Julia E. Vogt
Rethinking Machine Unlearning: Models Designed to Forget via Key Deletion
Mar 16, 2026Abstract:Machine unlearning is rapidly becoming a practical requirement, driven by privacy regulations, data errors, and the need to remove harmful or corrupted training samples. Despite this, most existing methods tackle the problem purely from a post-hoc perspective. They attempt to erase the influence of targeted training samples through parameter updates that typically require access to the full training data. This creates a mismatch with real deployment scenarios where unlearning requests can be anticipated, revealing a fundamental limitation of post-hoc approaches. We propose \textit{unlearning by design}, a novel paradigm in which models are directly trained to support forgetting as an inherent capability. We instantiate this idea with Machine UNlearning via KEY deletion (MUNKEY), a memory augmented transformer that decouples instance-specific memorization from model weights. Here, unlearning corresponds to removing the instance-identifying key, enabling direct zero-shot forgetting without weight updates or access to the original samples or labels. Across natural image benchmarks, fine-grained recognition, and medical datasets, MUNKEY outperforms all post-hoc baselines. Our results establish that unlearning by design enables fast, deployment-oriented unlearning while preserving predictive performance.
Steering Generative Models for Accessibility: EasyRead Image Generation
Mar 14, 2026Abstract:EasyRead pictograms are simple, visually clear images that represent specific concepts and support comprehension for people with intellectual disabilities, low literacy, or language barriers. The large-scale production of EasyRead content has traditionally been constrained by the cost and expertise required to manually design pictograms. In contrast, automatic generation of such images could significantly reduce production time and cost, enabling broader accessibility across digital and printed materials. However, modern diffusion-based image generation models tend to produce outputs that exhibit excessive visual detail and lack stylistic stability across random seeds, limiting their suitability for clear and consistent pictogram generation. This challenge highlights the need for methods specifically tailored to accessibility-oriented visual content. In this work, we present a unified pipeline for generating EasyRead pictograms by fine-tuning a Stable Diffusion model using LoRA adapters on a curated corpus that combines augmented samples from multiple pictogram datasets. Since EasyRead pictograms lack a unified formal definition, we introduce an EasyRead score to benchmark pictogram quality and consistency. Our results demonstrate that diffusion models can be effectively steered toward producing coherent EasyRead-style images, indicating that generative models can serve as practical tools for scalable and accessible pictogram production.
Reference-Guided Machine Unlearning
Mar 11, 2026Abstract:Machine unlearning aims to remove the influence of specific data from trained models while preserving general utility. Existing approximate unlearning methods often rely on performance-degradation heuristics, such as loss maximization or random labeling. However, these signals can be poorly conditioned, leading to unstable optimization and harming the model's generalization. We argue that unlearning should instead prioritize distributional indistinguishability, aligning the model's behavior on forget data with its behavior on truly unseen data. Motivated by this, we propose Reference-Guided Unlearning (ReGUn), a framework that leverages a disjoint held-out dataset to provide a principled, class-conditioned reference for distillation. We demonstrate across various model architectures, natural image datasets, and varying forget fractions that ReGUn consistently outperforms standard approximate baselines, achieving a superior forgetting-utility trade-off.
You Only Train Once: Differentiable Subset Selection for Omics Data
Dec 19, 2025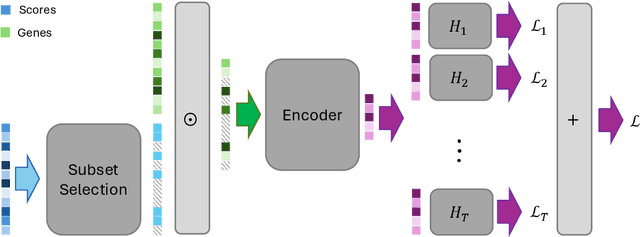
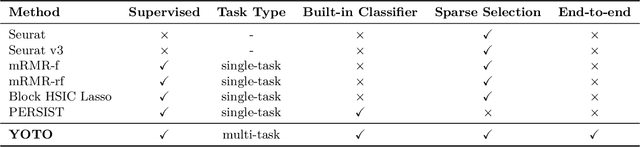

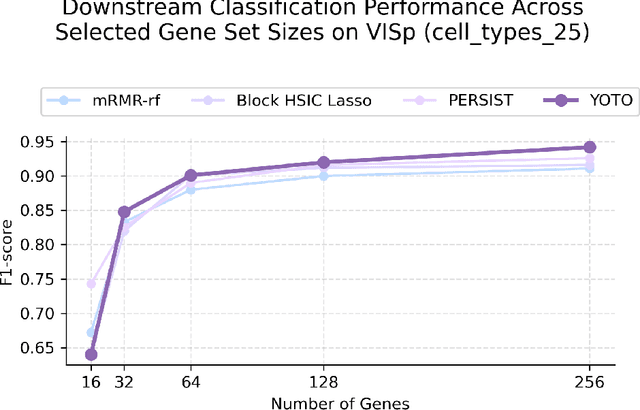
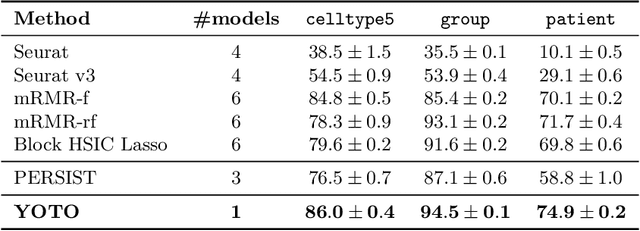
Abstract:Selecting compact and informative gene subsets from single-cell transcriptomic data is essential for biomarker discovery, improving interpretability, and cost-effective profiling. However, most existing feature selection approaches either operate as multi-stage pipelines or rely on post hoc feature attribution, making selection and prediction weakly coupled. In this work, we present YOTO (you only train once), an end-to-end framework that jointly identifies discrete gene subsets and performs prediction within a single differentiable architecture. In our model, the prediction task directly guides which genes are selected, while the learned subsets, in turn, shape the predictive representation. This closed feedback loop enables the model to iteratively refine both what it selects and how it predicts during training. Unlike existing approaches, YOTO enforces sparsity so that only the selected genes contribute to inference, eliminating the need to train additional downstream classifiers. Through a multi-task learning design, the model learns shared representations across related objectives, allowing partially labeled datasets to inform one another, and discovering gene subsets that generalize across tasks without additional training steps. We evaluate YOTO on two representative single-cell RNA-seq datasets, showing that it consistently outperforms state-of-the-art baselines. These results demonstrate that sparse, end-to-end, multi-task gene subset selection improves predictive performance and yields compact and meaningful gene subsets, advancing biomarker discovery and single-cell analysis.
Enhancing Radiology Report Generation and Visual Grounding using Reinforcement Learning
Dec 11, 2025Abstract:Recent advances in vision-language models (VLMs) have improved Chest X-ray (CXR) interpretation in multiple aspects. However, many medical VLMs rely solely on supervised fine-tuning (SFT), which optimizes next-token prediction without evaluating answer quality. In contrast, reinforcement learning (RL) can incorporate task-specific feedback, and its combination with explicit intermediate reasoning ("thinking") has demonstrated substantial gains on verifiable math and coding tasks. To investigate the effects of RL and thinking in a CXR VLM, we perform large-scale SFT on CXR data to build an updated RadVLM based on Qwen3-VL, followed by a cold-start SFT stage that equips the model with basic thinking ability. We then apply Group Relative Policy Optimization (GRPO) with clinically grounded, task-specific rewards for report generation and visual grounding, and run matched RL experiments on both domain-specific and general-domain Qwen3-VL variants, with and without thinking. Across these settings, we find that while strong SFT remains crucial for high base performance, RL provides additional gains on both tasks, whereas explicit thinking does not appear to further improve results. Under a unified evaluation pipeline, the RL-optimized RadVLM models outperform their baseline counterparts and reach state-of-the-art performance on both report generation and grounding, highlighting clinically aligned RL as a powerful complement to SFT for medical VLMs.
From Slices to Structures: Unsupervised 3D Reconstruction of Female Pelvic Anatomy from Freehand Transvaginal Ultrasound
Aug 20, 2025Abstract:Volumetric ultrasound has the potential to significantly improve diagnostic accuracy and clinical decision-making, yet its widespread adoption remains limited by dependence on specialized hardware and restrictive acquisition protocols. In this work, we present a novel unsupervised framework for reconstructing 3D anatomical structures from freehand 2D transvaginal ultrasound (TVS) sweeps, without requiring external tracking or learned pose estimators. Our method adapts the principles of Gaussian Splatting to the domain of ultrasound, introducing a slice-aware, differentiable rasterizer tailored to the unique physics and geometry of ultrasound imaging. We model anatomy as a collection of anisotropic 3D Gaussians and optimize their parameters directly from image-level supervision, leveraging sensorless probe motion estimation and domain-specific geometric priors. The result is a compact, flexible, and memory-efficient volumetric representation that captures anatomical detail with high spatial fidelity. This work demonstrates that accurate 3D reconstruction from 2D ultrasound images can be achieved through purely computational means, offering a scalable alternative to conventional 3D systems and enabling new opportunities for AI-assisted analysis and diagnosis.
From Pixels to Perception: Interpretable Predictions via Instance-wise Grouped Feature Selection
May 09, 2025Abstract:Understanding the decision-making process of machine learning models provides valuable insights into the task, the data, and the reasons behind a model's failures. In this work, we propose a method that performs inherently interpretable predictions through the instance-wise sparsification of input images. To align the sparsification with human perception, we learn the masking in the space of semantically meaningful pixel regions rather than on pixel-level. Additionally, we introduce an explicit way to dynamically determine the required level of sparsity for each instance. We show empirically on semi-synthetic and natural image datasets that our inherently interpretable classifier produces more meaningful, human-understandable predictions than state-of-the-art benchmarks.
Revisiting Automatic Data Curation for Vision Foundation Models in Digital Pathology
Mar 24, 2025Abstract:Vision foundation models (FMs) are accelerating the development of digital pathology algorithms and transforming biomedical research. These models learn, in a self-supervised manner, to represent histological features in highly heterogeneous tiles extracted from whole-slide images (WSIs) of real-world patient samples. The performance of these FMs is significantly influenced by the size, diversity, and balance of the pre-training data. However, data selection has been primarily guided by expert knowledge at the WSI level, focusing on factors such as disease classification and tissue types, while largely overlooking the granular details available at the tile level. In this paper, we investigate the potential of unsupervised automatic data curation at the tile-level, taking into account 350 million tiles. Specifically, we apply hierarchical clustering trees to pre-extracted tile embeddings, allowing us to sample balanced datasets uniformly across the embedding space of the pretrained FM. We further identify these datasets are subject to a trade-off between size and balance, potentially compromising the quality of representations learned by FMs, and propose tailored batch sampling strategies to mitigate this effect. We demonstrate the effectiveness of our method through improved performance on a diverse range of clinically relevant downstream tasks.
From Pixels to Components: Eigenvector Masking for Visual Representation Learning
Feb 10, 2025



Abstract:Predicting masked from visible parts of an image is a powerful self-supervised approach for visual representation learning. However, the common practice of masking random patches of pixels exhibits certain failure modes, which can prevent learning meaningful high-level features, as required for downstream tasks. We propose an alternative masking strategy that operates on a suitable transformation of the data rather than on the raw pixels. Specifically, we perform principal component analysis and then randomly mask a subset of components, which accounts for a fixed ratio of the data variance. The learning task then amounts to reconstructing the masked components from the visible ones. Compared to local patches of pixels, the principal components of images carry more global information. We thus posit that predicting masked from visible components involves more high-level features, allowing our masking strategy to extract more useful representations. This is corroborated by our empirical findings which demonstrate improved image classification performance for component over pixel masking. Our method thus constitutes a simple and robust data-driven alternative to traditional masked image modeling approaches.
RadVLM: A Multitask Conversational Vision-Language Model for Radiology
Feb 05, 2025



Abstract:The widespread use of chest X-rays (CXRs), coupled with a shortage of radiologists, has driven growing interest in automated CXR analysis and AI-assisted reporting. While existing vision-language models (VLMs) show promise in specific tasks such as report generation or abnormality detection, they often lack support for interactive diagnostic capabilities. In this work we present RadVLM, a compact, multitask conversational foundation model designed for CXR interpretation. To this end, we curate a large-scale instruction dataset comprising over 1 million image-instruction pairs containing both single-turn tasks -- such as report generation, abnormality classification, and visual grounding -- and multi-turn, multi-task conversational interactions. After fine-tuning RadVLM on this instruction dataset, we evaluate it across different tasks along with re-implemented baseline VLMs. Our results show that RadVLM achieves state-of-the-art performance in conversational capabilities and visual grounding while remaining competitive in other radiology tasks. Ablation studies further highlight the benefit of joint training across multiple tasks, particularly for scenarios with limited annotated data. Together, these findings highlight the potential of RadVLM as a clinically relevant AI assistant, providing structured CXR interpretation and conversational capabilities to support more effective and accessible diagnostic workflows.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge