Kesi Xu
Tissue Aware Nuclei Detection and Classification Model for Histopathology Images
Nov 17, 2025Abstract:Accurate nuclei detection and classification are fundamental to computational pathology, yet existing approaches are hindered by reliance on detailed expert annotations and insufficient use of tissue context. We present Tissue-Aware Nuclei Detection (TAND), a novel framework achieving joint nuclei detection and classification using point-level supervision enhanced by tissue mask conditioning. TAND couples a ConvNeXt-based encoder-decoder with a frozen Virchow-2 tissue segmentation branch, where semantic tissue probabilities selectively modulate the classification stream through a novel multi-scale Spatial Feature-wise Linear Modulation (Spatial-FiLM). On the PUMA benchmark, TAND achieves state-of-the-art performance, surpassing both tissue-agnostic baselines and mask-supervised methods. Notably, our approach demonstrates remarkable improvements in tissue-dependent cell types such as epithelium, endothelium, and stroma. To the best of our knowledge, this is the first method to condition per-cell classification on learned tissue masks, offering a practical pathway to reduce annotation burden.
Benchmarking Domain Generalization Algorithms in Computational Pathology
Sep 25, 2024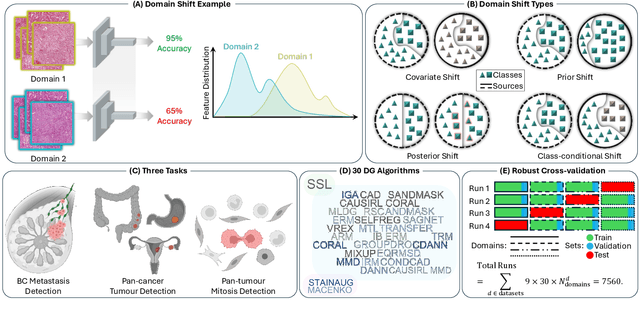
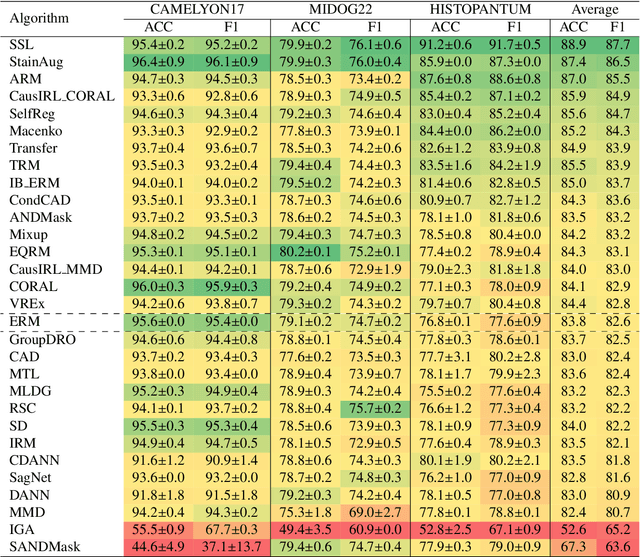
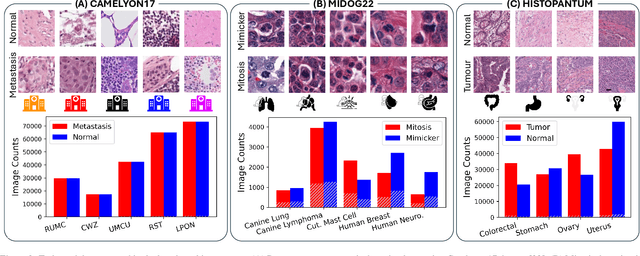
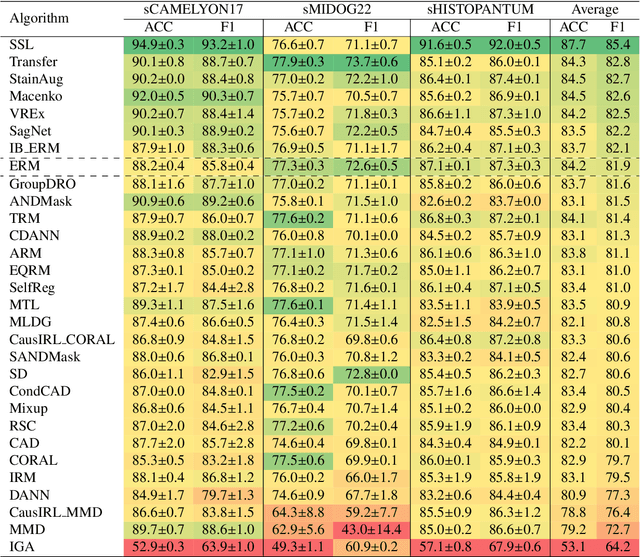
Abstract:Deep learning models have shown immense promise in computational pathology (CPath) tasks, but their performance often suffers when applied to unseen data due to domain shifts. Addressing this requires domain generalization (DG) algorithms. However, a systematic evaluation of DG algorithms in the CPath context is lacking. This study aims to benchmark the effectiveness of 30 DG algorithms on 3 CPath tasks of varying difficulty through 7,560 cross-validation runs. We evaluate these algorithms using a unified and robust platform, incorporating modality-specific techniques and recent advances like pretrained foundation models. Our extensive cross-validation experiments provide insights into the relative performance of various DG strategies. We observe that self-supervised learning and stain augmentation consistently outperform other methods, highlighting the potential of pretrained models and data augmentation. Furthermore, we introduce a new pan-cancer tumor detection dataset (HISTOPANTUM) as a benchmark for future research. This study offers valuable guidance to researchers in selecting appropriate DG approaches for CPath tasks.
On generalisability of segment anything model for nuclear instance segmentation in histology images
Jan 25, 2024Abstract:Pre-trained on a large and diverse dataset, the segment anything model (SAM) is the first promptable foundation model in computer vision aiming at object segmentation tasks. In this work, we evaluate SAM for the task of nuclear instance segmentation performance with zero-shot learning and finetuning. We compare SAM with other representative methods in nuclear instance segmentation, especially in the context of model generalisability. To achieve automatic nuclear instance segmentation, we propose using a nuclei detection model to provide bounding boxes or central points of nu-clei as visual prompts for SAM in generating nuclear instance masks from histology images.
Domain Generalization in Computational Pathology: Survey and Guidelines
Oct 30, 2023



Abstract:Deep learning models have exhibited exceptional effectiveness in Computational Pathology (CPath) by tackling intricate tasks across an array of histology image analysis applications. Nevertheless, the presence of out-of-distribution data (stemming from a multitude of sources such as disparate imaging devices and diverse tissue preparation methods) can cause \emph{domain shift} (DS). DS decreases the generalization of trained models to unseen datasets with slightly different data distributions, prompting the need for innovative \emph{domain generalization} (DG) solutions. Recognizing the potential of DG methods to significantly influence diagnostic and prognostic models in cancer studies and clinical practice, we present this survey along with guidelines on achieving DG in CPath. We rigorously define various DS types, systematically review and categorize existing DG approaches and resources in CPath, and provide insights into their advantages, limitations, and applicability. We also conduct thorough benchmarking experiments with 28 cutting-edge DG algorithms to address a complex DG problem. Our findings suggest that careful experiment design and CPath-specific Stain Augmentation technique can be very effective. However, there is no one-size-fits-all solution for DG in CPath. Therefore, we establish clear guidelines for detecting and managing DS depending on different scenarios. While most of the concepts, guidelines, and recommendations are given for applications in CPath, we believe that they are applicable to most medical image analysis tasks as well.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge