Denis Jered McInerney
Open (Clinical) LLMs are Sensitive to Instruction Phrasings
Jul 12, 2024



Abstract:Instruction-tuned Large Language Models (LLMs) can perform a wide range of tasks given natural language instructions to do so, but they are sensitive to how such instructions are phrased. This issue is especially concerning in healthcare, as clinicians are unlikely to be experienced prompt engineers and the potential consequences of inaccurate outputs are heightened in this domain. This raises a practical question: How robust are instruction-tuned LLMs to natural variations in the instructions provided for clinical NLP tasks? We collect prompts from medical doctors across a range of tasks and quantify the sensitivity of seven LLMs -- some general, others specialized -- to natural (i.e., non-adversarial) instruction phrasings. We find that performance varies substantially across all models, and that -- perhaps surprisingly -- domain-specific models explicitly trained on clinical data are especially brittle, compared to their general domain counterparts. Further, arbitrary phrasing differences can affect fairness, e.g., valid but distinct instructions for mortality prediction yield a range both in overall performance, and in terms of differences between demographic groups.
Towards Reducing Diagnostic Errors with Interpretable Risk Prediction
Feb 15, 2024Abstract:Many diagnostic errors occur because clinicians cannot easily access relevant information in patient Electronic Health Records (EHRs). In this work we propose a method to use LLMs to identify pieces of evidence in patient EHR data that indicate increased or decreased risk of specific diagnoses; our ultimate aim is to increase access to evidence and reduce diagnostic errors. In particular, we propose a Neural Additive Model to make predictions backed by evidence with individualized risk estimates at time-points where clinicians are still uncertain, aiming to specifically mitigate delays in diagnosis and errors stemming from an incomplete differential. To train such a model, it is necessary to infer temporally fine-grained retrospective labels of eventual "true" diagnoses. We do so with LLMs, to ensure that the input text is from before a confident diagnosis can be made. We use an LLM to retrieve an initial pool of evidence, but then refine this set of evidence according to correlations learned by the model. We conduct an in-depth evaluation of the usefulness of our approach by simulating how it might be used by a clinician to decide between a pre-defined list of differential diagnoses.
Leveraging Generative AI for Clinical Evidence Summarization Needs to Achieve Trustworthiness
Nov 19, 2023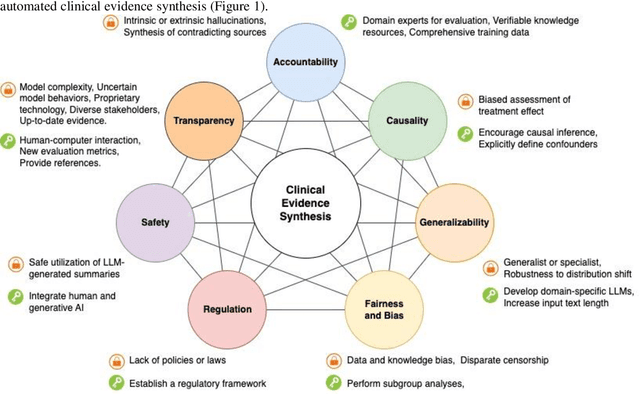
Abstract:Evidence-based medicine aims to improve the quality of healthcare by empowering medical decisions and practices with the best available evidence. The rapid growth of medical evidence, which can be obtained from various sources, poses a challenge in collecting, appraising, and synthesizing the evidential information. Recent advancements in generative AI, exemplified by large language models, hold promise in facilitating the arduous task. However, developing accountable, fair, and inclusive models remains a complicated undertaking. In this perspective, we discuss the trustworthiness of generative AI in the context of automated summarization of medical evidence.
Retrieving Evidence from EHRs with LLMs: Possibilities and Challenges
Sep 08, 2023



Abstract:Unstructured Electronic Health Record (EHR) data often contains critical information complementary to imaging data that would inform radiologists' diagnoses. However, time constraints and the large volume of notes frequently associated with individual patients renders manual perusal of such data to identify relevant evidence infeasible in practice. Modern Large Language Models (LLMs) provide a flexible means of interacting with unstructured EHR data, and may provide a mechanism to efficiently retrieve and summarize unstructured evidence relevant to a given query. In this work, we propose and evaluate an LLM (Flan-T5 XXL) for this purpose. Specifically, in a zero-shot setting we task the LLM to infer whether a patient has or is at risk of a particular condition; if so, we prompt the model to summarize the supporting evidence. Enlisting radiologists for manual evaluation, we find that this LLM-based approach provides outputs consistently preferred to a standard information retrieval baseline, but we also highlight the key outstanding challenge: LLMs are prone to hallucinating evidence. However, we provide results indicating that model confidence in outputs might indicate when LLMs are hallucinating, potentially providing a means to address this.
Automatically Summarizing Evidence from Clinical Trials: A Prototype Highlighting Current Challenges
Mar 07, 2023
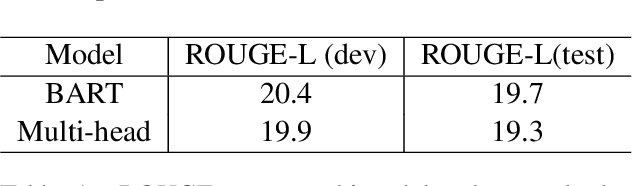

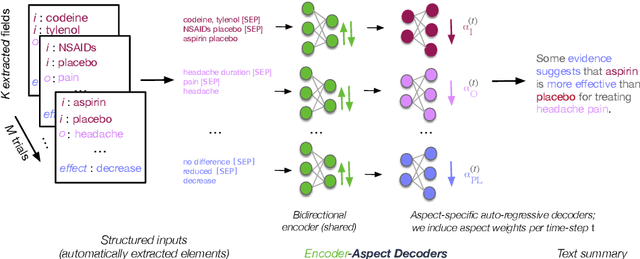

Abstract:We present TrialsSummarizer, a system that aims to automatically summarize evidence presented in the set of randomized controlled trials most relevant to a given query. Building on prior work, the system retrieves trial publications matching a query specifying a combination of condition, intervention(s), and outcome(s), and ranks these according to sample size and estimated study quality. The top-k such studies are passed through a neural multi-document summarization system, yielding a synopsis of these trials. We consider two architectures: A standard sequence-to-sequence model based on BART and a multi-headed architecture intended to provide greater transparency to end-users. Both models produce fluent and relevant summaries of evidence retrieved for queries, but their tendency to introduce unsupported statements render them inappropriate for use in this domain at present. The proposed architecture may help users verify outputs allowing users to trace generated tokens back to inputs.
CHiLL: Zero-shot Custom Interpretable Feature Extraction from Clinical Notes with Large Language Models
Feb 23, 2023



Abstract:Large Language Models (LLMs) have yielded fast and dramatic progress in NLP, and now offer strong few- and zero-shot capabilities on new tasks, reducing the need for annotation. This is especially exciting for the medical domain, in which supervision is often scant and expensive. At the same time, model predictions are rarely so accurate that they can be trusted blindly. Clinicians therefore tend to favor "interpretable" classifiers over opaque LLMs. For example, risk prediction tools are often linear models defined over manually crafted predictors that must be laboriously extracted from EHRs. We propose CHiLL (Crafting High-Level Latents), which uses LLMs to permit natural language specification of high-level features for linear models via zero-shot feature extraction using expert-composed queries. This approach has the promise to empower physicians to use their domain expertise to craft features which are clinically meaningful for a downstream task of interest, without having to manually extract these from raw EHR (as often done now). We are motivated by a real-world risk prediction task, but as a reproducible proxy, we use MIMIC-III and MIMIC-CXR data and standard predictive tasks (e.g., 30-day readmission) to evaluate our approach. We find that linear models using automatically extracted features are comparably performant to models using reference features, and provide greater interpretability than linear models using "Bag-of-Words" features. We verify that learned feature weights align well with clinical expectations.
That's the Wrong Lung! Evaluating and Improving the Interpretability of Unsupervised Multimodal Encoders for Medical Data
Oct 12, 2022



Abstract:Pretraining multimodal models on Electronic Health Records (EHRs) provides a means of learning representations that can transfer to downstream tasks with minimal supervision. Recent multimodal models induce soft local alignments between image regions and sentences. This is of particular interest in the medical domain, where alignments might highlight regions in an image relevant to specific phenomena described in free-text. While past work has suggested that attention "heatmaps" can be interpreted in this manner, there has been little evaluation of such alignments. We compare alignments from a state-of-the-art multimodal (image and text) model for EHR with human annotations that link image regions to sentences. Our main finding is that the text has an often weak or unintuitive influence on attention; alignments do not consistently reflect basic anatomical information. Moreover, synthetic modifications -- such as substituting "left" for "right" -- do not substantially influence highlights. Simple techniques such as allowing the model to opt out of attending to the image and few-shot finetuning show promise in terms of their ability to improve alignments with very little or no supervision.
Kronecker Factorization for Preventing Catastrophic Forgetting in Large-scale Medical Entity Linking
Nov 11, 2021
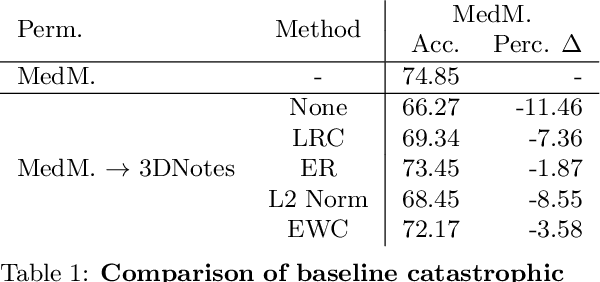

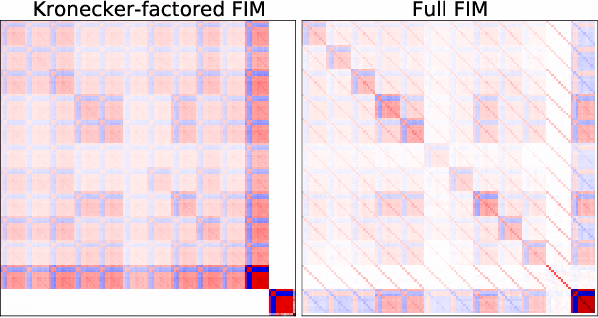
Abstract:Multi-task learning is useful in NLP because it is often practically desirable to have a single model that works across a range of tasks. In the medical domain, sequential training on tasks may sometimes be the only way to train models, either because access to the original (potentially sensitive) data is no longer available, or simply owing to the computational costs inherent to joint retraining. A major issue inherent to sequential learning, however, is catastrophic forgetting, i.e., a substantial drop in accuracy on prior tasks when a model is updated for a new task. Elastic Weight Consolidation is a recently proposed method to address this issue, but scaling this approach to the modern large models used in practice requires making strong independence assumptions about model parameters, limiting its effectiveness. In this work, we apply Kronecker Factorization--a recent approach that relaxes independence assumptions--to prevent catastrophic forgetting in convolutional and Transformer-based neural networks at scale. We show the effectiveness of this technique on the important and illustrative task of medical entity linking across three datasets, demonstrating the capability of the technique to be used to make efficient updates to existing methods as new medical data becomes available. On average, the proposed method reduces catastrophic forgetting by 51% when using a BERT-based model, compared to a 27% reduction using standard Elastic Weight Consolidation, while maintaining spatial complexity proportional to the number of model parameters.
Query-Focused EHR Summarization to Aid Imaging Diagnosis
Apr 26, 2020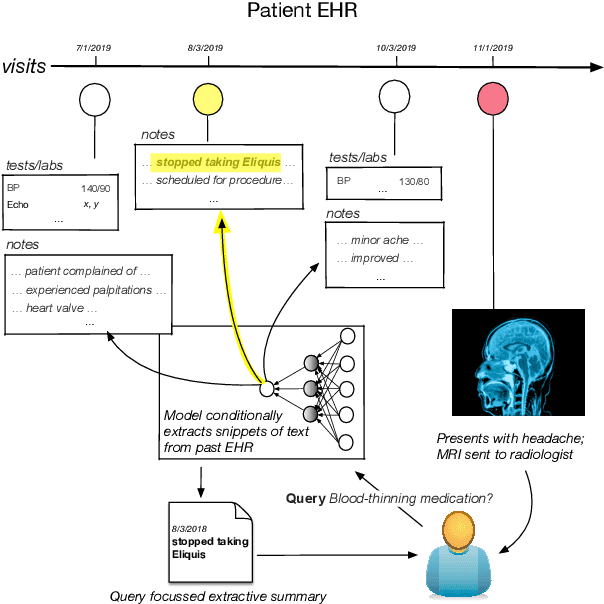
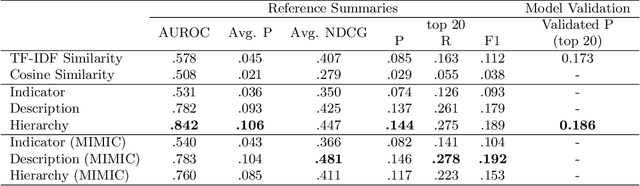


Abstract:Electronic Health Records (EHRs) provide vital contextual information to radiologists and other physicians when making a diagnosis. Unfortunately, because a given patient's record may contain hundreds of notes and reports, identifying relevant information within these in the short time typically allotted to a case is very difficult. We propose and evaluate models that extract relevant text snippets from patient records to provide a rough case summary intended to aid physicians considering one or more diagnoses. This is hard because direct supervision (i.e., physician annotations of snippets relevant to specific diagnoses in medical records) is prohibitively expensive to collect at scale. We propose a distantly supervised strategy in which we use groups of International Classification of Diseases (ICD) codes observed in 'future' records as noisy proxies for 'downstream' diagnoses. Using this we train a transformer-based neural model to perform extractive summarization conditioned on potential diagnoses. This model defines an attention mechanism that is conditioned on potential diagnoses (queries) provided by the diagnosing physician. We train (via distant supervision) and evaluate variants of this model on EHR data from Brigham and Women's Hospital in Boston and MIMIC-III (the latter to facilitate reproducibility). Evaluations performed by radiologists demonstrate that these distantly supervised models yield better extractive summaries than do unsupervised approaches. Such models may aid diagnosis by identifying sentences in past patient reports that are clinically relevant to a potential diagnosis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge