Liang Hong
Hierarchical Flow Decomposition for Turning Movement Prediction at Signalized Intersections
Apr 10, 2026Abstract:Accurate prediction of intersection turning movements is essential for adaptive signal control but remains difficult due to the high volatility of directional flows. This study proposes HFD-TM (Hierarchical Flow-Decomposition for Turning Movement Prediction), a hierarchical deep learning framework that predicts turning movements by first forecasting corridor through-movements and then expanding these predictions to individual turning streams. This design is motivated by empirical traffic structure, where corridor flows account for 65.1% of total volume, exhibit lower volatility than turning movements, and explain 35.5% of turning-movement variance. A physics-informed loss function enforces flow conservation to maintain structural consistency. Evaluated on six months of 15-minute interval LiDAR (Light Detection and Ranging) data from a six-intersection corridor in Nashville, Tennessee, HFD-TM achieves a mean absolute error of 2.49 vehicles per interval, reducing MAE by 5.7% compared to a Transformer and by 27.0% compared to a GRU (Gated Recurrent Unit). Ablation results show that hierarchical decomposition provides the largest performance gain, while training time is 12.8 times lower than DCRNN (Diffusion Convolutional Recurrent Neural Network), demonstrating suitability for real-time traffic applications.
Physics-Embedded Feature Learning for AI in Medical Imaging
Mar 30, 2026Abstract:Deep learning (DL) models have achieved strong performance in an intelligence healthcare setting, yet most existing approaches operate as black boxes and ignore the physical processes that govern tumor growth, limiting interpretability, robustness, and clinical trust. To address this limitation, we propose PhysNet, a physics-embedded DL framework that integrates tumor growth dynamics directly into the feature learning process of a convolutional neural network (CNN). Unlike conventional physics-informed methods that impose physical constraints only at the output level, PhysNet embeds a reaction diffusion model of tumor growth within intermediate feature representations of a ResNet backbone. The architecture jointly performs multi-class tumor classification while learning a latent tumor density field, its temporal evolution, and biologically meaningful physical parameters, including tumor diffusion and growth rates, through end-to-end training. This design is necessary because purely data-driven models, even when highly accurate or ensemble-based, cannot guarantee physically consistent predictions or provide insight into tumor behavior. Experimental results on a large brain MRI dataset demonstrate that PhysNet outperforms multiple state-of-the-art DL baselines, including MobileNetV2, VGG16, VGG19, and ensemble models, achieving superior classification accuracy and F1-score. In addition to improved performance, PhysNet produces interpretable latent representations and learned bio-physical parameters that align with established medical knowledge, highlighting physics-embedded representation learning as a practical pathway toward more trustworthy and clinically meaningful medical AI systems.
Self-evolving AI agents for protein discovery and directed evolution
Mar 28, 2026Abstract:Protein scientific discovery is bottlenecked by the manual orchestration of information and algorithms, while general agents are insufficient in complex domain projects. VenusFactory2 provides an autonomous framework that shifts from static tool usage to dynamic workflow synthesis via a self-evolving multi-agent infrastructure to address protein-related demands. It outperforms a set of well-known agents on the VenusAgentEval benchmark, and autonomously organizes the discovery and optimization of proteins from a single natural language prompt.
Rank-and-Reason: Multi-Agent Collaboration Accelerates Zero-Shot Protein Mutation Prediction
Feb 03, 2026Abstract:Zero-shot mutation prediction is vital for low-resource protein engineering, yet existing protein language models (PLMs) often yield statistically confident results that ignore fundamental biophysical constraints. Currently, selecting candidates for wet-lab validation relies on manual expert auditing of PLM outputs, a process that is inefficient, subjective, and highly dependent on domain expertise. To address this, we propose Rank-and-Reason (VenusRAR), a two-stage agentic framework to automate this workflow and maximize expected wet-lab fitness. In the Rank-Stage, a Computational Expert and Virtual Biologist aggregate a context-aware multi-modal ensemble, establishing a new Spearman correlation record of 0.551 (vs. 0.518) on ProteinGym. In the Reason-Stage, an agentic Expert Panel employs chain-of-thought reasoning to audit candidates against geometric and structural constraints, improving the Top-5 Hit Rate by up to 367% on ProteinGym-DMS99. The wet-lab validation on Cas12i3 nuclease further confirms the framework's efficacy, achieving a 46.7% positive rate and identifying two novel mutants with 4.23-fold and 5.05-fold activity improvements. Code and datasets are released on GitHub (https://github.com/ai4protein/VenusRAR/).
A new strategy for finite-sample valid prediction of future insurance claims in the regression setting
Jan 29, 2026Abstract:The extant insurance literature demonstrates a paucity of finite-sample valid prediction intervals of future insurance claims in the regression setting. To address this challenge, this article proposes a new strategy that converts a predictive method in the unsupervised iid (independent identically distributed) setting to a predictive method in the regression setting. In particular, it enables an actuary to obtain infinitely many finite-sample valid prediction intervals in the regression setting.
VenusX: Unlocking Fine-Grained Functional Understanding of Proteins
May 17, 2025



Abstract:Deep learning models have driven significant progress in predicting protein function and interactions at the protein level. While these advancements have been invaluable for many biological applications such as enzyme engineering and function annotation, a more detailed perspective is essential for understanding protein functional mechanisms and evaluating the biological knowledge captured by models. To address this demand, we introduce VenusX, the first large-scale benchmark for fine-grained functional annotation and function-based protein pairing at the residue, fragment, and domain levels. VenusX comprises three major task categories across six types of annotations, including residue-level binary classification, fragment-level multi-class classification, and pairwise functional similarity scoring for identifying critical active sites, binding sites, conserved sites, motifs, domains, and epitopes. The benchmark features over 878,000 samples curated from major open-source databases such as InterPro, BioLiP, and SAbDab. By providing mixed-family and cross-family splits at three sequence identity thresholds, our benchmark enables a comprehensive assessment of model performance on both in-distribution and out-of-distribution scenarios. For baseline evaluation, we assess a diverse set of popular and open-source models, including pre-trained protein language models, sequence-structure hybrids, structure-based methods, and alignment-based techniques. Their performance is reported across all benchmark datasets and evaluation settings using multiple metrics, offering a thorough comparison and a strong foundation for future research. Code and data are publicly available at https://github.com/ai4protein/VenusX.
VenusFactory: A Unified Platform for Protein Engineering Data Retrieval and Language Model Fine-Tuning
Mar 19, 2025Abstract:Natural language processing (NLP) has significantly influenced scientific domains beyond human language, including protein engineering, where pre-trained protein language models (PLMs) have demonstrated remarkable success. However, interdisciplinary adoption remains limited due to challenges in data collection, task benchmarking, and application. This work presents VenusFactory, a versatile engine that integrates biological data retrieval, standardized task benchmarking, and modular fine-tuning of PLMs. VenusFactory supports both computer science and biology communities with choices of both a command-line execution and a Gradio-based no-code interface, integrating $40+$ protein-related datasets and $40+$ popular PLMs. All implementations are open-sourced on https://github.com/tyang816/VenusFactory.
Finite-sample valid prediction of future insurance claims in the regression problem
Mar 05, 2025Abstract:In the current insurance literature, prediction of insurance claims in the regression problem is often performed with a statistical model. This model-based approach may suffer from several drawbacks: (i) model misspecification, (ii) selection effect, and (iii) lack of finite-sample validity. This article addresses these three issues simultaneously by employing conformal prediction-a general machine learning strategy for valid predictions. The proposed method is both model-free and tuning-parameter-free. It also guarantees finite-sample validity at a pre-assigned coverage probability level.
AI-Driven Secure Data Sharing: A Trustworthy and Privacy-Preserving Approach
Jan 26, 2025



Abstract:In the era of data-driven decision-making, ensuring the privacy and security of shared data is paramount across various domains. Applying existing deep neural networks (DNNs) to encrypted data is critical and often compromises performance, security, and computational overhead. To address these limitations, this research introduces a secure framework consisting of a learnable encryption method based on the block-pixel operation to encrypt the data and subsequently integrate it with the Vision Transformer (ViT). The proposed framework ensures data privacy and security by creating unique scrambling patterns per key, providing robust performance against adversarial attacks without compromising computational efficiency and data integrity. The framework was tested on sensitive medical datasets to validate its efficacy, proving its ability to handle highly confidential information securely. The suggested framework was validated with a 94\% success rate after extensive testing on real-world datasets, such as MRI brain tumors and histological scans of lung and colon cancers. Additionally, the framework was tested under diverse adversarial attempts against secure data sharing with optimum performance and demonstrated its effectiveness in various threat scenarios. These comprehensive analyses underscore its robustness, making it a trustworthy solution for secure data sharing in critical applications.
Retrieval-Enhanced Mutation Mastery: Augmenting Zero-Shot Prediction of Protein Language Model
Oct 28, 2024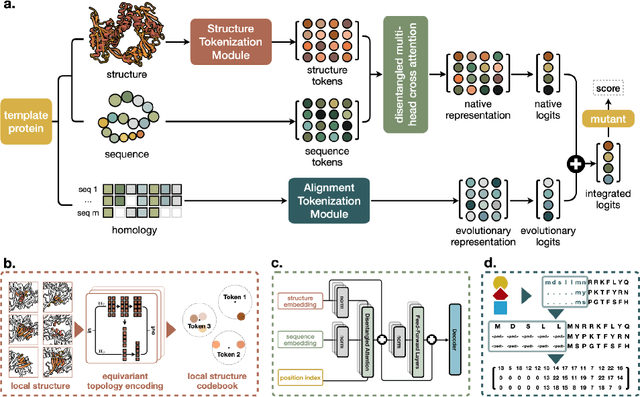
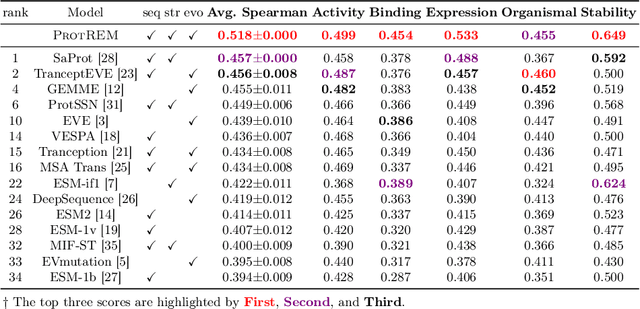
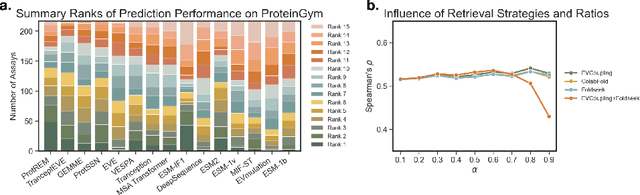
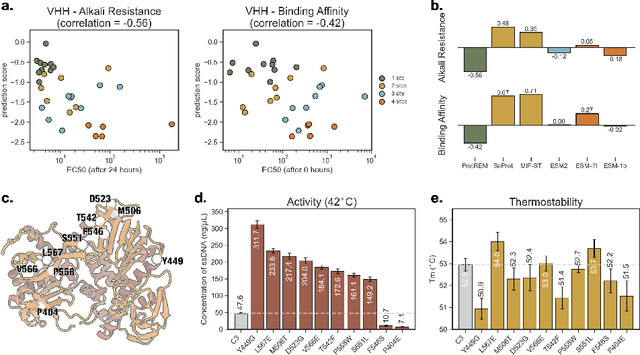
Abstract:Enzyme engineering enables the modification of wild-type proteins to meet industrial and research demands by enhancing catalytic activity, stability, binding affinities, and other properties. The emergence of deep learning methods for protein modeling has demonstrated superior results at lower costs compared to traditional approaches such as directed evolution and rational design. In mutation effect prediction, the key to pre-training deep learning models lies in accurately interpreting the complex relationships among protein sequence, structure, and function. This study introduces a retrieval-enhanced protein language model for comprehensive analysis of native properties from sequence and local structural interactions, as well as evolutionary properties from retrieved homologous sequences. The state-of-the-art performance of the proposed ProtREM is validated on over 2 million mutants across 217 assays from an open benchmark (ProteinGym). We also conducted post-hoc analyses of the model's ability to improve the stability and binding affinity of a VHH antibody. Additionally, we designed 10 new mutants on a DNA polymerase and conducted wet-lab experiments to evaluate their enhanced activity at higher temperatures. Both in silico and experimental evaluations confirmed that our method provides reliable predictions of mutation effects, offering an auxiliary tool for biologists aiming to evolve existing enzymes. The implementation is publicly available at https://github.com/tyang816/ProtREM.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge