Purang Abolmaesumi
Point Tracking as a Temporal Cue for Robust Myocardial Segmentation in Echocardiography Videos
Jan 14, 2026Abstract:Purpose: Myocardium segmentation in echocardiography videos is a challenging task due to low contrast, noise, and anatomical variability. Traditional deep learning models either process frames independently, ignoring temporal information, or rely on memory-based feature propagation, which accumulates error over time. Methods: We propose Point-Seg, a transformer-based segmentation framework that integrates point tracking as a temporal cue to ensure stable and consistent segmentation of myocardium across frames. Our method leverages a point-tracking module trained on a synthetic echocardiography dataset to track key anatomical landmarks across video sequences. These tracked trajectories provide an explicit motion-aware signal that guides segmentation, reducing drift and eliminating the need for memory-based feature accumulation. Additionally, we incorporate a temporal smoothing loss to further enhance temporal consistency across frames. Results: We evaluate our approach on both public and private echocardiography datasets. Experimental results demonstrate that Point-Seg has statistically similar accuracy in terms of Dice to state-of-the-art segmentation models in high quality echo data, while it achieves better segmentation accuracy in lower quality echo with improved temporal stability. Furthermore, Point-Seg has the key advantage of pixel-level myocardium motion information as opposed to other segmentation methods. Such information is essential in the computation of other downstream tasks such as myocardial strain measurement and regional wall motion abnormality detection. Conclusion: Point-Seg demonstrates that point tracking can serve as an effective temporal cue for consistent video segmentation, offering a reliable and generalizable approach for myocardium segmentation in echocardiography videos. The code is available at https://github.com/DeepRCL/PointSeg.
A Dataset and Benchmarks for Atrial Fibrillation Detection from Electrocardiograms of Intensive Care Unit Patients
Dec 19, 2025Abstract:Objective: Atrial fibrillation (AF) is the most common cardiac arrhythmia experienced by intensive care unit (ICU) patients and can cause adverse health effects. In this study, we publish a labelled ICU dataset and benchmarks for AF detection. Methods: We compared machine learning models across three data-driven artificial intelligence (AI) approaches: feature-based classifiers, deep learning (DL), and ECG foundation models (FMs). This comparison addresses a critical gap in the literature and aims to pinpoint which AI approach is best for accurate AF detection. Electrocardiograms (ECGs) from a Canadian ICU and the 2021 PhysioNet/Computing in Cardiology Challenge were used to conduct the experiments. Multiple training configurations were tested, ranging from zero-shot inference to transfer learning. Results: On average and across both datasets, ECG FMs performed best, followed by DL, then feature-based classifiers. The model that achieved the top F1 score on our ICU test set was ECG-FM through a transfer learning strategy (F1=0.89). Conclusion: This study demonstrates promising potential for using AI to build an automatic patient monitoring system. Significance: By publishing our labelled ICU dataset (LinkToBeAdded) and performance benchmarks, this work enables the research community to continue advancing the state-of-the-art in AF detection in the ICU.
EchoAgent: Guideline-Centric Reasoning Agent for Echocardiography Measurement and Interpretation
Nov 17, 2025Abstract:Purpose: Echocardiographic interpretation requires video-level reasoning and guideline-based measurement analysis, which current deep learning models for cardiac ultrasound do not support. We present EchoAgent, a framework that enables structured, interpretable automation for this domain. Methods: EchoAgent orchestrates specialized vision tools under Large Language Model (LLM) control to perform temporal localization, spatial measurement, and clinical interpretation. A key contribution is a measurement-feasibility prediction model that determines whether anatomical structures are reliably measurable in each frame, enabling autonomous tool selection. We curated a benchmark of diverse, clinically validated video-query pairs for evaluation. Results: EchoAgent achieves accurate, interpretable results despite added complexity of spatiotemporal video analysis. Outputs are grounded in visual evidence and clinical guidelines, supporting transparency and traceability. Conclusion: This work demonstrates the feasibility of agentic, guideline-aligned reasoning for echocardiographic video analysis, enabled by task-specific tools and full video-level automation. EchoAgent sets a new direction for trustworthy AI in cardiac ultrasound.
ProstNFound+: A Prospective Study using Medical Foundation Models for Prostate Cancer Detection
Oct 30, 2025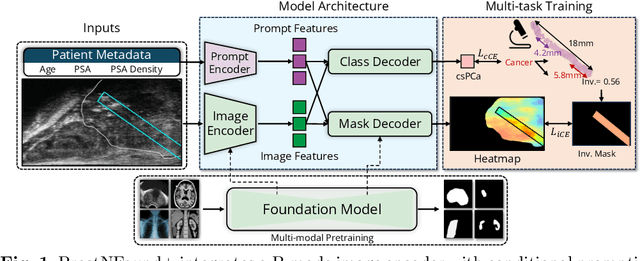
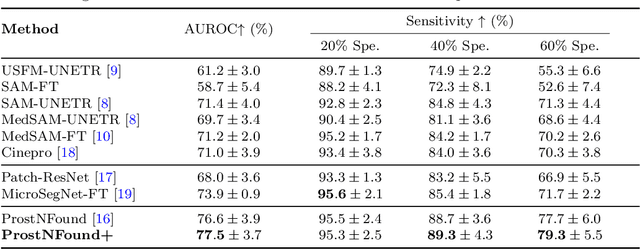
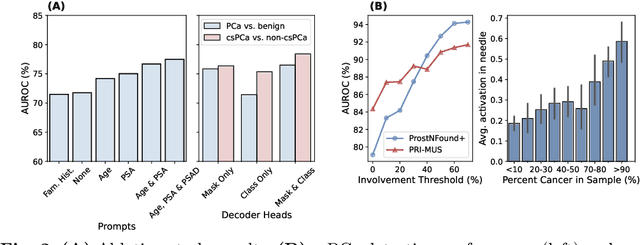
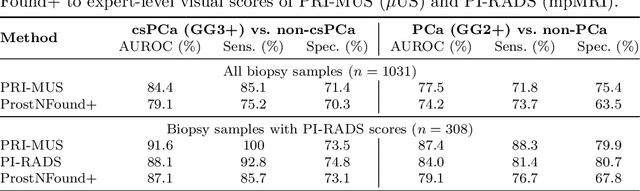
Abstract:Purpose: Medical foundation models (FMs) offer a path to build high-performance diagnostic systems. However, their application to prostate cancer (PCa) detection from micro-ultrasound ({\mu}US) remains untested in clinical settings. We present ProstNFound+, an adaptation of FMs for PCa detection from {\mu}US, along with its first prospective validation. Methods: ProstNFound+ incorporates a medical FM, adapter tuning, and a custom prompt encoder that embeds PCa-specific clinical biomarkers. The model generates a cancer heatmap and a risk score for clinically significant PCa. Following training on multi-center retrospective data, the model is prospectively evaluated on data acquired five years later from a new clinical site. Model predictions are benchmarked against standard clinical scoring protocols (PRI-MUS and PI-RADS). Results: ProstNFound+ shows strong generalization to the prospective data, with no performance degradation compared to retrospective evaluation. It aligns closely with clinical scores and produces interpretable heatmaps consistent with biopsy-confirmed lesions. Conclusion: The results highlight its potential for clinical deployment, offering a scalable and interpretable alternative to expert-driven protocols.
Domain Knowledge is Power: Leveraging Physiological Priors for Self Supervised Representation Learning in Electrocardiography
Sep 09, 2025Abstract:Objective: Electrocardiograms (ECGs) play a crucial role in diagnosing heart conditions; however, the effectiveness of artificial intelligence (AI)-based ECG analysis is often hindered by the limited availability of labeled data. Self-supervised learning (SSL) can address this by leveraging large-scale unlabeled data. We introduce PhysioCLR (Physiology-aware Contrastive Learning Representation for ECG), a physiology-aware contrastive learning framework that incorporates domain-specific priors to enhance the generalizability and clinical relevance of ECG-based arrhythmia classification. Methods: During pretraining, PhysioCLR learns to bring together embeddings of samples that share similar clinically relevant features while pushing apart those that are dissimilar. Unlike existing methods, our method integrates ECG physiological similarity cues into contrastive learning, promoting the learning of clinically meaningful representations. Additionally, we introduce ECG- specific augmentations that preserve the ECG category post augmentation and propose a hybrid loss function to further refine the quality of learned representations. Results: We evaluate PhysioCLR on two public ECG datasets, Chapman and Georgia, for multilabel ECG diagnoses, as well as a private ICU dataset labeled for binary classification. Across the Chapman, Georgia, and private cohorts, PhysioCLR boosts the mean AUROC by 12% relative to the strongest baseline, underscoring its robust cross-dataset generalization. Conclusion: By embedding physiological knowledge into contrastive learning, PhysioCLR enables the model to learn clinically meaningful and transferable ECG eatures. Significance: PhysioCLR demonstrates the potential of physiology-informed SSL to offer a promising path toward more effective and label-efficient ECG diagnostics.
ControlEchoSynth: Boosting Ejection Fraction Estimation Models via Controlled Video Diffusion
Aug 25, 2025



Abstract:Synthetic data generation represents a significant advancement in boosting the performance of machine learning (ML) models, particularly in fields where data acquisition is challenging, such as echocardiography. The acquisition and labeling of echocardiograms (echo) for heart assessment, crucial in point-of-care ultrasound (POCUS) settings, often encounter limitations due to the restricted number of echo views available, typically captured by operators with varying levels of experience. This study proposes a novel approach for enhancing clinical diagnosis accuracy by synthetically generating echo views. These views are conditioned on existing, real views of the heart, focusing specifically on the estimation of ejection fraction (EF), a critical parameter traditionally measured from biplane apical views. By integrating a conditional generative model, we demonstrate an improvement in EF estimation accuracy, providing a comparative analysis with traditional methods. Preliminary results indicate that our synthetic echoes, when used to augment existing datasets, not only enhance EF estimation but also show potential in advancing the development of more robust, accurate, and clinically relevant ML models. This approach is anticipated to catalyze further research in synthetic data applications, paving the way for innovative solutions in medical imaging diagnostics.
Diverse Prototypical Ensembles Improve Robustness to Subpopulation Shift
May 29, 2025Abstract:The subpopulationtion shift, characterized by a disparity in subpopulation distributibetween theween the training and target datasets, can significantly degrade the performance of machine learning models. Current solutions to subpopulation shift involve modifying empirical risk minimization with re-weighting strategies to improve generalization. This strategy relies on assumptions about the number and nature of subpopulations and annotations on group membership, which are unavailable for many real-world datasets. Instead, we propose using an ensemble of diverse classifiers to adaptively capture risk associated with subpopulations. Given a feature extractor network, we replace its standard linear classification layer with a mixture of prototypical classifiers, where each member is trained to classify the data while focusing on different features and samples from other members. In empirical evaluation on nine real-world datasets, covering diverse domains and kinds of subpopulation shift, our method of Diverse Prototypical Ensembles (DPEs) often outperforms the prior state-of-the-art in worst-group accuracy. The code is available at https://github.com/minhto2802/dpe4subpop
TRUSWorthy: Toward Clinically Applicable Deep Learning for Confident Detection of Prostate Cancer in Micro-Ultrasound
Feb 20, 2025Abstract:While deep learning methods have shown great promise in improving the effectiveness of prostate cancer (PCa) diagnosis by detecting suspicious lesions from trans-rectal ultrasound (TRUS), they must overcome multiple simultaneous challenges. There is high heterogeneity in tissue appearance, significant class imbalance in favor of benign examples, and scarcity in the number and quality of ground truth annotations available to train models. Failure to address even a single one of these problems can result in unacceptable clinical outcomes.We propose TRUSWorthy, a carefully designed, tuned, and integrated system for reliable PCa detection. Our pipeline integrates self-supervised learning, multiple-instance learning aggregation using transformers, random-undersampled boosting and ensembling: these address label scarcity, weak labels, class imbalance, and overconfidence, respectively. We train and rigorously evaluate our method using a large, multi-center dataset of micro-ultrasound data. Our method outperforms previous state-of-the-art deep learning methods in terms of accuracy and uncertainty calibration, with AUROC and balanced accuracy scores of 79.9% and 71.5%, respectively. On the top 20% of predictions with the highest confidence, we can achieve a balanced accuracy of up to 91%. The success of TRUSWorthy demonstrates the potential of integrated deep learning solutions to meet clinical needs in a highly challenging deployment setting, and is a significant step towards creating a trustworthy system for computer-assisted PCa diagnosis.
Cinepro: Robust Training of Foundation Models for Cancer Detection in Prostate Ultrasound Cineloops
Jan 21, 2025



Abstract:Prostate cancer (PCa) detection using deep learning (DL) models has shown potential for enhancing real-time guidance during biopsies. However, prostate ultrasound images lack pixel-level cancer annotations, introducing label noise. Current approaches often focus on limited regions of interest (ROIs), disregarding anatomical context necessary for accurate diagnosis. Foundation models can overcome this limitation by analyzing entire images to capture global spatial relationships; however, they still encounter challenges stemming from the weak labels associated with coarse pathology annotations in ultrasound data. We introduce Cinepro, a novel framework that strengthens foundation models' ability to localize PCa in ultrasound cineloops. Cinepro adapts robust training by integrating the proportion of cancer tissue reported by pathology in a biopsy core into its loss function to address label noise, providing a more nuanced supervision. Additionally, it leverages temporal data across multiple frames to apply robust augmentations, enhancing the model's ability to learn stable cancer-related features. Cinepro demonstrates superior performance on a multi-center prostate ultrasound dataset, achieving an AUROC of 77.1% and a balanced accuracy of 83.8%, surpassing current benchmarks. These findings underscore Cinepro's promise in advancing foundation models for weakly labeled ultrasound data.
Reliable Multi-View Learning with Conformal Prediction for Aortic Stenosis Classification in Echocardiography
Sep 15, 2024Abstract:The fundamental problem with ultrasound-guided diagnosis is that the acquired images are often 2-D cross-sections of a 3-D anatomy, potentially missing important anatomical details. This limitation leads to challenges in ultrasound echocardiography, such as poor visualization of heart valves or foreshortening of ventricles. Clinicians must interpret these images with inherent uncertainty, a nuance absent in machine learning's one-hot labels. We propose Re-Training for Uncertainty (RT4U), a data-centric method to introduce uncertainty to weakly informative inputs in the training set. This simple approach can be incorporated to existing state-of-the-art aortic stenosis classification methods to further improve their accuracy. When combined with conformal prediction techniques, RT4U can yield adaptively sized prediction sets which are guaranteed to contain the ground truth class to a high accuracy. We validate the effectiveness of RT4U on three diverse datasets: a public (TMED-2) and a private AS dataset, along with a CIFAR-10-derived toy dataset. Results show improvement on all the datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge