and PET scans are fused and then combined with patients' electronic health records for the prediction. The proposed model is trained and tested on 224 and 101 patient records respectively. Experimental results show that our proposed ensemble solution achieves a C-index of 0.72 on The HECKTOR test set that saved us the first place in prognosis task of the HECKTOR challenge. The full implementation based on PyTorch is available on \url{https://github.com/numanai/BioMedIA-Hecktor2021}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge