Yaoqin Xie
Visualizing the Invisible: Enhancing Radiologist Performance in Breast Mammography via Task-Driven Chromatic Encoding
Feb 07, 2026Abstract:Purpose:Mammography screening is less sensitive in dense breasts, where tissue overlap and subtle findings increase perceptual difficulty. We present MammoColor, an end-to-end framework with a Task-Driven Chromatic Encoding (TDCE) module that converts single-channel mammograms into TDCE-encoded views for visual augmentation. Materials and Methods:MammoColor couples a lightweight TDCE module with a BI-RADS triage classifier and was trained end-to-end on VinDr-Mammo. Performance was evaluated on an internal test set, two public datasets (CBIS-DDSM and INBreast), and three external clinical cohorts. We also conducted a multi-reader, multi-case (MRMC) observer study with a washout period, comparing (1) grayscale-only, (2) TDCE-only, and (3) side-by-side grayscale+TDCE. Results:On VinDr-Mammo, MammoColor improved AUC from 0.7669 to 0.8461 (P=0.004). Gains were larger in dense breasts (AUC 0.749 to 0.835). In the MRMC study, TDCE-encoded images improved specificity (0.90 to 0.96; P=0.052) with comparable sensitivity. Conclusion:TDCE provides a task-optimized chromatic representation that may improve perceptual salience and reduce false-positive recalls in mammography triage.
Three-Dimensional Medical Image Fusion with Deformable Cross-Attention
Oct 10, 2023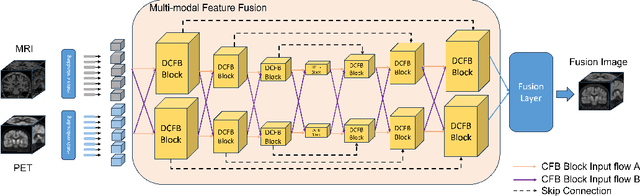
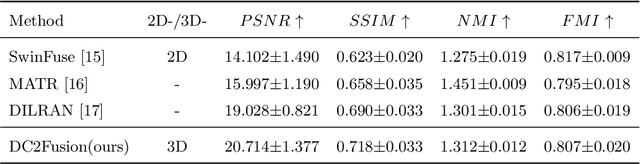
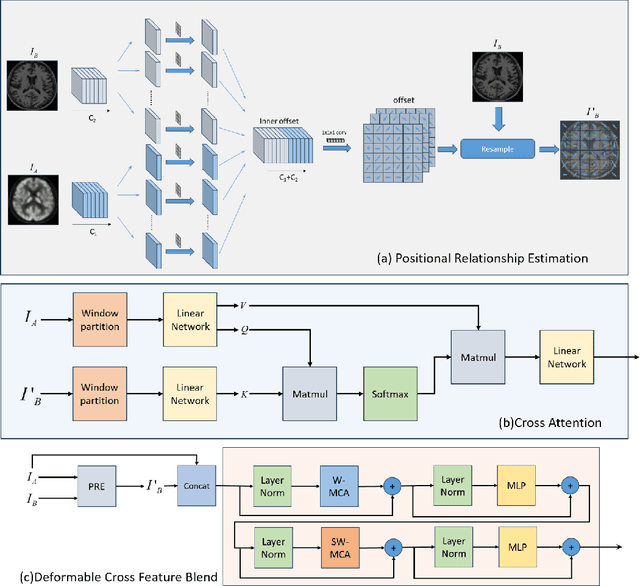
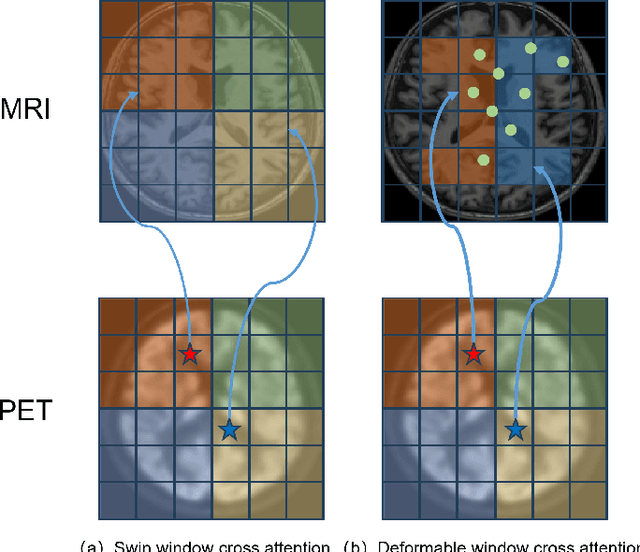
Abstract:Multimodal medical image fusion plays an instrumental role in several areas of medical image processing, particularly in disease recognition and tumor detection. Traditional fusion methods tend to process each modality independently before combining the features and reconstructing the fusion image. However, this approach often neglects the fundamental commonalities and disparities between multimodal information. Furthermore, the prevailing methodologies are largely confined to fusing two-dimensional (2D) medical image slices, leading to a lack of contextual supervision in the fusion images and subsequently, a decreased information yield for physicians relative to three-dimensional (3D) images. In this study, we introduce an innovative unsupervised feature mutual learning fusion network designed to rectify these limitations. Our approach incorporates a Deformable Cross Feature Blend (DCFB) module that facilitates the dual modalities in discerning their respective similarities and differences. We have applied our model to the fusion of 3D MRI and PET images obtained from 660 patients in the Alzheimer's Disease Neuroimaging Initiative (ADNI) dataset. Through the application of the DCFB module, our network generates high-quality MRI-PET fusion images. Experimental results demonstrate that our method surpasses traditional 2D image fusion methods in performance metrics such as Peak Signal to Noise Ratio (PSNR) and Structural Similarity Index Measure (SSIM). Importantly, the capacity of our method to fuse 3D images enhances the information available to physicians and researchers, thus marking a significant step forward in the field. The code will soon be available online.
QUIZ: An Arbitrary Volumetric Point Matching Method for Medical Image Registration
Sep 30, 2023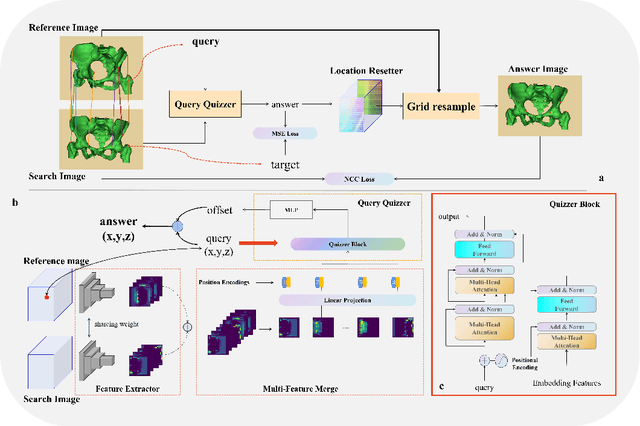
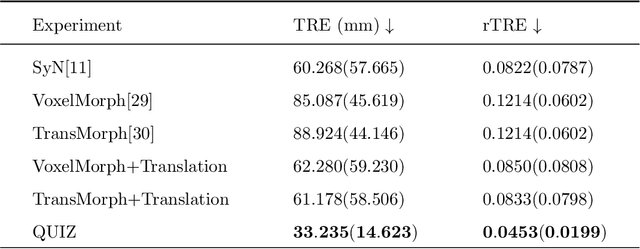
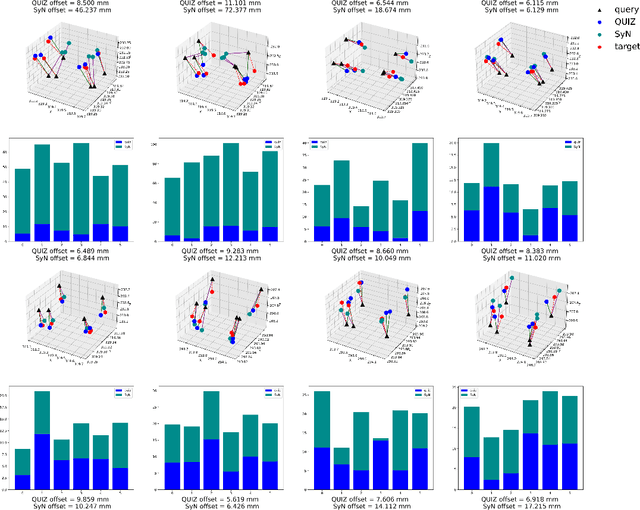
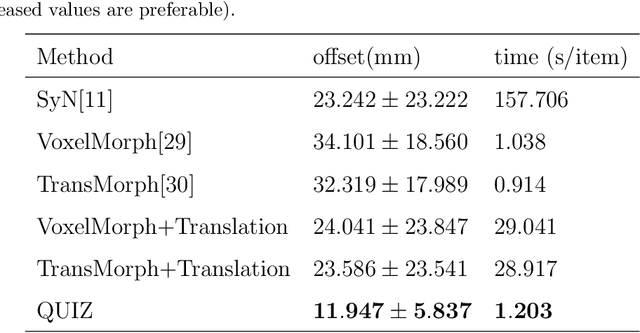
Abstract:Rigid pre-registration involving local-global matching or other large deformation scenarios is crucial. Current popular methods rely on unsupervised learning based on grayscale similarity, but under circumstances where different poses lead to varying tissue structures, or where image quality is poor, these methods tend to exhibit instability and inaccuracies. In this study, we propose a novel method for medical image registration based on arbitrary voxel point of interest matching, called query point quizzer (QUIZ). QUIZ focuses on the correspondence between local-global matching points, specifically employing CNN for feature extraction and utilizing the Transformer architecture for global point matching queries, followed by applying average displacement for local image rigid transformation. We have validated this approach on a large deformation dataset of cervical cancer patients, with results indicating substantially smaller deviations compared to state-of-the-art methods. Remarkably, even for cross-modality subjects, it achieves results surpassing the current state-of-the-art.
Unsupervised CT Metal Artifact Reduction by Plugging Diffusion Priors in Dual Domains
Aug 31, 2023



Abstract:During the process of computed tomography (CT), metallic implants often cause disruptive artifacts in the reconstructed images, impeding accurate diagnosis. Several supervised deep learning-based approaches have been proposed for reducing metal artifacts (MAR). However, these methods heavily rely on training with simulated data, as obtaining paired metal artifact CT and clean CT data in clinical settings is challenging. This limitation can lead to decreased performance when applying these methods in clinical practice. Existing unsupervised MAR methods, whether based on learning or not, typically operate within a single domain, either in the image domain or the sinogram domain. In this paper, we propose an unsupervised MAR method based on the diffusion model, a generative model with a high capacity to represent data distributions. Specifically, we first train a diffusion model using CT images without metal artifacts. Subsequently, we iteratively utilize the priors embedded within the pre-trained diffusion model in both the sinogram and image domains to restore the degraded portions caused by metal artifacts. This dual-domain processing empowers our approach to outperform existing unsupervised MAR methods, including another MAR method based on the diffusion model, which we have qualitatively and quantitatively validated using synthetic datasets. Moreover, our method demonstrates superior visual results compared to both supervised and unsupervised methods on clinical datasets.
XTransCT: Ultra-Fast Volumetric CT Reconstruction using Two Orthogonal X-Ray Projections via a Transformer Network
May 31, 2023Abstract:Computed tomography (CT) scans offer a detailed, three-dimensional representation of patients' internal organs. However, conventional CT reconstruction techniques necessitate acquiring hundreds or thousands of x-ray projections through a complete rotational scan of the body, making navigation or positioning during surgery infeasible. In image-guided radiation therapy, a method that reconstructs ultra-sparse X-ray projections into CT images, we can exploit the substantially reduced radiation dose and minimize equipment burden for localization and navigation. In this study, we introduce a novel Transformer architecture, termed XTransCT, devised to facilitate real-time reconstruction of CT images from two-dimensional X-ray images. We assess our approach regarding image quality and structural reliability using a dataset of fifty patients, supplied by a hospital, as well as the larger public dataset LIDC-IDRI, which encompasses thousands of patients. Additionally, we validated our algorithm's generalizability on the LNDb dataset. Our findings indicate that our algorithm surpasses other methods in image quality, structural precision, and generalizability. Moreover, in comparison to previous 3D convolution-based approaches, we note a substantial speed increase of approximately 300 $\%$, achieving 44 ms per 3D image reconstruction. To guarantee the replicability of our results, we have made our code publicly available.
A Diffusion Probabilistic Prior for Low-Dose CT Image Denoising
May 25, 2023Abstract:Low-dose computed tomography (CT) image denoising is crucial in medical image computing. Recent years have been remarkable improvement in deep learning-based methods for this task. However, training deep denoising neural networks requires low-dose and normal-dose CT image pairs, which are difficult to obtain in the clinic settings. To address this challenge, we propose a novel fully unsupervised method for low-dose CT image denoising, which is based on denoising diffusion probabilistic model -- a powerful generative model. First, we train an unconditional denoising diffusion probabilistic model capable of generating high-quality normal-dose CT images from random noise. Subsequently, the probabilistic priors of the pre-trained diffusion model are incorporated into a Maximum A Posteriori (MAP) estimation framework for iteratively solving the image denoising problem. Our method ensures the diffusion model produces high-quality normal-dose CT images while keeping the image content consistent with the input low-dose CT images. We evaluate our method on a widely used low-dose CT image denoising benchmark, and it outperforms several supervised low-dose CT image denoising methods in terms of both quantitative and visual performance.
A Hybrid Deep Feature-Based Deformable Image Registration Method for Pathological Images
Aug 17, 2022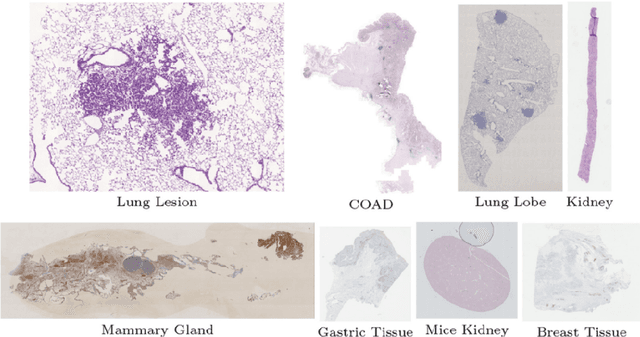
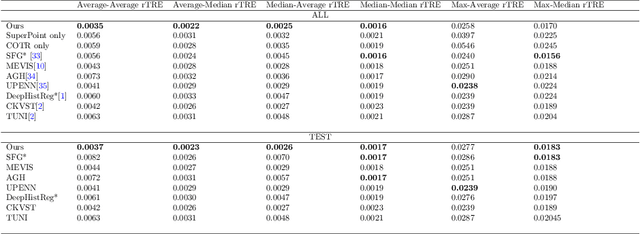
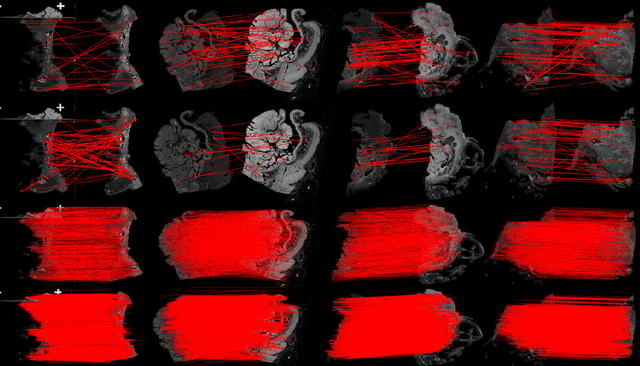
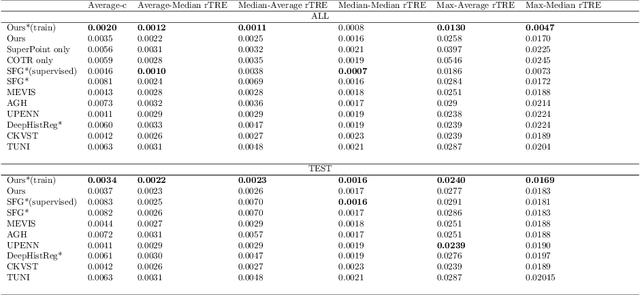
Abstract:Pathologists need to combine information from differently stained pathological slices to obtain accurate diagnostic results. Deformable image registration is a necessary technique for fusing multi-modal pathological slices. This paper proposes a hybrid deep feature-based deformable image registration framework for stained pathological samples. We first extract dense feature points and perform points matching by two deep learning feature networks. Then, to further reduce false matches, an outlier detection method combining the isolation forest statistical model and the local affine correction model is proposed. Finally, the interpolation method generates the DVF for pathology image registration based on the above matching points. We evaluate our method on the dataset of the Non-rigid Histology Image Registration (ANHIR) challenge, which is co-organized with the IEEE ISBI 2019 conference. Our technique outperforms the traditional approaches by 17% with the Average-Average registration target error (rTRE) reaching 0.0034. The proposed method achieved state-of-the-art performance and ranking it 1 in evaluating the test dataset. The proposed hybrid deep feature-based registration method can potentially become a reliable method for pathology image registration.
Elastic Net based Feature Ranking and Selection
Dec 30, 2020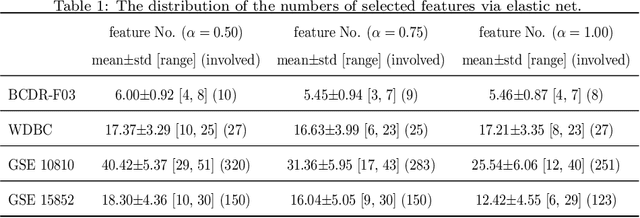
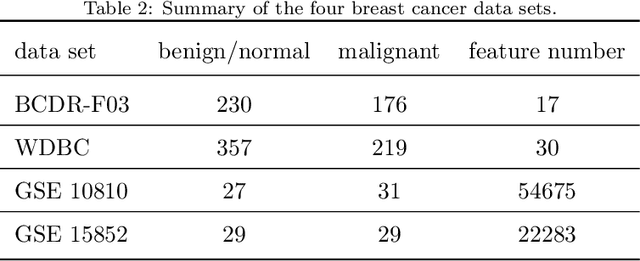
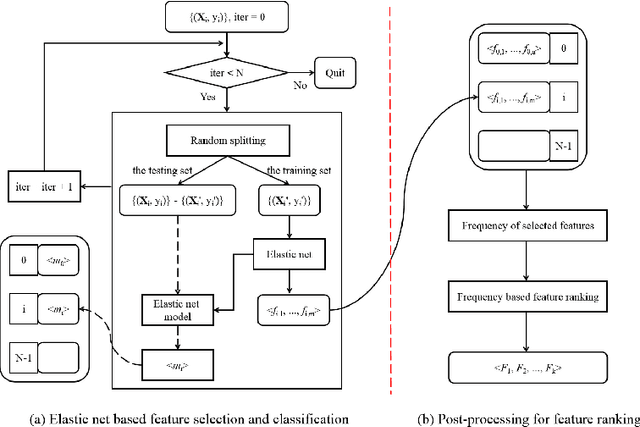
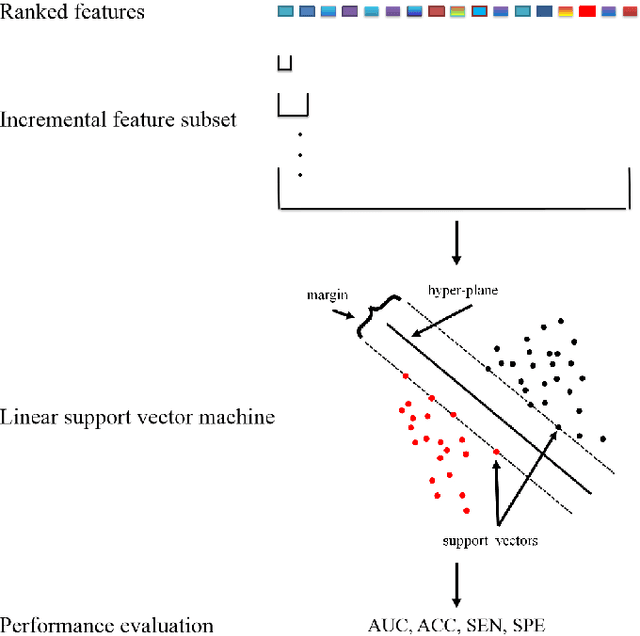

Abstract:Feature selection is important in data representation and intelligent diagnosis. Elastic net is one of the most widely used feature selectors. However, the features selected are dependant on the training data, and their weights dedicated for regularized regression are irrelevant to their importance if used for feature ranking, that degrades the model interpretability and extension. In this study, an intuitive idea is put at the end of multiple times of data splitting and elastic net based feature selection. It concerns the frequency of selected features and uses the frequency as an indicator of feature importance. After features are sorted according to their frequency, linear support vector machine performs the classification in an incremental manner. At last, a compact subset of discriminative features is selected by comparing the prediction performance. Experimental results on breast cancer data sets (BCDR-F03, WDBC, GSE 10810, and GSE 15852) suggest that the proposed framework achieves competitive or superior performance to elastic net and with consistent selection of fewer features. How to further enhance its consistency on high-dimension small-sample-size data sets should be paid more attention in our future work. The proposed framework is accessible online (https://github.com/NicoYuCN/elasticnetFR).
A Matlab Toolbox for Feature Importance Ranking
Mar 10, 2020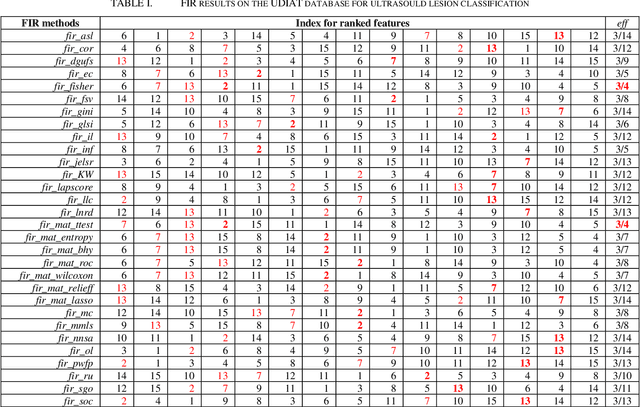
Abstract:More attention is being paid for feature importance ranking (FIR), in particular when thousands of features can be extracted for intelligent diagnosis and personalized medicine. A large number of FIR approaches have been proposed, while few are integrated for comparison and real-life applications. In this study, a matlab toolbox is presented and a total of 30 algorithms are collected. Moreover, the toolbox is evaluated on a database of 163 ultrasound images. To each breast mass lesion, 15 features are extracted. To figure out the optimal subset of features for classification, all combinations of features are tested and linear support vector machine is used for the malignancy prediction of lesions annotated in ultrasound images. At last, the effectiveness of FIR is analyzed according to performance comparison. The toolbox is online (https://github.com/NicoYuCN/matFIR). In our future work, more FIR methods, feature selection methods and machine learning classifiers will be integrated.
Performance Evaluation of Edge-Directed Interpolation Methods for Images
Mar 26, 2013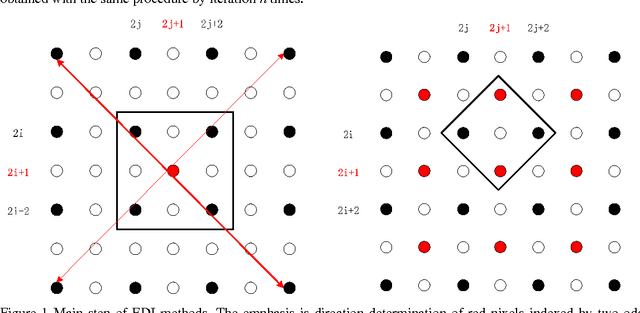
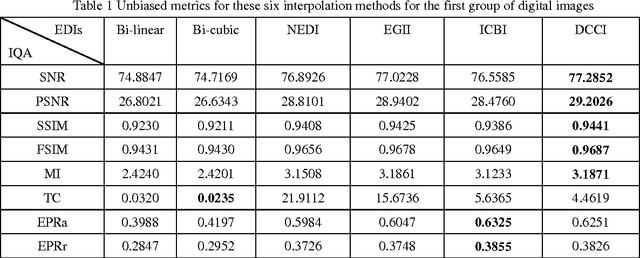
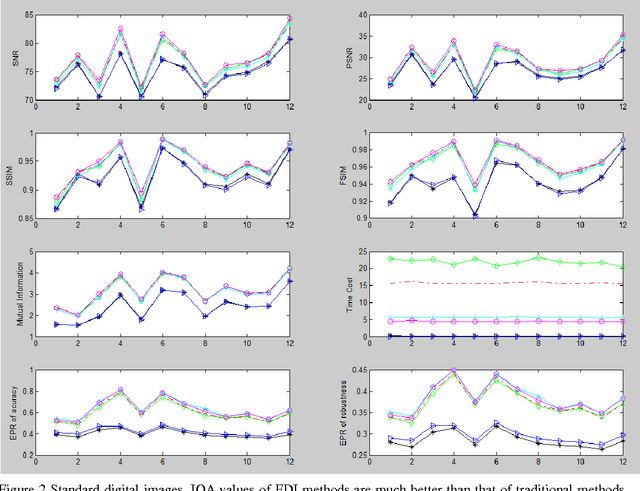
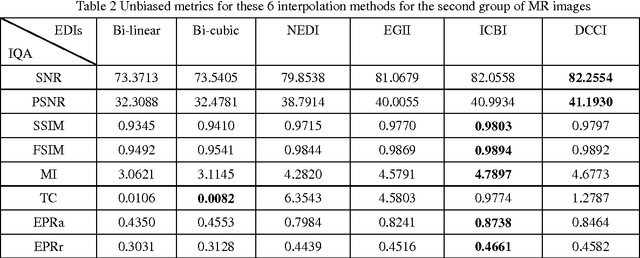
Abstract:Many interpolation methods have been developed for high visual quality, but fail for inability to preserve image structures. Edges carry heavy structural information for detection, determination and classification. Edge-adaptive interpolation approaches become a center of focus. In this paper, performance of four edge-directed interpolation methods comparing with two traditional methods is evaluated on two groups of images. These methods include new edge-directed interpolation (NEDI), edge-guided image interpolation (EGII), iterative curvature-based interpolation (ICBI), directional cubic convolution interpolation (DCCI) and two traditional approaches, bi-linear and bi-cubic. Meanwhile, no parameters are mentioned to measure edge-preserving ability of edge-adaptive interpolation approaches and we proposed two. One evaluates accuracy and the other measures robustness of edge-preservation ability. Performance evaluation is based on six parameters. Objective assessment and visual analysis are illustrated and conclusions are drawn from theoretical backgrounds and practical results.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge