Curtis L. Cole
DIRI: Adversarial Patient Reidentification with Large Language Models for Evaluating Clinical Text Anonymization
Oct 22, 2024Abstract:Sharing protected health information (PHI) is critical for furthering biomedical research. Before data can be distributed, practitioners often perform deidentification to remove any PHI contained in the text. Contemporary deidentification methods are evaluated on highly saturated datasets (tools achieve near-perfect accuracy) which may not reflect the full variability or complexity of real-world clinical text and annotating them is resource intensive, which is a barrier to real-world applications. To address this gap, we developed an adversarial approach using a large language model (LLM) to re-identify the patient corresponding to a redacted clinical note and evaluated the performance with a novel De-Identification/Re-Identification (DIRI) method. Our method uses a large language model to reidentify the patient corresponding to a redacted clinical note. We demonstrate our method on medical data from Weill Cornell Medicine anonymized with three deidentification tools: rule-based Philter and two deep-learning-based models, BiLSTM-CRF and ClinicalBERT. Although ClinicalBERT was the most effective, masking all identified PII, our tool still reidentified 9% of clinical notes Our study highlights significant weaknesses in current deidentification technologies while providing a tool for iterative development and improvement.
Leveraging Generative AI for Clinical Evidence Summarization Needs to Achieve Trustworthiness
Nov 19, 2023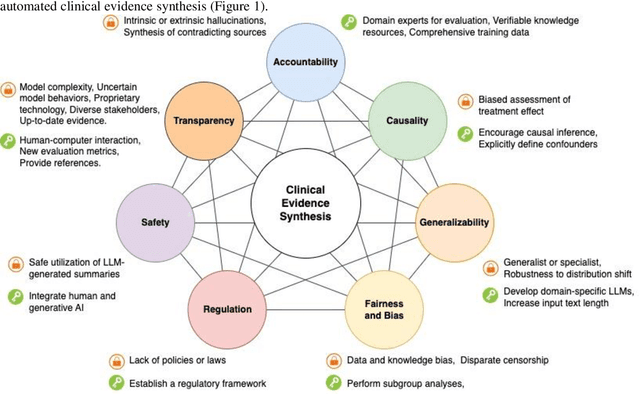
Abstract:Evidence-based medicine aims to improve the quality of healthcare by empowering medical decisions and practices with the best available evidence. The rapid growth of medical evidence, which can be obtained from various sources, poses a challenge in collecting, appraising, and synthesizing the evidential information. Recent advancements in generative AI, exemplified by large language models, hold promise in facilitating the arduous task. However, developing accountable, fair, and inclusive models remains a complicated undertaking. In this perspective, we discuss the trustworthiness of generative AI in the context of automated summarization of medical evidence.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge