Wenxue Li
RationalVLA: A Rational Vision-Language-Action Model with Dual System
Jun 12, 2025Abstract:A fundamental requirement for real-world robotic deployment is the ability to understand and respond to natural language instructions. Existing language-conditioned manipulation tasks typically assume that instructions are perfectly aligned with the environment. This assumption limits robustness and generalization in realistic scenarios where instructions may be ambiguous, irrelevant, or infeasible. To address this problem, we introduce RAtional MAnipulation (RAMA), a new benchmark that challenges models with both unseen executable instructions and defective ones that should be rejected. In RAMA, we construct a dataset with over 14,000 samples, including diverse defective instructions spanning six dimensions: visual, physical, semantic, motion, safety, and out-of-context. We further propose the Rational Vision-Language-Action model (RationalVLA). It is a dual system for robotic arms that integrates the high-level vision-language model with the low-level manipulation policy by introducing learnable latent space embeddings. This design enables RationalVLA to reason over instructions, reject infeasible commands, and execute manipulation effectively. Experiments demonstrate that RationalVLA outperforms state-of-the-art baselines on RAMA by a 14.5% higher success rate and 0.94 average task length, while maintaining competitive performance on standard manipulation tasks. Real-world trials further validate its effectiveness and robustness in practical applications. Our project page is https://irpn-eai.github.io/rationalvla.
mFabric: An Efficient and Scalable Fabric for Mixture-of-Experts Training
Jan 07, 2025



Abstract:Mixture-of-Expert (MoE) models outperform conventional models by selectively activating different subnets, named \emph{experts}, on a per-token basis. This gated computation generates dynamic communications that cannot be determined beforehand, challenging the existing GPU interconnects that remain \emph{static} during the distributed training process. In this paper, we advocate for a first-of-its-kind system, called mFabric, that unlocks topology reconfiguration \emph{during} distributed MoE training. Towards this vision, we first perform a production measurement study and show that the MoE dynamic communication pattern has \emph{strong locality}, alleviating the requirement of global reconfiguration. Based on this, we design and implement a \emph{regionally reconfigurable high-bandwidth domain} on top of existing electrical interconnects using optical circuit switching (OCS), achieving scalability while maintaining rapid adaptability. We have built a fully functional mFabric prototype with commodity hardware and a customized collective communication runtime that trains state-of-the-art MoE models with \emph{in-training} topology reconfiguration across 32 A100 GPUs. Large-scale packet-level simulations show that mFabric delivers comparable performance as the non-blocking fat-tree fabric while boosting the training cost efficiency (e.g., performance per dollar) of four representative MoE models by 1.2$\times$--1.5$\times$ and 1.9$\times$--2.3$\times$ at 100 Gbps and 400 Gbps link bandwidths, respectively.
Neighbor Does Matter: Density-Aware Contrastive Learning for Medical Semi-supervised Segmentation
Dec 27, 2024



Abstract:In medical image analysis, multi-organ semi-supervised segmentation faces challenges such as insufficient labels and low contrast in soft tissues. To address these issues, existing studies typically employ semi-supervised segmentation techniques using pseudo-labeling and consistency regularization. However, these methods mainly rely on individual data samples for training, ignoring the rich neighborhood information present in the feature space. In this work, we argue that supervisory information can be directly extracted from the geometry of the feature space. Inspired by the density-based clustering hypothesis, we propose using feature density to locate sparse regions within feature clusters. Our goal is to increase intra-class compactness by addressing sparsity issues. To achieve this, we propose a Density-Aware Contrastive Learning (DACL) strategy, pushing anchored features in sparse regions towards cluster centers approximated by high-density positive samples, resulting in more compact clusters. Specifically, our method constructs density-aware neighbor graphs using labeled and unlabeled data samples to estimate feature density and locate sparse regions. We also combine label-guided co-training with density-guided geometric regularization to form complementary supervision for unlabeled data. Experiments on the Multi-Organ Segmentation Challenge dataset demonstrate that our proposed method outperforms state-of-the-art methods, highlighting its efficacy in medical image segmentation tasks.
SAM2-UNet: Segment Anything 2 Makes Strong Encoder for Natural and Medical Image Segmentation
Aug 16, 2024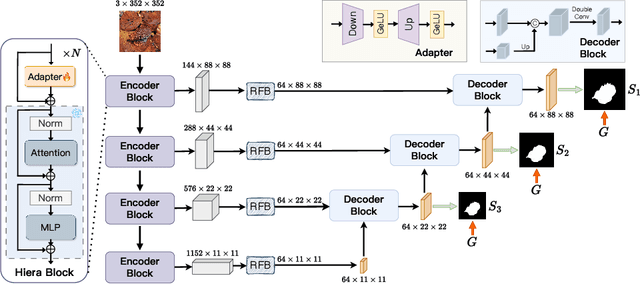



Abstract:Image segmentation plays an important role in vision understanding. Recently, the emerging vision foundation models continuously achieved superior performance on various tasks. Following such success, in this paper, we prove that the Segment Anything Model 2 (SAM2) can be a strong encoder for U-shaped segmentation models. We propose a simple but effective framework, termed SAM2-UNet, for versatile image segmentation. Specifically, SAM2-UNet adopts the Hiera backbone of SAM2 as the encoder, while the decoder uses the classic U-shaped design. Additionally, adapters are inserted into the encoder to allow parameter-efficient fine-tuning. Preliminary experiments on various downstream tasks, such as camouflaged object detection, salient object detection, marine animal segmentation, mirror detection, and polyp segmentation, demonstrate that our SAM2-UNet can simply beat existing specialized state-of-the-art methods without bells and whistles. Project page: \url{https://github.com/WZH0120/SAM2-UNet}.
TP-DRSeg: Improving Diabetic Retinopathy Lesion Segmentation with Explicit Text-Prompts Assisted SAM
Jun 22, 2024



Abstract:Recent advances in large foundation models, such as the Segment Anything Model (SAM), have demonstrated considerable promise across various tasks. Despite their progress, these models still encounter challenges in specialized medical image analysis, especially in recognizing subtle inter-class differences in Diabetic Retinopathy (DR) lesion segmentation. In this paper, we propose a novel framework that customizes SAM for text-prompted DR lesion segmentation, termed TP-DRSeg. Our core idea involves exploiting language cues to inject medical prior knowledge into the vision-only segmentation network, thereby combining the advantages of different foundation models and enhancing the credibility of segmentation. Specifically, to unleash the potential of vision-language models in the recognition of medical concepts, we propose an explicit prior encoder that transfers implicit medical concepts into explicit prior knowledge, providing explainable clues to excavate low-level features associated with lesions. Furthermore, we design a prior-aligned injector to inject explicit priors into the segmentation process, which can facilitate knowledge sharing across multi-modality features and allow our framework to be trained in a parameter-efficient fashion. Experimental results demonstrate the superiority of our framework over other traditional models and foundation model variants.
Diffusion Model Driven Test-Time Image Adaptation for Robust Skin Lesion Classification
May 18, 2024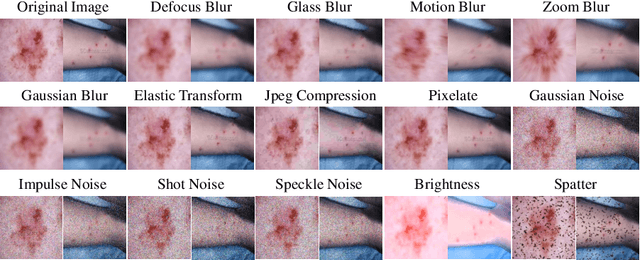

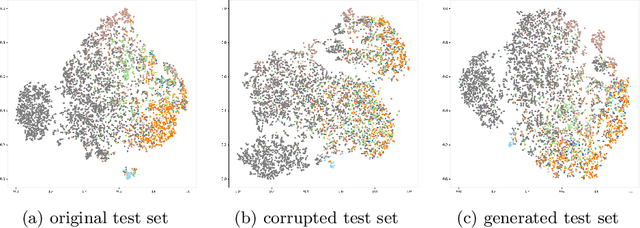

Abstract:Deep learning-based diagnostic systems have demonstrated potential in skin disease diagnosis. However, their performance can easily degrade on test domains due to distribution shifts caused by input-level corruptions, such as imaging equipment variability, brightness changes, and image blur. This will reduce the reliability of model deployment in real-world scenarios. Most existing solutions focus on adapting the source model through retraining on different target domains. Although effective, this retraining process is sensitive to the amount of data and the hyperparameter configuration for optimization. In this paper, we propose a test-time image adaptation method to enhance the accuracy of the model on test data by simultaneously updating and predicting test images. We modify the target test images by projecting them back to the source domain using a diffusion model. Specifically, we design a structure guidance module that adds refinement operations through low-pass filtering during reverse sampling, regularizing the diffusion to preserve structural information. Additionally, we introduce a self-ensembling scheme automatically adjusts the reliance on adapted and unadapted inputs, enhancing adaptation robustness by rejecting inappropriate generative modeling results. To facilitate this study, we constructed the ISIC2019-C and Dermnet-C corruption robustness evaluation benchmarks. Extensive experiments on the proposed benchmarks demonstrate that our method makes the classifier more robust across various corruptions, architectures, and data regimes. Our datasets and code will be available at \url{https://github.com/minghu0830/Skin-TTA_Diffusion}.
Semi- and Weakly-Supervised Learning for Mammogram Mass Segmentation with Limited Annotations
Mar 14, 2024



Abstract:Accurate identification of breast masses is crucial in diagnosing breast cancer; however, it can be challenging due to their small size and being camouflaged in surrounding normal glands. Worse still, it is also expensive in clinical practice to obtain adequate pixel-wise annotations for training deep neural networks. To overcome these two difficulties with one stone, we propose a semi- and weakly-supervised learning framework for mass segmentation that utilizes limited strongly-labeled samples and sufficient weakly-labeled samples to achieve satisfactory performance. The framework consists of an auxiliary branch to exclude lesion-irrelevant background areas, a segmentation branch for final prediction, and a spatial prompting module to integrate the complementary information of the two branches. We further disentangle encoded obscure features into lesion-related and others to boost performance. Experiments on CBIS-DDSM and INbreast datasets demonstrate the effectiveness of our method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge