Mohamed Harmanani
Vision-Language Models Encode Clinical Guidelines for Concept-Based Medical Reasoning
Mar 09, 2026Abstract:Concept Bottleneck Models (CBMs) are a prominent framework for interpretable AI that map learned visual features to a set of meaningful concepts for task-specific downstream predictions. Their sequential structure enhances transparency by connecting model predictions to the underlying concepts that support them. In medical imaging, where transparency is essential, CBMs offer an appealing foundation for explainable model design. However, discrete concept representations often overlook broader clinical context such as diagnostic guidelines and expert heuristics, reducing reliability in complex cases. We propose MedCBR, a concept-based reasoning framework that integrates clinical guidelines with vision-language and reasoning models. Labeled clinical descriptors are transformed into guideline-conformant text, and a concept-based model is trained with a multitask objective combining multimodal contrastive alignment, concept supervision, and diagnostic classification to jointly ground image features, concepts, and pathology. A reasoning model then converts these predictions into structured clinical narratives that explain the diagnosis, emulating expert reasoning based on established guidelines. MedCBR achieves superior diagnostic and concept-level performance, with AUROCs of 94.2% on ultrasound and 84.0% on mammography. Further experiments on non-medical datasets achieve 86.1% accuracy. Our framework enhances interpretability and forms an end-to-end bridge from medical image analysis to decision-making.
ProstNFound+: A Prospective Study using Medical Foundation Models for Prostate Cancer Detection
Oct 30, 2025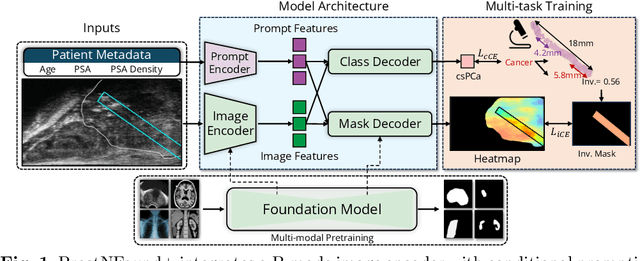
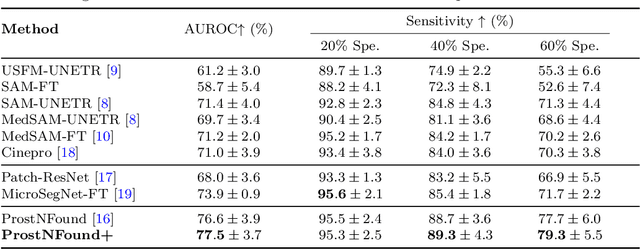
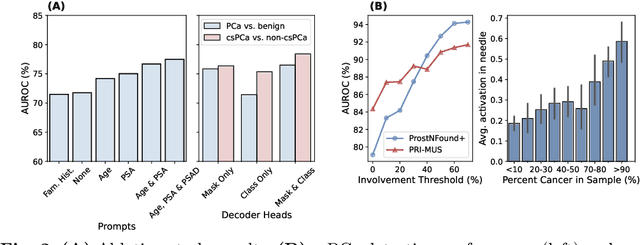
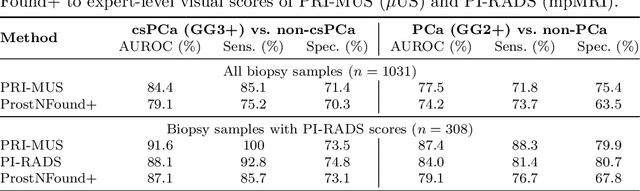
Abstract:Purpose: Medical foundation models (FMs) offer a path to build high-performance diagnostic systems. However, their application to prostate cancer (PCa) detection from micro-ultrasound ({\mu}US) remains untested in clinical settings. We present ProstNFound+, an adaptation of FMs for PCa detection from {\mu}US, along with its first prospective validation. Methods: ProstNFound+ incorporates a medical FM, adapter tuning, and a custom prompt encoder that embeds PCa-specific clinical biomarkers. The model generates a cancer heatmap and a risk score for clinically significant PCa. Following training on multi-center retrospective data, the model is prospectively evaluated on data acquired five years later from a new clinical site. Model predictions are benchmarked against standard clinical scoring protocols (PRI-MUS and PI-RADS). Results: ProstNFound+ shows strong generalization to the prospective data, with no performance degradation compared to retrospective evaluation. It aligns closely with clinical scores and produces interpretable heatmaps consistent with biopsy-confirmed lesions. Conclusion: The results highlight its potential for clinical deployment, offering a scalable and interpretable alternative to expert-driven protocols.
Diverse Prototypical Ensembles Improve Robustness to Subpopulation Shift
May 29, 2025Abstract:The subpopulationtion shift, characterized by a disparity in subpopulation distributibetween theween the training and target datasets, can significantly degrade the performance of machine learning models. Current solutions to subpopulation shift involve modifying empirical risk minimization with re-weighting strategies to improve generalization. This strategy relies on assumptions about the number and nature of subpopulations and annotations on group membership, which are unavailable for many real-world datasets. Instead, we propose using an ensemble of diverse classifiers to adaptively capture risk associated with subpopulations. Given a feature extractor network, we replace its standard linear classification layer with a mixture of prototypical classifiers, where each member is trained to classify the data while focusing on different features and samples from other members. In empirical evaluation on nine real-world datasets, covering diverse domains and kinds of subpopulation shift, our method of Diverse Prototypical Ensembles (DPEs) often outperforms the prior state-of-the-art in worst-group accuracy. The code is available at https://github.com/minhto2802/dpe4subpop
TRUSWorthy: Toward Clinically Applicable Deep Learning for Confident Detection of Prostate Cancer in Micro-Ultrasound
Feb 20, 2025Abstract:While deep learning methods have shown great promise in improving the effectiveness of prostate cancer (PCa) diagnosis by detecting suspicious lesions from trans-rectal ultrasound (TRUS), they must overcome multiple simultaneous challenges. There is high heterogeneity in tissue appearance, significant class imbalance in favor of benign examples, and scarcity in the number and quality of ground truth annotations available to train models. Failure to address even a single one of these problems can result in unacceptable clinical outcomes.We propose TRUSWorthy, a carefully designed, tuned, and integrated system for reliable PCa detection. Our pipeline integrates self-supervised learning, multiple-instance learning aggregation using transformers, random-undersampled boosting and ensembling: these address label scarcity, weak labels, class imbalance, and overconfidence, respectively. We train and rigorously evaluate our method using a large, multi-center dataset of micro-ultrasound data. Our method outperforms previous state-of-the-art deep learning methods in terms of accuracy and uncertainty calibration, with AUROC and balanced accuracy scores of 79.9% and 71.5%, respectively. On the top 20% of predictions with the highest confidence, we can achieve a balanced accuracy of up to 91%. The success of TRUSWorthy demonstrates the potential of integrated deep learning solutions to meet clinical needs in a highly challenging deployment setting, and is a significant step towards creating a trustworthy system for computer-assisted PCa diagnosis.
Cinepro: Robust Training of Foundation Models for Cancer Detection in Prostate Ultrasound Cineloops
Jan 21, 2025



Abstract:Prostate cancer (PCa) detection using deep learning (DL) models has shown potential for enhancing real-time guidance during biopsies. However, prostate ultrasound images lack pixel-level cancer annotations, introducing label noise. Current approaches often focus on limited regions of interest (ROIs), disregarding anatomical context necessary for accurate diagnosis. Foundation models can overcome this limitation by analyzing entire images to capture global spatial relationships; however, they still encounter challenges stemming from the weak labels associated with coarse pathology annotations in ultrasound data. We introduce Cinepro, a novel framework that strengthens foundation models' ability to localize PCa in ultrasound cineloops. Cinepro adapts robust training by integrating the proportion of cancer tissue reported by pathology in a biopsy core into its loss function to address label noise, providing a more nuanced supervision. Additionally, it leverages temporal data across multiple frames to apply robust augmentations, enhancing the model's ability to learn stable cancer-related features. Cinepro demonstrates superior performance on a multi-center prostate ultrasound dataset, achieving an AUROC of 77.1% and a balanced accuracy of 83.8%, surpassing current benchmarks. These findings underscore Cinepro's promise in advancing foundation models for weakly labeled ultrasound data.
Calibrated Diverse Ensemble Entropy Minimization for Robust Test-Time Adaptation in Prostate Cancer Detection
Jul 17, 2024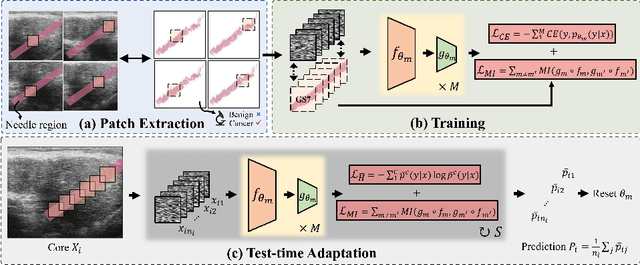
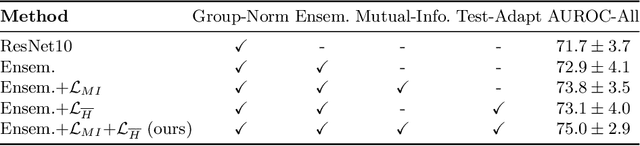
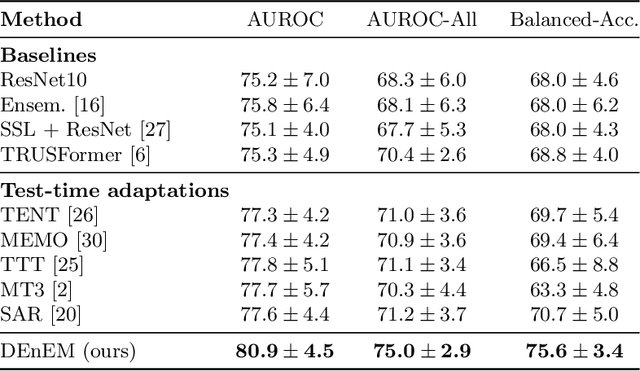

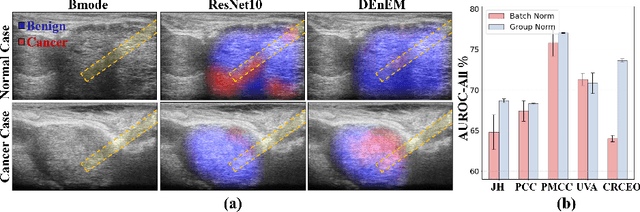
Abstract:High resolution micro-ultrasound has demonstrated promise in real-time prostate cancer detection, with deep learning becoming a prominent tool for learning complex tissue properties reflected on ultrasound. However, a significant roadblock to real-world deployment remains, which prior works often overlook: model performance suffers when applied to data from different clinical centers due to variations in data distribution. This distribution shift significantly impacts the model's robustness, posing major challenge to clinical deployment. Domain adaptation and specifically its test-time adaption (TTA) variant offer a promising solution to address this challenge. In a setting designed to reflect real-world conditions, we compare existing methods to state-of-the-art TTA approaches adopted for cancer detection, demonstrating the lack of robustness to distribution shifts in the former. We then propose Diverse Ensemble Entropy Minimization (DEnEM), questioning the effectiveness of current TTA methods on ultrasound data. We show that these methods, although outperforming baselines, are suboptimal due to relying on neural networks output probabilities, which could be uncalibrated, or relying on data augmentation, which is not straightforward to define on ultrasound data. Our results show a significant improvement of $5\%$ to $7\%$ in AUROC over the existing methods and $3\%$ to $5\%$ over TTA methods, demonstrating the advantage of DEnEM in addressing distribution shift. \keywords{Ultrasound Imaging \and Prostate Cancer \and Computer-aided Diagnosis \and Distribution Shift Robustness \and Test-time Adaptation.}
Benchmarking Image Transformers for Prostate Cancer Detection from Ultrasound Data
Mar 27, 2024Abstract:PURPOSE: Deep learning methods for classifying prostate cancer (PCa) in ultrasound images typically employ convolutional networks (CNNs) to detect cancer in small regions of interest (ROI) along a needle trace region. However, this approach suffers from weak labelling, since the ground-truth histopathology labels do not describe the properties of individual ROIs. Recently, multi-scale approaches have sought to mitigate this issue by combining the context awareness of transformers with a CNN feature extractor to detect cancer from multiple ROIs using multiple-instance learning (MIL). In this work, we present a detailed study of several image transformer architectures for both ROI-scale and multi-scale classification, and a comparison of the performance of CNNs and transformers for ultrasound-based prostate cancer classification. We also design a novel multi-objective learning strategy that combines both ROI and core predictions to further mitigate label noise. METHODS: We evaluate 3 image transformers on ROI-scale cancer classification, then use the strongest model to tune a multi-scale classifier with MIL. We train our MIL models using our novel multi-objective learning strategy and compare our results to existing baselines. RESULTS: We find that for both ROI-scale and multi-scale PCa detection, image transformer backbones lag behind their CNN counterparts. This deficit in performance is even more noticeable for larger models. When using multi-objective learning, we can improve performance of MIL, with a 77.9% AUROC, a sensitivity of 75.9%, and a specificity of 66.3%. CONCLUSION: Convolutional networks are better suited for modelling sparse datasets of prostate ultrasounds, producing more robust features than transformers in PCa detection. Multi-scale methods remain the best architecture for this task, with multi-objective learning presenting an effective way to improve performance.
Modelling the Spread of COVID-19 in Indoor Spaces using Automated Probabilistic Planning
Aug 16, 2023Abstract:The coronavirus disease 2019 (COVID-19) pandemic has been ongoing for around 3 years, and has infected over 750 million people and caused over 6 million deaths worldwide at the time of writing. Throughout the pandemic, several strategies for controlling the spread of the disease have been debated by healthcare professionals, government authorities, and international bodies. To anticipate the potential impact of the disease, and to simulate the effectiveness of different mitigation strategies, a robust model of disease spread is needed. In this work, we explore a novel approach based on probabilistic planning and dynamic graph analysis to model the spread of COVID-19 in indoor spaces. We endow the planner with means to control the spread of the disease through non-pharmaceutical interventions (NPIs) such as mandating masks and vaccines, and we compare the impact of crowds and capacity limits on the spread of COVID-19 in these settings. We demonstrate that the use of probabilistic planning is effective in predicting the amount of infections that are likely to occur in shared spaces, and that automated planners have the potential to design competent interventions to limit the spread of the disease. Our code is fully open-source and is available at: https://github.com/mharmanani/prob-planning-covid19 .
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge