Nasir Rajpoot
*: shared first/last authors
MPath: Multimodal Pathology Report Generation from Whole Slide Images
Dec 10, 2025Abstract:Automated generation of diagnostic pathology reports directly from whole slide images (WSIs) is an emerging direction in computational pathology. Translating high-resolution tissue patterns into clinically coherent text remains difficult due to large morphological variability and the complex structure of pathology narratives. We introduce MPath, a lightweight multimodal framework that conditions a pretrained biomedical language model (BioBART) on WSI-derived visual embeddings through a learned visual-prefix prompting mechanism. Instead of end-to-end vision-language pretraining, MPath leverages foundation-model WSI features (CONCH + Titan) and injects them into BioBART via a compact projection module, keeping the language backbone frozen for stability and data efficiency. MPath was developed and evaluated on the RED 2025 Grand Challenge dataset and ranked 4th in Test Phase 2, despite limited submission opportunities. The results highlight the potential of prompt-based multimodal conditioning as a scalable and interpretable strategy for pathology report generation.
Tissue Aware Nuclei Detection and Classification Model for Histopathology Images
Nov 17, 2025Abstract:Accurate nuclei detection and classification are fundamental to computational pathology, yet existing approaches are hindered by reliance on detailed expert annotations and insufficient use of tissue context. We present Tissue-Aware Nuclei Detection (TAND), a novel framework achieving joint nuclei detection and classification using point-level supervision enhanced by tissue mask conditioning. TAND couples a ConvNeXt-based encoder-decoder with a frozen Virchow-2 tissue segmentation branch, where semantic tissue probabilities selectively modulate the classification stream through a novel multi-scale Spatial Feature-wise Linear Modulation (Spatial-FiLM). On the PUMA benchmark, TAND achieves state-of-the-art performance, surpassing both tissue-agnostic baselines and mask-supervised methods. Notably, our approach demonstrates remarkable improvements in tissue-dependent cell types such as epithelium, endothelium, and stroma. To the best of our knowledge, this is the first method to condition per-cell classification on learned tissue masks, offering a practical pathway to reduce annotation burden.
A Deep Learning Framework for Thyroid Nodule Segmentation and Malignancy Classification from Ultrasound Images
Nov 14, 2025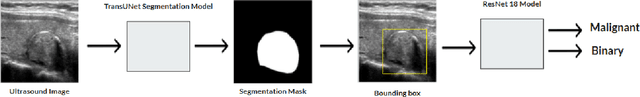
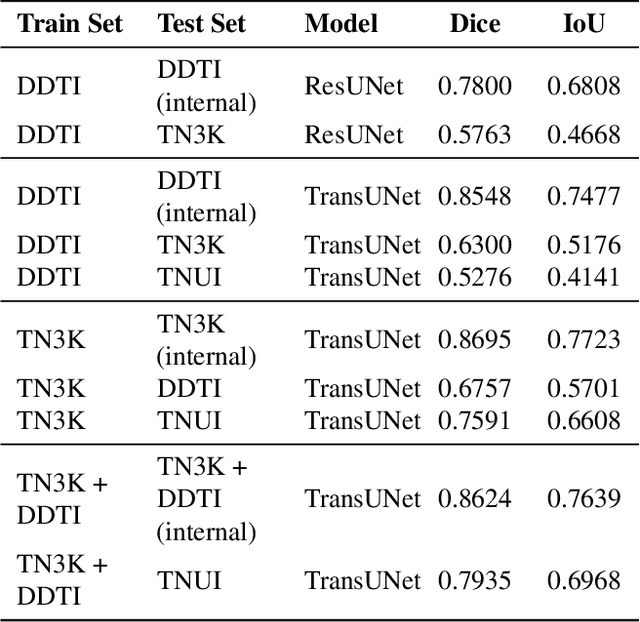
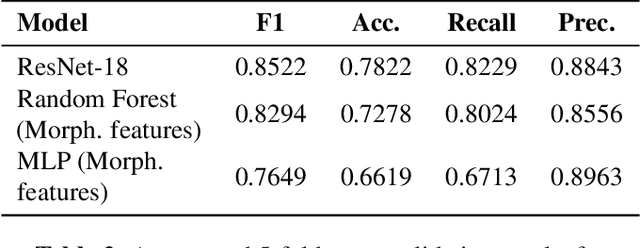
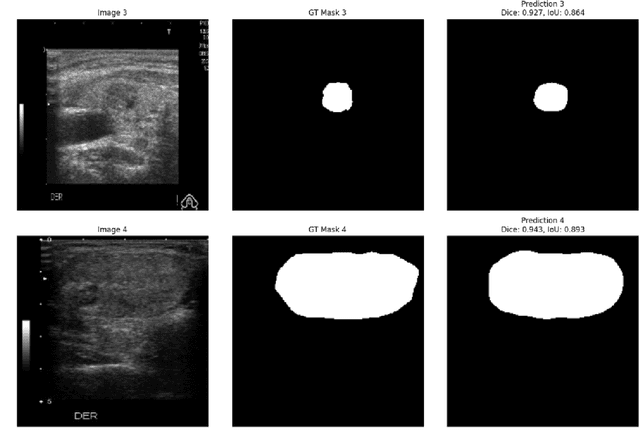
Abstract:Ultrasound-based risk stratification of thyroid nodules is a critical clinical task, but it suffers from high inter-observer variability. While many deep learning (DL) models function as "black boxes," we propose a fully automated, two-stage framework for interpretable malignancy prediction. Our method achieves interpretability by forcing the model to focus only on clinically relevant regions. First, a TransUNet model automatically segments the thyroid nodule. The resulting mask is then used to create a region of interest around the nodule, and this localised image is fed directly into a ResNet-18 classifier. We evaluated our framework using 5-fold cross-validation on a clinical dataset of 349 images, where it achieved a high F1-score of 0.852 for predicting malignancy. To validate its performance, we compared it against a strong baseline using a Random Forest classifier with hand-crafted morphological features, which achieved an F1-score of 0.829. The superior performance of our DL framework suggests that the implicit visual features learned from the localised nodule are more predictive than explicit shape features alone. This is the first fully automated end-to-end pipeline for both detecting thyroid nodules on ultrasound images and predicting their malignancy.
Synergy vs. Noise: Performance-Guided Multimodal Fusion For Biochemical Recurrence-Free Survival in Prostate Cancer
Nov 14, 2025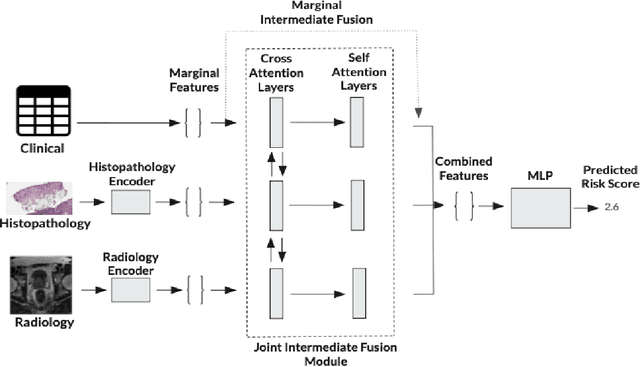
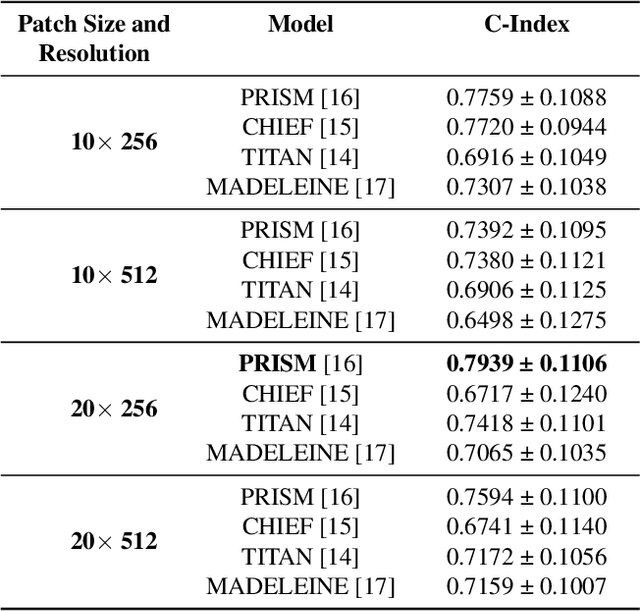
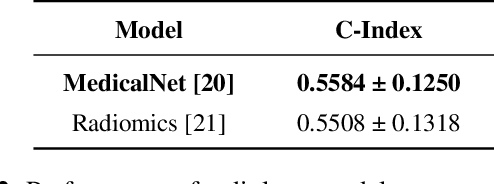
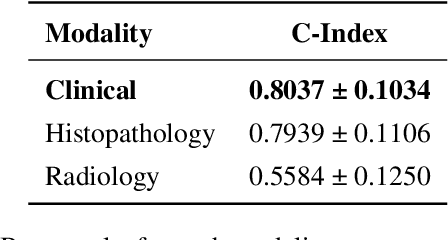
Abstract:Multimodal deep learning (MDL) has emerged as a transformative approach in computational pathology. By integrating complementary information from multiple data sources, MDL models have demonstrated superior predictive performance across diverse clinical tasks compared to unimodal models. However, the assumption that combining modalities inherently improves performance remains largely unexamined. We hypothesise that multimodal gains depend critically on the predictive quality of individual modalities, and that integrating weak modalities may introduce noise rather than complementary information. We test this hypothesis on a prostate cancer dataset with histopathology, radiology, and clinical data to predict time-to-biochemical recurrence. Our results confirm that combining high-performing modalities yield superior performance compared to unimodal approaches. However, integrating a poor-performing modality with other higher-performing modalities degrades predictive accuracy. These findings demonstrate that multimodal benefit requires selective, performance-guided integration rather than indiscriminate modality combination, with implications for MDL design across computational pathology and medical imaging.
CORE - A Cell-Level Coarse-to-Fine Image Registration Engine for Multi-stain Image Alignment
Nov 12, 2025Abstract:Accurate and efficient registration of whole slide images (WSIs) is essential for high-resolution, nuclei-level analysis in multi-stained tissue slides. We propose a novel coarse-to-fine framework CORE for accurate nuclei-level registration across diverse multimodal whole-slide image (WSI) datasets. The coarse registration stage leverages prompt-based tissue mask extraction to effectively filter out artefacts and non-tissue regions, followed by global alignment using tissue morphology and ac- celerated dense feature matching with a pre-trained feature extractor. From the coarsely aligned slides, nuclei centroids are detected and subjected to fine-grained rigid registration using a custom, shape-aware point-set registration model. Finally, non-rigid alignment at the cellular level is achieved by estimating a non-linear dis- placement field using Coherent Point Drift (CPD). Our approach benefits from automatically generated nuclei that enhance the accuracy of deformable registra- tion and ensure precise nuclei-level correspondence across modalities. The pro- posed model is evaluated on three publicly available WSI registration datasets, and two private datasets. We show that CORE outperforms current state-of-the-art methods in terms of generalisability, precision, and robustness in bright-field and immunofluorescence microscopy WSIs
Foundation Models in Computational Pathology: A Review of Challenges, Opportunities, and Impact
Feb 12, 2025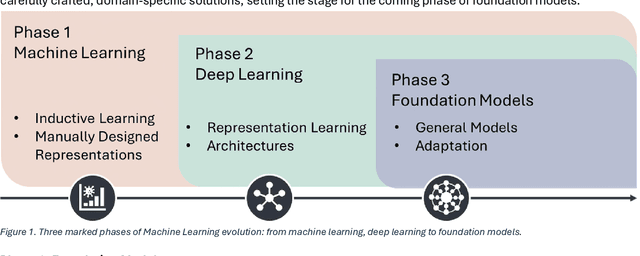
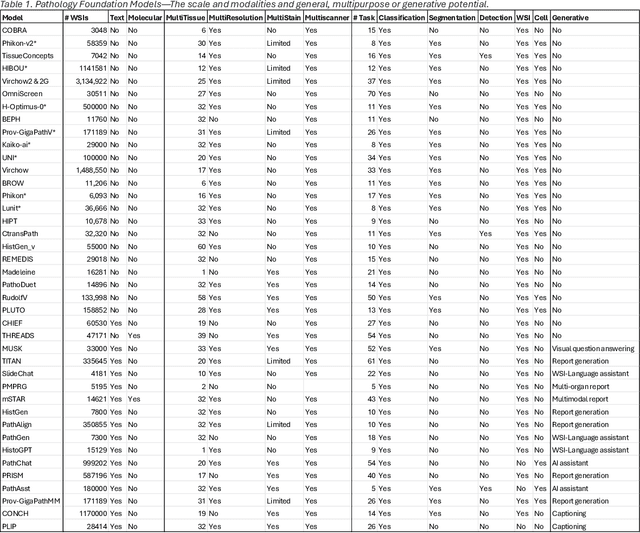
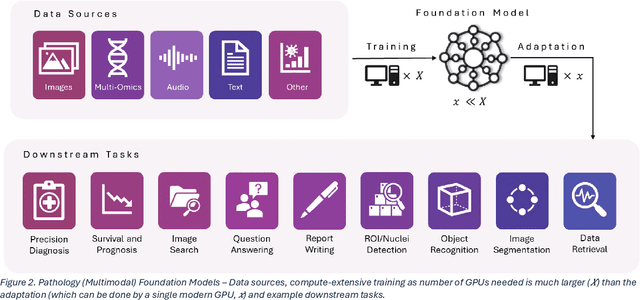
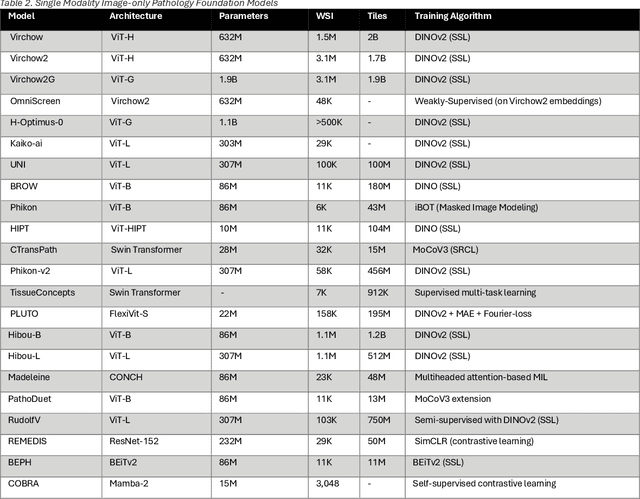
Abstract:From self-supervised, vision-only models to contrastive visual-language frameworks, computational pathology has rapidly evolved in recent years. Generative AI "co-pilots" now demonstrate the ability to mine subtle, sub-visual tissue cues across the cellular-to-pathology spectrum, generate comprehensive reports, and respond to complex user queries. The scale of data has surged dramatically, growing from tens to millions of multi-gigapixel tissue images, while the number of trainable parameters in these models has risen to several billion. The critical question remains: how will this new wave of generative and multi-purpose AI transform clinical diagnostics? In this article, we explore the true potential of these innovations and their integration into clinical practice. We review the rapid progress of foundation models in pathology, clarify their applications and significance. More precisely, we examine the very definition of foundational models, identifying what makes them foundational, general, or multipurpose, and assess their impact on computational pathology. Additionally, we address the unique challenges associated with their development and evaluation. These models have demonstrated exceptional predictive and generative capabilities, but establishing global benchmarks is crucial to enhancing evaluation standards and fostering their widespread clinical adoption. In computational pathology, the broader impact of frontier AI ultimately depends on widespread adoption and societal acceptance. While direct public exposure is not strictly necessary, it remains a powerful tool for dispelling misconceptions, building trust, and securing regulatory support.
Stain-Invariant Representation for Tissue Classification in Histology Images
Nov 21, 2024
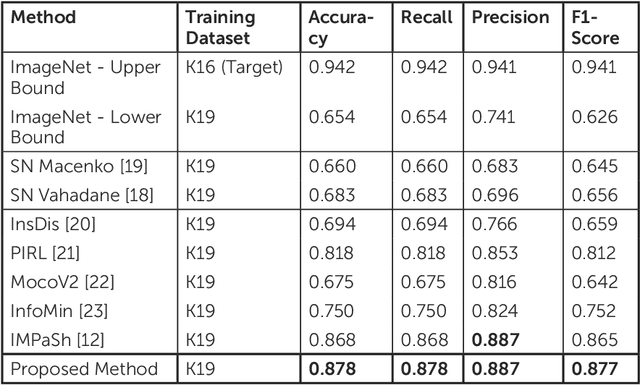
Abstract:The process of digitising histology slides involves multiple factors that can affect a whole slide image's (WSI) final appearance, including the staining protocol, scanner, and tissue type. This variability constitutes a domain shift and results in significant problems when training and testing deep learning (DL) algorithms in multi-cohort settings. As such, developing robust and generalisable DL models in computational pathology (CPath) remains an open challenge. In this regard, we propose a framework that generates stain-augmented versions of the training images using stain matrix perturbation. Thereafter, we employed a stain regularisation loss to enforce consistency between the feature representations of the source and augmented images. Doing so encourages the model to learn stain-invariant and, consequently, domain-invariant feature representations. We evaluate the performance of the proposed model on cross-domain multi-class tissue type classification of colorectal cancer images and have achieved improved performance compared to other state-of-the-art methods.
Benchmarking Domain Generalization Algorithms in Computational Pathology
Sep 25, 2024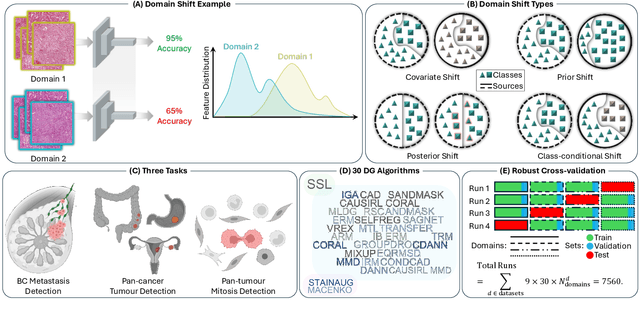
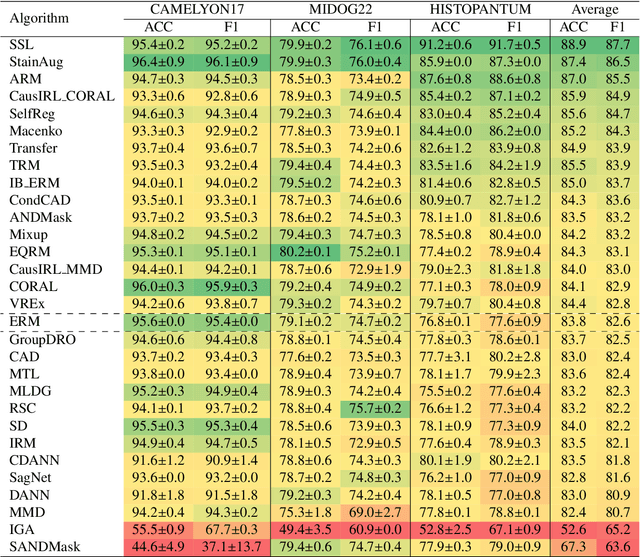
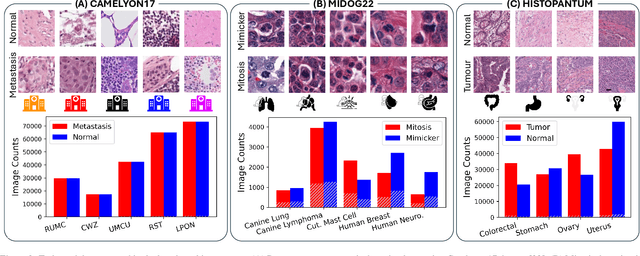
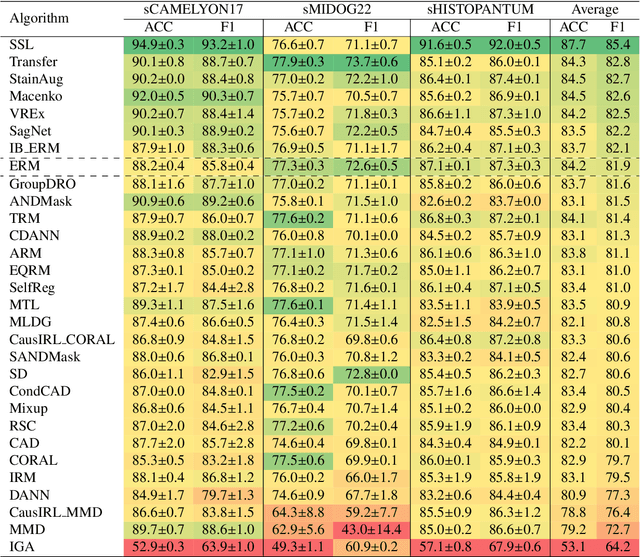
Abstract:Deep learning models have shown immense promise in computational pathology (CPath) tasks, but their performance often suffers when applied to unseen data due to domain shifts. Addressing this requires domain generalization (DG) algorithms. However, a systematic evaluation of DG algorithms in the CPath context is lacking. This study aims to benchmark the effectiveness of 30 DG algorithms on 3 CPath tasks of varying difficulty through 7,560 cross-validation runs. We evaluate these algorithms using a unified and robust platform, incorporating modality-specific techniques and recent advances like pretrained foundation models. Our extensive cross-validation experiments provide insights into the relative performance of various DG strategies. We observe that self-supervised learning and stain augmentation consistently outperform other methods, highlighting the potential of pretrained models and data augmentation. Furthermore, we introduce a new pan-cancer tumor detection dataset (HISTOPANTUM) as a benchmark for future research. This study offers valuable guidance to researchers in selecting appropriate DG approaches for CPath tasks.
StainFuser: Controlling Diffusion for Faster Neural Style Transfer in Multi-Gigapixel Histology Images
Mar 14, 2024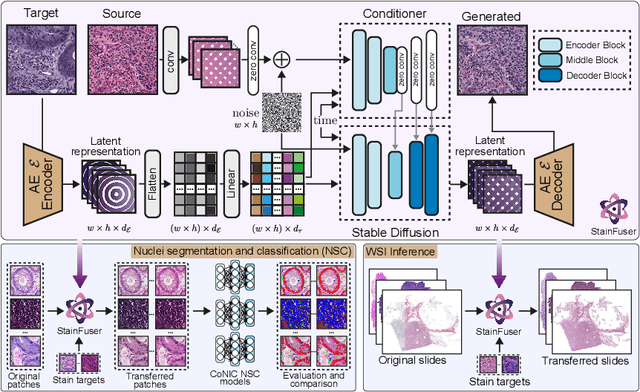
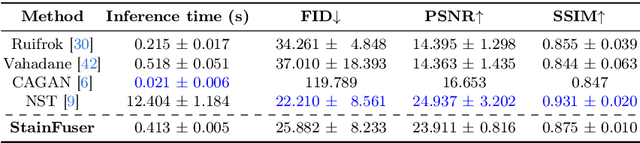

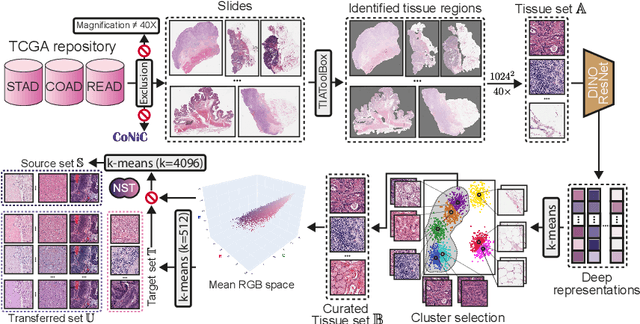
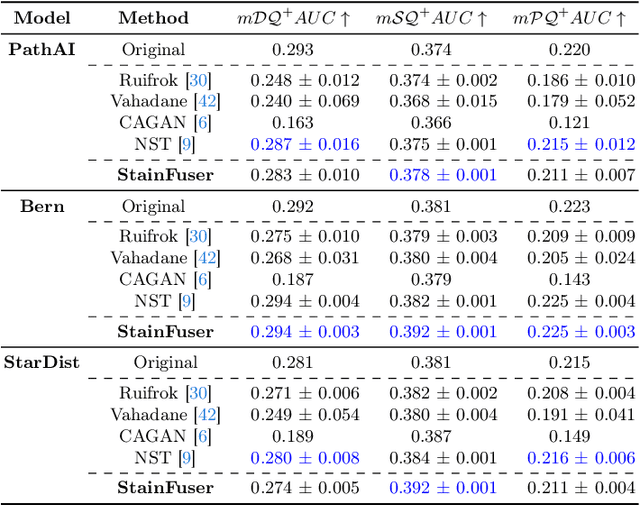
Abstract:Stain normalization algorithms aim to transform the color and intensity characteristics of a source multi-gigapixel histology image to match those of a target image, mitigating inconsistencies in the appearance of stains used to highlight cellular components in the images. We propose a new approach, StainFuser, which treats this problem as a style transfer task using a novel Conditional Latent Diffusion architecture, eliminating the need for handcrafted color components. With this method, we curate SPI-2M the largest stain normalization dataset to date of over 2 million histology images with neural style transfer for high-quality transformations. Trained on this data, StainFuser outperforms current state-of-the-art GAN and handcrafted methods in terms of the quality of normalized images. Additionally, compared to existing approaches, it improves the performance of nuclei instance segmentation and classification models when used as a test time augmentation method on the challenging CoNIC dataset. Finally, we apply StainFuser on multi-gigapixel Whole Slide Images (WSIs) and demonstrate improved performance in terms of computational efficiency, image quality and consistency across tiles over current methods.
An AI based Digital Score of Tumour-Immune Microenvironment Predicts Benefit to Maintenance Immunotherapy in Advanced Oesophagogastric Adenocarcinoma
Feb 29, 2024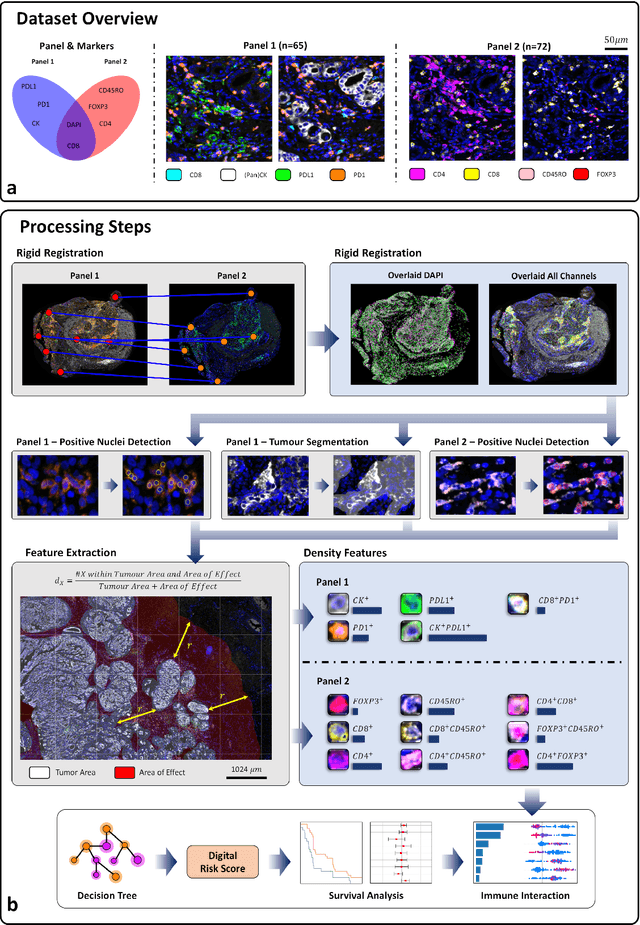
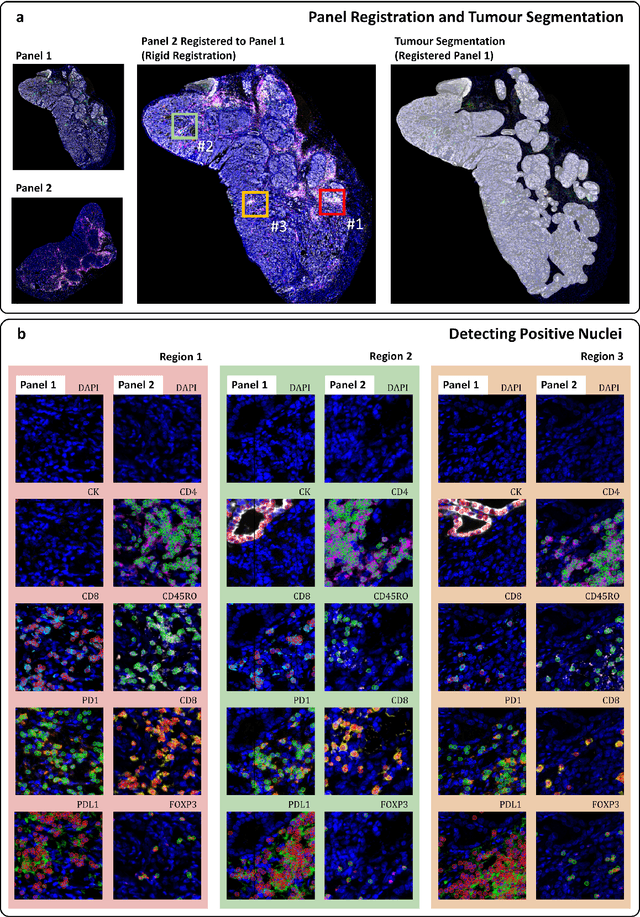
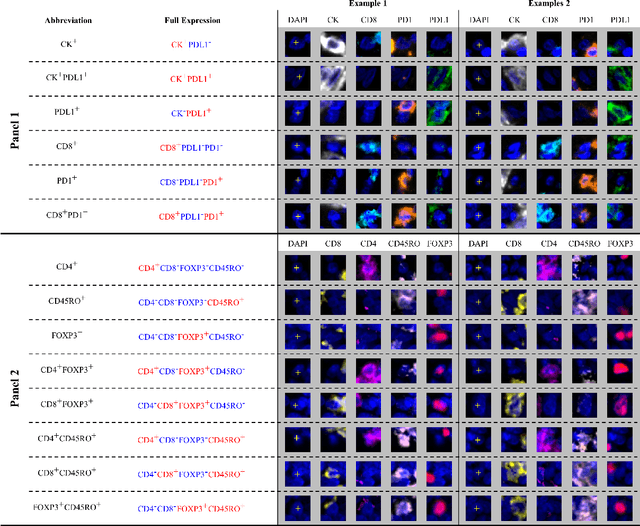
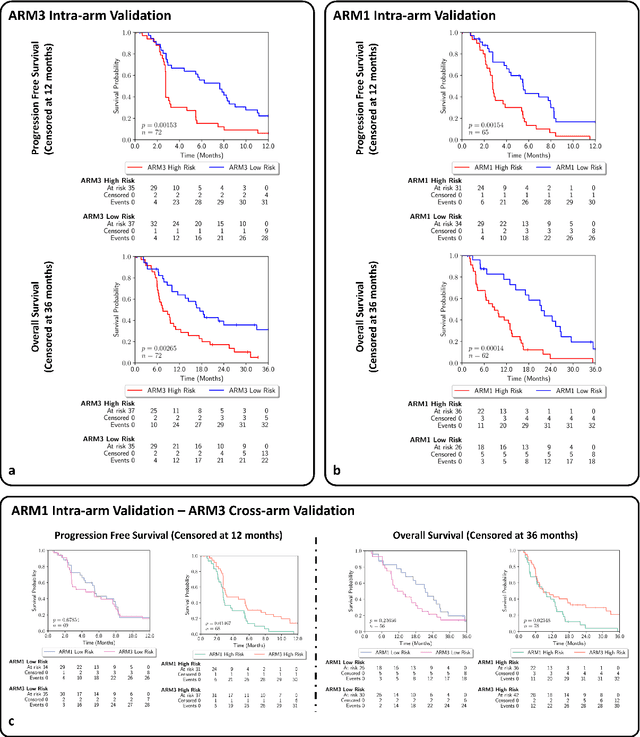
Abstract:Gastric and oesophageal (OG) cancers are the leading causes of cancer mortality worldwide. In OG cancers, recent studies have showed that PDL1 immune checkpoint inhibitors (ICI) in combination with chemotherapy improves patient survival. However, our understanding of the tumour immune microenvironment in OG cancers remains limited. In this study, we interrogate multiplex immunofluorescence (mIF) images taken from patients with advanced Oesophagogastric Adenocarcinoma (OGA) who received first-line fluoropyrimidine and platinum-based chemotherapy in the PLATFORM trial (NCT02678182) to predict the efficacy of the treatment and to explore the biological basis of patients responding to maintenance durvalumab (PDL1 inhibitor). Our proposed Artificial Intelligence (AI) based marker successfully identified responder from non-responder (p < 0.05) as well as those who could potentially benefit from ICI with statistical significance (p < 0.05) for both progression free and overall survival. Our findings suggest that T cells that express FOXP3 seem to heavily influence the patient treatment response and survival outcome. We also observed that higher levels of CD8+PD1+ cells are consistently linked to poor prognosis for both OS and PFS, regardless of ICI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge