Zhi Hong
MASPOB: Bandit-Based Prompt Optimization for Multi-Agent Systems with Graph Neural Networks
Mar 03, 2026Abstract:Large Language Models (LLMs) have achieved great success in many real-world applications, especially the one serving as the cognitive backbone of Multi-Agent Systems (MAS) to orchestrate complex workflows in practice. Since many deployment scenarios preclude MAS workflow modifications and its performance is highly sensitive to the input prompts, prompt optimization emerges as a more natural approach to improve its performance. However, real-world prompt optimization for MAS is impeded by three key challenges: (1) the need of sample efficiency due to prohibitive evaluation costs, (2) topology-induced coupling among prompts, and (3) the combinatorial explosion of the search space. To address these challenges, we introduce MASPOB (Multi-Agent System Prompt Optimization via Bandits), a novel sample-efficient framework based on bandits. By leveraging Upper Confidence Bound (UCB) to quantify uncertainty, the bandit framework balances exploration and exploitation, maximizing gains within a strictly limited budget. To handle topology-induced coupling, MASPOB integrates Graph Neural Networks (GNNs) to capture structural priors, learning topology-aware representations of prompt semantics. Furthermore, it employs coordinate ascent to decompose the optimization into univariate sub-problems, reducing search complexity from exponential to linear. Extensive experiments across diverse benchmarks demonstrate that MASPOB achieves state-of-the-art performance, consistently outperforming existing baselines.
Workflow-R1: Group Sub-sequence Policy Optimization for Multi-turn Workflow Construction
Feb 01, 2026Abstract:The rapid evolution of agentic workflows has demonstrated strong performance of LLM-based agents in addressing complex reasoning tasks. However, existing workflow optimization methods typically formulate workflow synthesis as a static, one-shot code-centric generation problem. This paradigm imposes excessive constraints on the model's coding capabilities and restricts the flexibility required for dynamic problem-solving. In this paper, we present Workflow-R1, a framework that reformulates workflow construction as a multi-turn, natural language-based sequential decision-making process. To resolve the optimization granularity mismatch inherent in such multi-turn interactions, we introduce Group Sub-sequence Policy Optimization (GSsPO). While explicitly tailored to align with the interleaved Think-Action dynamics of agentic reasoning, GSsPO fundamentally functions as a structure-aware RL algorithm generalizable to a broad class of multi-turn agentic sequential decision-making tasks. By recalibrating the optimization unit to the composite sub-sequence, specifically the atomic Think-Action cycle, it aligns gradient updates with the semantic boundaries of these interactions, ensuring robust learning in complex multi-turn reasoning tasks. Through extensive experiments on multiple QA benchmarks, Workflow-R1 outperforms competitive baselines, validating GSsPO as a generalized solution for sequential reasoning and establishing Workflow-R1 as a promising new paradigm for automated workflow optimization.
LLM-based Robot Task Planning with Exceptional Handling for General Purpose Service Robots
May 24, 2024Abstract:The development of a general purpose service robot for daily life necessitates the robot's ability to deploy a myriad of fundamental behaviors judiciously. Recent advancements in training Large Language Models (LLMs) can be used to generate action sequences directly, given an instruction in natural language with no additional domain information. However, while the outputs of LLMs are semantically correct, the generated task plans may not accurately map to acceptable actions and might encompass various linguistic ambiguities. LLM hallucinations pose another challenge for robot task planning, which results in content that is inconsistent with real-world facts or user inputs. In this paper, we propose a task planning method based on a constrained LLM prompt scheme, which can generate an executable action sequence from a command. An exceptional handling module is further proposed to deal with LLM hallucinations problem. This module can ensure the LLM-generated results are admissible in the current environment. We evaluate our method on the commands generated by the RoboCup@Home Command Generator, observing that the robot demonstrates exceptional performance in both comprehending instructions and executing tasks.
Combining Language and Graph Models for Semi-structured Information Extraction on the Web
Feb 21, 2024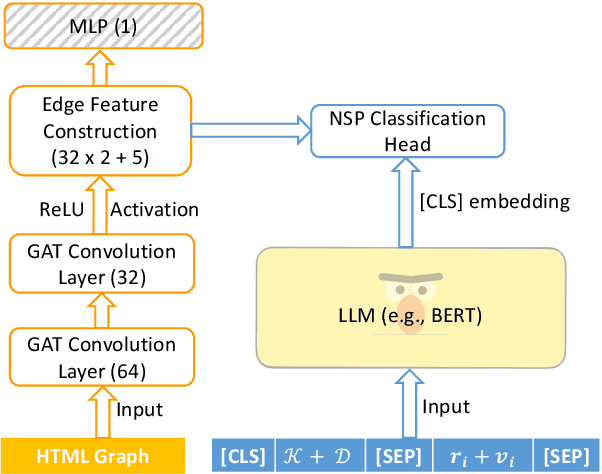

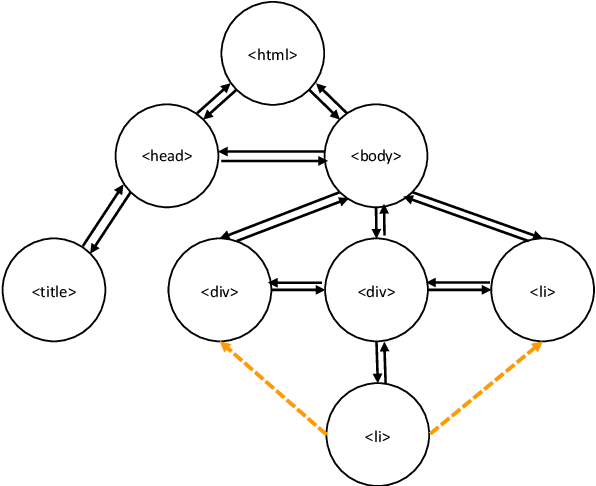

Abstract:Relation extraction is an efficient way of mining the extraordinary wealth of human knowledge on the Web. Existing methods rely on domain-specific training data or produce noisy outputs. We focus here on extracting targeted relations from semi-structured web pages given only a short description of the relation. We present GraphScholarBERT, an open-domain information extraction method based on a joint graph and language model structure. GraphScholarBERT can generalize to previously unseen domains without additional data or training and produces only clean extraction results matched to the search keyword. Experiments show that GraphScholarBERT can improve extraction F1 scores by as much as 34.8\% compared to previous work in a zero-shot domain and zero-shot website setting.
14 Examples of How LLMs Can Transform Materials Science and Chemistry: A Reflection on a Large Language Model Hackathon
Jun 13, 2023



Abstract:Chemistry and materials science are complex. Recently, there have been great successes in addressing this complexity using data-driven or computational techniques. Yet, the necessity of input structured in very specific forms and the fact that there is an ever-growing number of tools creates usability and accessibility challenges. Coupled with the reality that much data in these disciplines is unstructured, the effectiveness of these tools is limited. Motivated by recent works that indicated that large language models (LLMs) might help address some of these issues, we organized a hackathon event on the applications of LLMs in chemistry, materials science, and beyond. This article chronicles the projects built as part of this hackathon. Participants employed LLMs for various applications, including predicting properties of molecules and materials, designing novel interfaces for tools, extracting knowledge from unstructured data, and developing new educational applications. The diverse topics and the fact that working prototypes could be generated in less than two days highlight that LLMs will profoundly impact the future of our fields. The rich collection of ideas and projects also indicates that the applications of LLMs are not limited to materials science and chemistry but offer potential benefits to a wide range of scientific disciplines.
ScholarBERT: Bigger is Not Always Better
May 23, 2022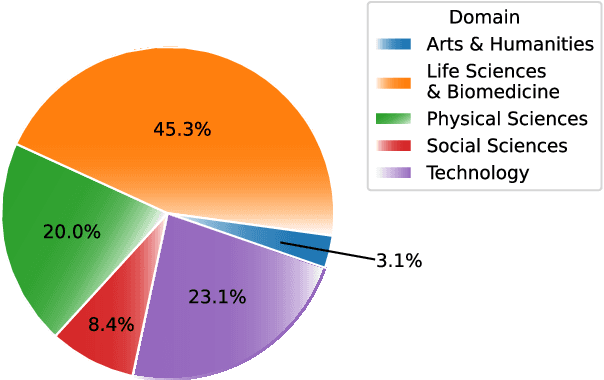
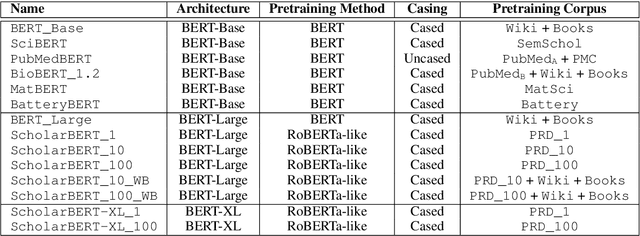

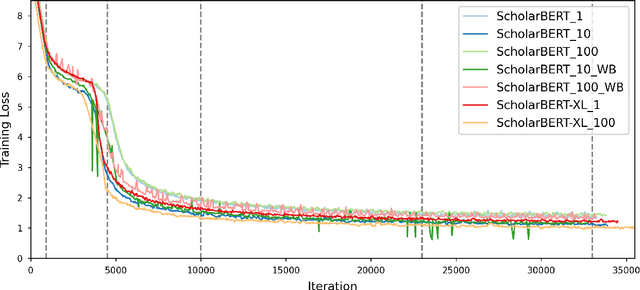
Abstract:Transformer-based masked language models trained on general corpora, such as BERT and RoBERTa, have shown impressive performance on various downstream tasks. Increasingly, researchers are "finetuning" these models to improve performance on domain-specific tasks. Here, we report a broad study in which we applied 14 transformer-based models to 11 scientific tasks in order to evaluate how downstream performance is affected by changes along various dimensions (e.g., training data, model size, pretraining time, finetuning length). In this process, we created the largest and most diverse scientific language model to date, ScholarBERT, by training a 770M-parameter BERT model on an 221B token scientific literature dataset spanning many disciplines. Counterintuitively, our evaluation of the 14 BERT-based models (seven versions of ScholarBERT, five science-specific large language models from the literature, BERT-Base, and BERT-Large) reveals little difference in performance across the 11 science-focused tasks, despite major differences in model size and training data. We argue that our results establish an upper bound for the performance achievable with BERT-based architectures on tasks from the scientific domain.
AI- and HPC-enabled Lead Generation for SARS-CoV-2: Models and Processes to Extract Druglike Molecules Contained in Natural Language Text
Jan 12, 2021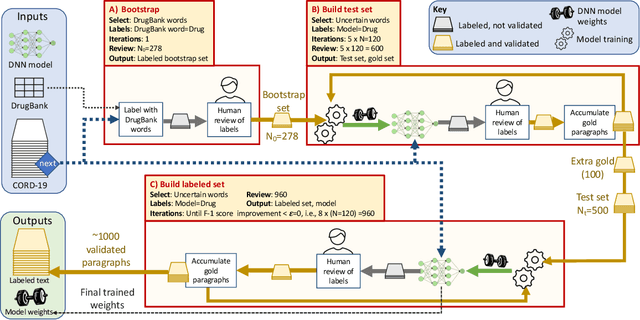
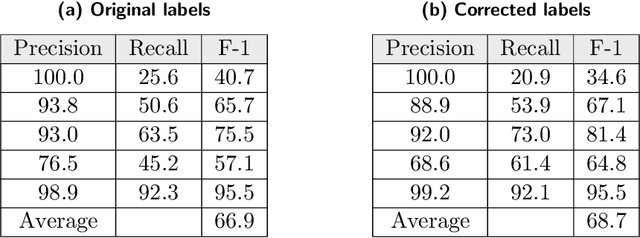
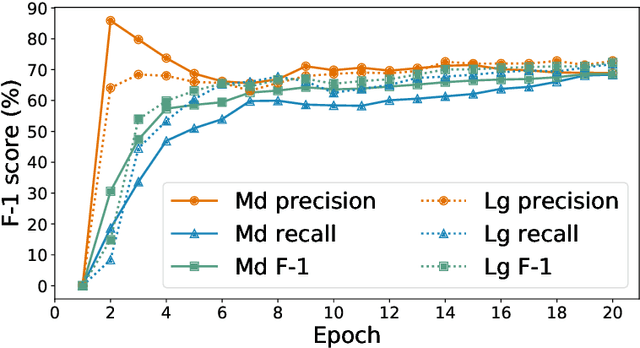
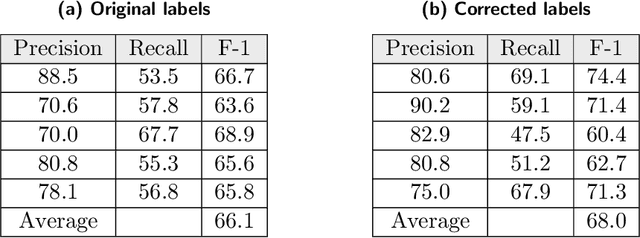
Abstract:Researchers worldwide are seeking to repurpose existing drugs or discover new drugs to counter the disease caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). A promising source of candidates for such studies is molecules that have been reported in the scientific literature to be drug-like in the context of coronavirus research. We report here on a project that leverages both human and artificial intelligence to detect references to drug-like molecules in free text. We engage non-expert humans to create a corpus of labeled text, use this labeled corpus to train a named entity recognition model, and employ the trained model to extract 10912 drug-like molecules from the COVID-19 Open Research Dataset Challenge (CORD-19) corpus of 198875 papers. Performance analyses show that our automated extraction model can achieve performance on par with that of non-expert humans.
Targeting SARS-CoV-2 with AI- and HPC-enabled Lead Generation: A First Data Release
May 28, 2020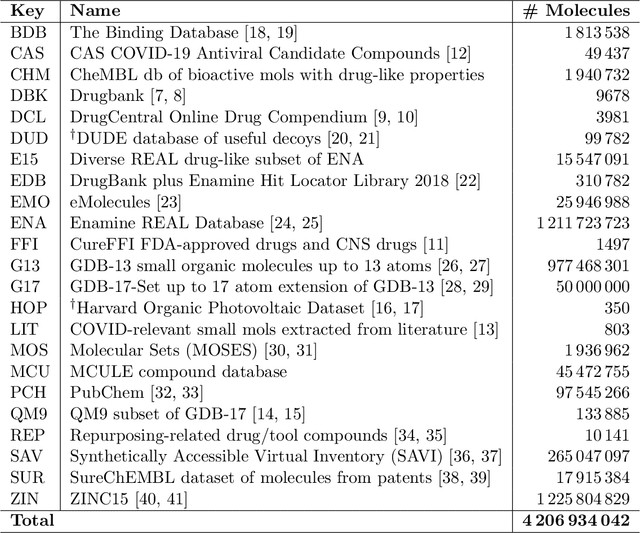
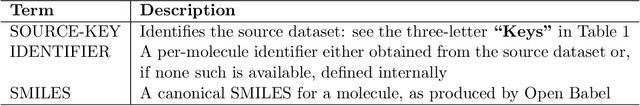
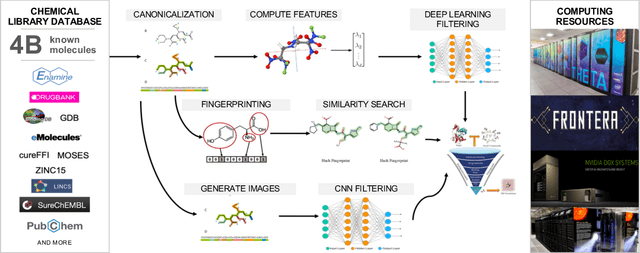

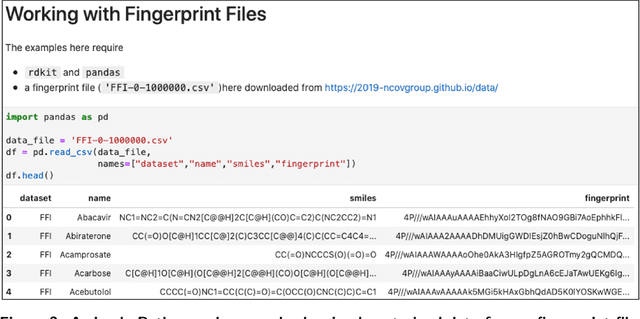
Abstract:Researchers across the globe are seeking to rapidly repurpose existing drugs or discover new drugs to counter the the novel coronavirus disease (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). One promising approach is to train machine learning (ML) and artificial intelligence (AI) tools to screen large numbers of small molecules. As a contribution to that effort, we are aggregating numerous small molecules from a variety of sources, using high-performance computing (HPC) to computer diverse properties of those molecules, using the computed properties to train ML/AI models, and then using the resulting models for screening. In this first data release, we make available 23 datasets collected from community sources representing over 4.2 B molecules enriched with pre-computed: 1) molecular fingerprints to aid similarity searches, 2) 2D images of molecules to enable exploration and application of image-based deep learning methods, and 3) 2D and 3D molecular descriptors to speed development of machine learning models. This data release encompasses structural information on the 4.2 B molecules and 60 TB of pre-computed data. Future releases will expand the data to include more detailed molecular simulations, computed models, and other products.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge