Daniel Sobotka
MRI-derived quantification of hepatic vessel-to-volume ratios in chronic liver disease using a deep learning approach
Oct 09, 2025Abstract:Background: We aimed to quantify hepatic vessel volumes across chronic liver disease stages and healthy controls using deep learning-based magnetic resonance imaging (MRI) analysis, and assess correlations with biomarkers for liver (dys)function and fibrosis/portal hypertension. Methods: We assessed retrospectively healthy controls, non-advanced and advanced chronic liver disease (ACLD) patients using a 3D U-Net model for hepatic vessel segmentation on portal venous phase gadoxetic acid-enhanced 3-T MRI. Total (TVVR), hepatic (HVVR), and intrahepatic portal vein-to-volume ratios (PVVR) were compared between groups and correlated with: albumin-bilirubin (ALBI) and model for end-stage liver disease-sodium (MELD-Na) score, and fibrosis/portal hypertension (Fibrosis-4 [FIB-4] score, liver stiffness measurement [LSM], hepatic venous pressure gradient [HVPG], platelet count [PLT], and spleen volume). Results: We included 197 subjects, aged 54.9 $\pm$ 13.8 years (mean $\pm$ standard deviation), 111 males (56.3\%): 35 healthy controls, 44 non-ACLD, and 118 ACLD patients. TVVR and HVVR were highest in controls (3.9; 2.1), intermediate in non-ACLD (2.8; 1.7), and lowest in ACLD patients (2.3; 1.0) ($p \leq 0.001$). PVVR was reduced in both non-ACLD and ACLD patients (both 1.2) compared to controls (1.7) ($p \leq 0.001$), but showed no difference between CLD groups ($p = 0.999$). HVVR significantly correlated indirectly with FIB-4, ALBI, MELD-Na, LSM, and spleen volume ($\rho$ ranging from -0.27 to -0.40), and directly with PLT ($\rho = 0.36$). TVVR and PVVR showed similar but weaker correlations. Conclusions: Deep learning-based hepatic vessel volumetry demonstrated differences between healthy liver and chronic liver disease stages and shows correlations with established markers of disease severity.
Improving Vessel Segmentation with Multi-Task Learning and Auxiliary Data Available Only During Model Training
Sep 04, 2025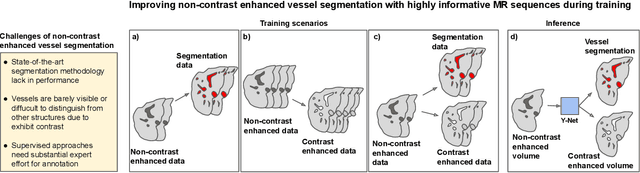

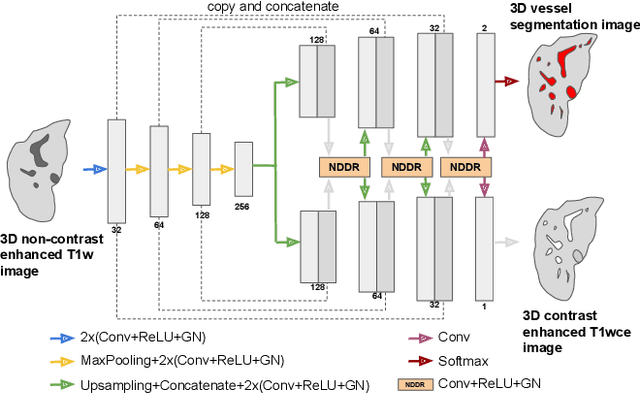

Abstract:Liver vessel segmentation in magnetic resonance imaging data is important for the computational analysis of vascular remodelling, associated with a wide spectrum of diffuse liver diseases. Existing approaches rely on contrast enhanced imaging data, but the necessary dedicated imaging sequences are not uniformly acquired. Images without contrast enhancement are acquired more frequently, but vessel segmentation is challenging, and requires large-scale annotated data. We propose a multi-task learning framework to segment vessels in liver MRI without contrast. It exploits auxiliary contrast enhanced MRI data available only during training to reduce the need for annotated training examples. Our approach draws on paired native and contrast enhanced data with and without vessel annotations for model training. Results show that auxiliary data improves the accuracy of vessel segmentation, even if they are not available during inference. The advantage is most pronounced if only few annotations are available for training, since the feature representation benefits from the shared task structure. A validation of this approach to augment a model for brain tumor segmentation confirms its benefits across different domains. An auxiliary informative imaging modality can augment expert annotations even if it is only available during training.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Fetal Brain Tissue Annotation and Segmentation Challenge Results
Apr 20, 2022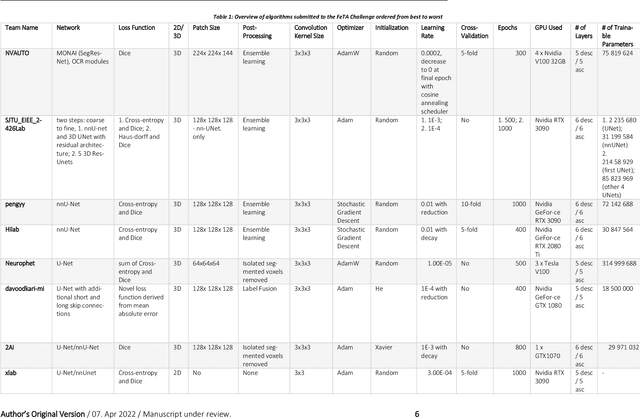
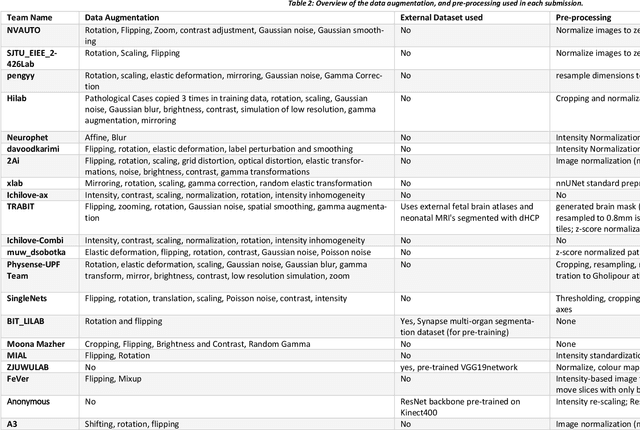
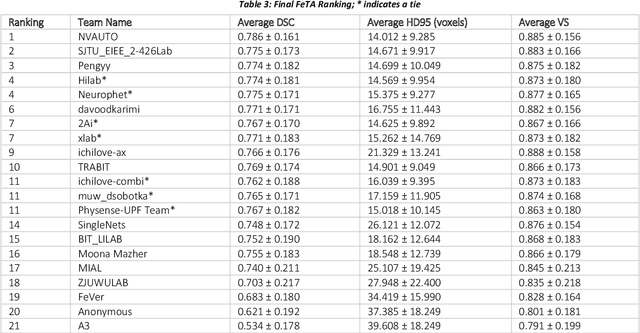
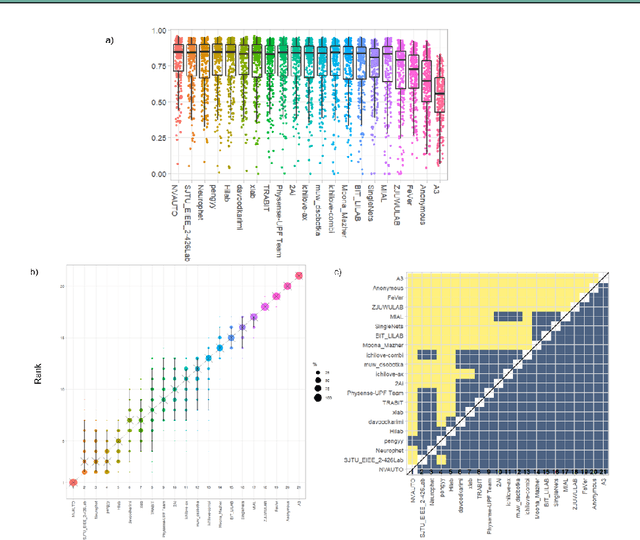
Abstract:In-utero fetal MRI is emerging as an important tool in the diagnosis and analysis of the developing human brain. Automatic segmentation of the developing fetal brain is a vital step in the quantitative analysis of prenatal neurodevelopment both in the research and clinical context. However, manual segmentation of cerebral structures is time-consuming and prone to error and inter-observer variability. Therefore, we organized the Fetal Tissue Annotation (FeTA) Challenge in 2021 in order to encourage the development of automatic segmentation algorithms on an international level. The challenge utilized FeTA Dataset, an open dataset of fetal brain MRI reconstructions segmented into seven different tissues (external cerebrospinal fluid, grey matter, white matter, ventricles, cerebellum, brainstem, deep grey matter). 20 international teams participated in this challenge, submitting a total of 21 algorithms for evaluation. In this paper, we provide a detailed analysis of the results from both a technical and clinical perspective. All participants relied on deep learning methods, mainly U-Nets, with some variability present in the network architecture, optimization, and image pre- and post-processing. The majority of teams used existing medical imaging deep learning frameworks. The main differences between the submissions were the fine tuning done during training, and the specific pre- and post-processing steps performed. The challenge results showed that almost all submissions performed similarly. Four of the top five teams used ensemble learning methods. However, one team's algorithm performed significantly superior to the other submissions, and consisted of an asymmetrical U-Net network architecture. This paper provides a first of its kind benchmark for future automatic multi-tissue segmentation algorithms for the developing human brain in utero.
Motion Correction and Volumetric Reconstruction for Fetal Functional Magnetic Resonance Imaging Data
Feb 11, 2022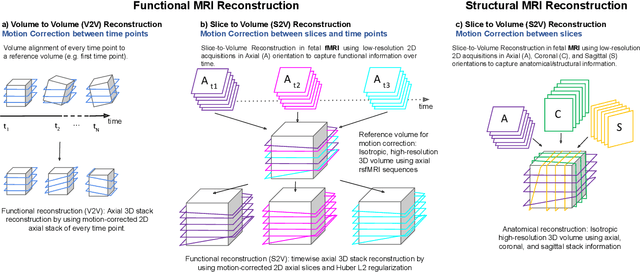
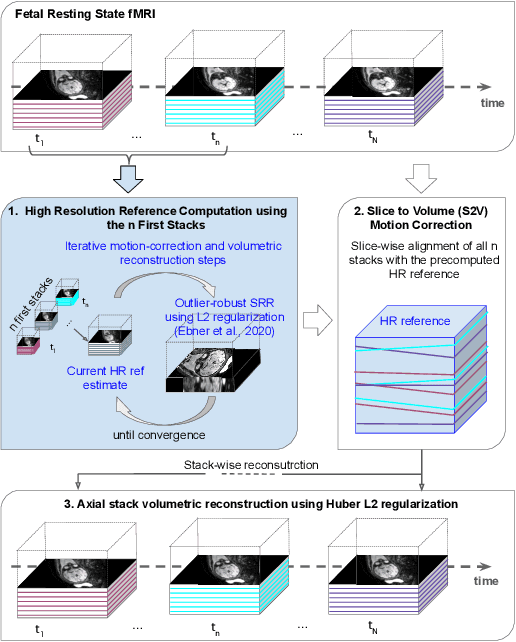
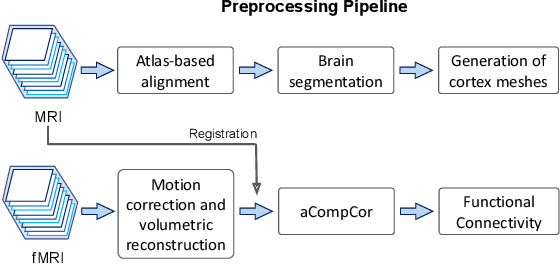
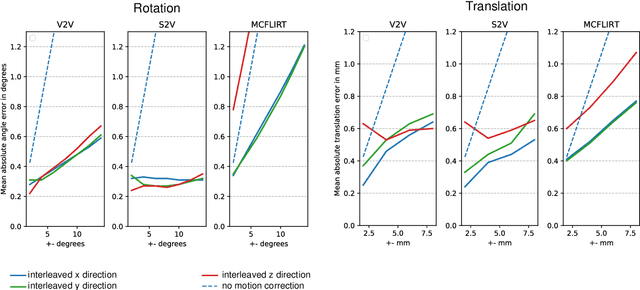
Abstract:Motion correction is an essential preprocessing step in functional Magnetic Resonance Imaging (fMRI) of the fetal brain with the aim to remove artifacts caused by fetal movement and maternal breathing and consequently to suppress erroneous signal correlations. Current motion correction approaches for fetal fMRI choose a single 3D volume from a specific acquisition timepoint with least motion artefacts as reference volume, and perform interpolation for the reconstruction of the motion corrected time series. The results can suffer, if no low-motion frame is available, and if reconstruction does not exploit any assumptions about the continuity of the fMRI signal. Here, we propose a novel framework, which estimates a high-resolution reference volume by using outlier-robust motion correction, and by utilizing Huber L2 regularization for intra-stack volumetric reconstruction of the motion-corrected fetal brain fMRI. We performed an extensive parameter study to investigate the effectiveness of motion estimation and present in this work benchmark metrics to quantify the effect of motion correction and regularised volumetric reconstruction approaches on functional connectivity computations. We demonstrate the proposed framework's ability to improve functional connectivity estimates, reproducibility and signal interpretability, which is clinically highly desirable for the establishment of prognostic noninvasive imaging biomarkers. The motion correction and volumetric reconstruction framework is made available as an open-source package of NiftyMIC.
4D iterative reconstruction of brain fMRI in the moving fetus
Nov 22, 2021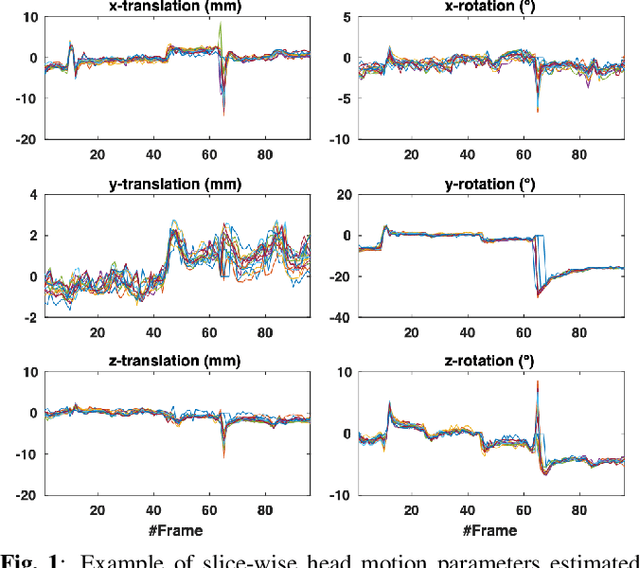
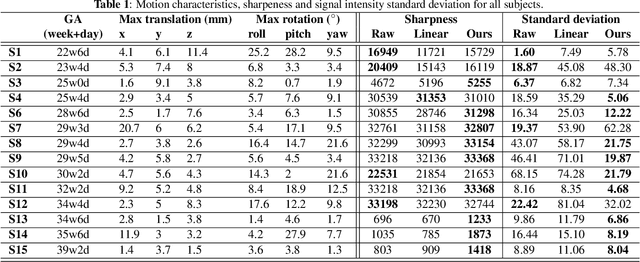
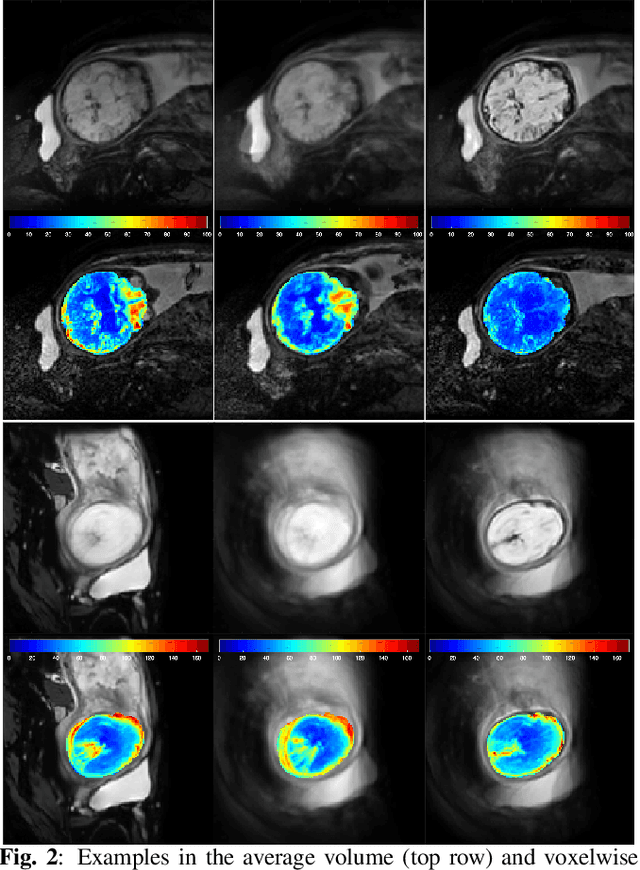
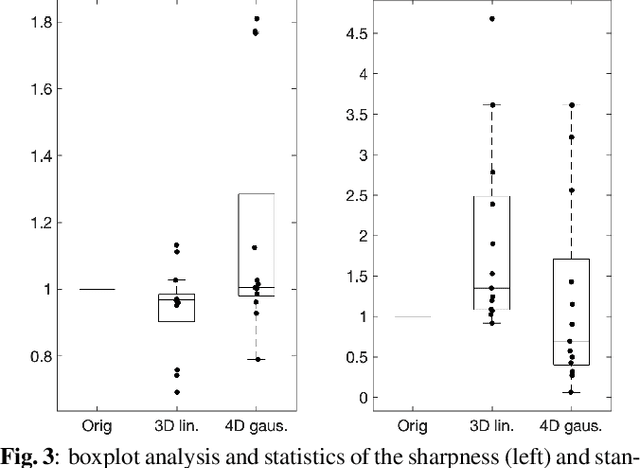
Abstract:Resting-state functional Magnetic Resonance Imaging (fMRI) is a powerful imaging technique for studying functional development of the brain in utero. However, unpredictable and excessive movement of fetuses has limited clinical application since it causes substantial signal fluctuations which can systematically alter observed patterns of functional connectivity. Previous studies have focused on the accurate estimation of the motion parameters in case of large fetal head movement and used a 3D single step interpolation approach at each timepoint to recover motion-free fMRI images. This does not guarantee that the reconstructed image corresponds to the minimum error representation of fMRI time series given the acquired data. Here, we propose a novel technique based on four dimensional iterative reconstruction of the scattered slices acquired during fetal fMRI. The accuracy of the proposed method was quantitatively evaluated on a group of real clinical fMRI fetuses. The results indicate improvements of reconstruction quality compared to the conventional 3D interpolation approach.
Unsupervised deep clustering for predictive texture pattern discovery in medical images
Jan 31, 2020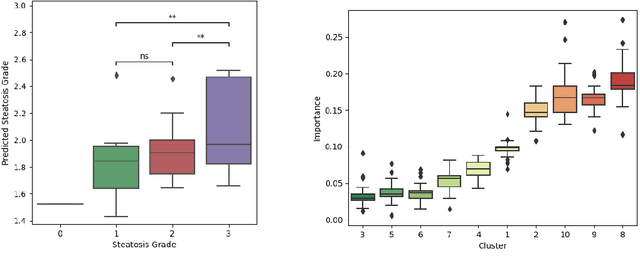
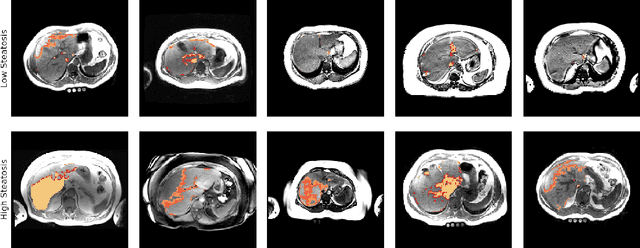
Abstract:Predictive marker patterns in imaging data are a means to quantify disease and progression, but their identification is challenging, if the underlying biology is poorly understood. Here, we present a method to identify predictive texture patterns in medical images in an unsupervised way. Based on deep clustering networks, we simultaneously encode and cluster medical image patches in a low-dimensional latent space. The resulting clusters serve as features for disease staging, linking them to the underlying disease. We evaluate the method on 70 T1-weighted magnetic resonance images of patients with different stages of liver steatosis. The deep clustering approach is able to find predictive clusters with a stable ranking, differentiating between low and high steatosis with an F1-Score of 0.78.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge