Tommy Löfstedt
Beyond a Single Mode: GAN Ensembles for Diverse Medical Data Generation
Mar 31, 2025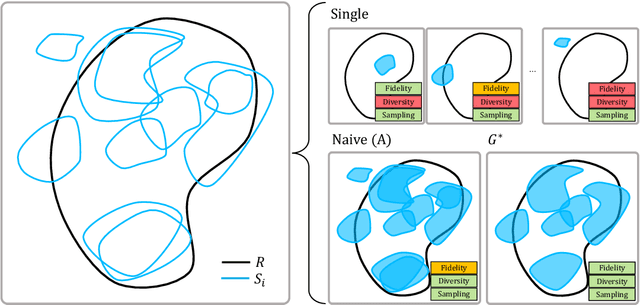

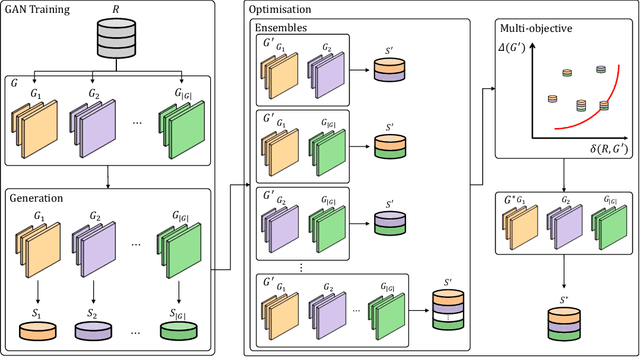
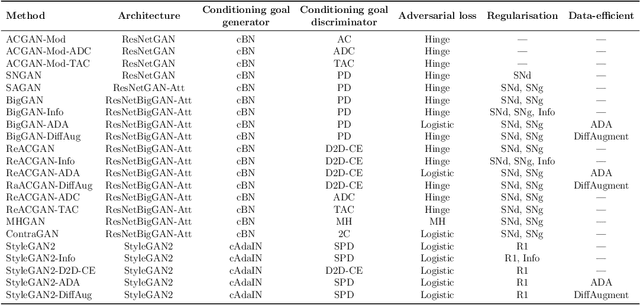
Abstract:The advancement of generative AI, particularly in medical imaging, confronts the trilemma of ensuring high fidelity, diversity, and efficiency in synthetic data generation. While Generative Adversarial Networks (GANs) have shown promise across various applications, they still face challenges like mode collapse and insufficient coverage of real data distributions. This work explores the use of GAN ensembles to overcome these limitations, specifically in the context of medical imaging. By solving a multi-objective optimisation problem that balances fidelity and diversity, we propose a method for selecting an optimal ensemble of GANs tailored for medical data. The selected ensemble is capable of generating diverse synthetic medical images that are representative of true data distributions and computationally efficient. Each model in the ensemble brings a unique contribution, ensuring minimal redundancy. We conducted a comprehensive evaluation using three distinct medical datasets, testing 22 different GAN architectures with various loss functions and regularisation techniques. By sampling models at different training epochs, we crafted 110 unique configurations. The results highlight the capability of GAN ensembles to enhance the quality and utility of synthetic medical images, thereby improving the efficacy of downstream tasks such as diagnostic modelling.
Provable Reduction in Communication Rounds for Non-Smooth Convex Federated Learning
Mar 27, 2025

Abstract:Multiple local steps are key to communication-efficient federated learning. However, theoretical guarantees for such algorithms, without data heterogeneity-bounding assumptions, have been lacking in general non-smooth convex problems. Leveraging projection-efficient optimization methods, we propose FedMLS, a federated learning algorithm with provable improvements from multiple local steps. FedMLS attains an $\epsilon$-suboptimal solution in $\mathcal{O}(1/\epsilon)$ communication rounds, requiring a total of $\mathcal{O}(1/\epsilon^2)$ stochastic subgradient oracle calls.
Using Synthetic Images to Augment Small Medical Image Datasets
Mar 02, 2025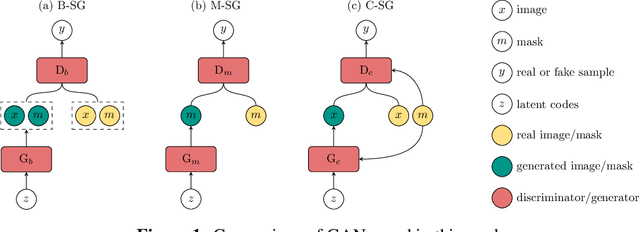

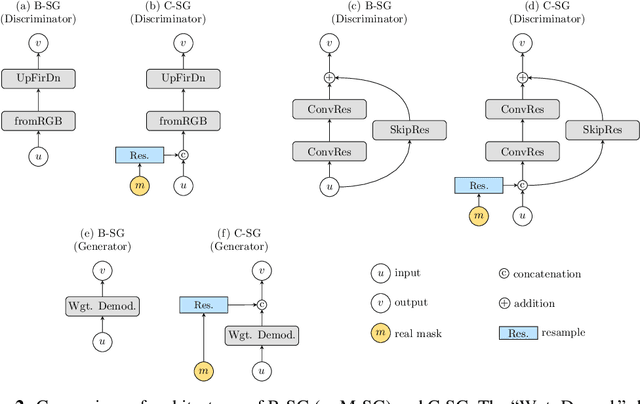

Abstract:Recent years have witnessed a growing academic and industrial interest in deep learning (DL) for medical imaging. To perform well, DL models require very large labeled datasets. However, most medical imaging datasets are small, with a limited number of annotated samples. The reason they are small is usually because delineating medical images is time-consuming and demanding for oncologists. There are various techniques that can be used to augment a dataset, for example, to apply affine transformations or elastic transformations to available images, or to add synthetic images generated by a Generative Adversarial Network (GAN). In this work, we have developed a novel conditional variant of a current GAN method, the StyleGAN2, to generate multi-modal high-resolution medical images with the purpose to augment small medical imaging datasets with these synthetic images. We use the synthetic and real images from six datasets to train models for the downstream task of semantic segmentation. The quality of the generated medical images and the effect of this augmentation on the segmentation performance were evaluated afterward. Finally, the results indicate that the downstream segmentation models did not benefit from the generated images. Further work and analyses are required to establish how this augmentation affects the segmentation performance.
A Cost-Aware Approach to Adversarial Robustness in Neural Networks
Sep 11, 2024



Abstract:Considering the growing prominence of production-level AI and the threat of adversarial attacks that can evade a model at run-time, evaluating the robustness of models to these evasion attacks is of critical importance. Additionally, testing model changes likely means deploying the models to (e.g. a car or a medical imaging device), or a drone to see how it affects performance, making un-tested changes a public problem that reduces development speed, increases cost of development, and makes it difficult (if not impossible) to parse cause from effect. In this work, we used survival analysis as a cloud-native, time-efficient and precise method for predicting model performance in the presence of adversarial noise. For neural networks in particular, the relationships between the learning rate, batch size, training time, convergence time, and deployment cost are highly complex, so researchers generally rely on benchmark datasets to assess the ability of a model to generalize beyond the training data. To address this, we propose using accelerated failure time models to measure the effect of hardware choice, batch size, number of epochs, and test-set accuracy by using adversarial attacks to induce failures on a reference model architecture before deploying the model to the real world. We evaluate several GPU types and use the Tree Parzen Estimator to maximize model robustness and minimize model run-time simultaneously. This provides a way to evaluate the model and optimise it in a single step, while simultaneously allowing us to model the effect of model parameters on training time, prediction time, and accuracy. Using this technique, we demonstrate that newer, more-powerful hardware does decrease the training time, but with a monetary and power cost that far outpaces the marginal gains in accuracy.
A Correlation- and Mean-Aware Loss Function and Benchmarking Framework to Improve GAN-based Tabular Data Synthesis
May 27, 2024Abstract:Advancements in science rely on data sharing. In medicine, where personal data are often involved, synthetic tabular data generated by generative adversarial networks (GANs) offer a promising avenue. However, existing GANs struggle to capture the complexities of real-world tabular data, which often contain a mix of continuous and categorical variables with potential imbalances and dependencies. We propose a novel correlation- and mean-aware loss function designed to address these challenges as a regularizer for GANs. To ensure a rigorous evaluation, we establish a comprehensive benchmarking framework using ten real-world datasets and eight established tabular GAN baselines. The proposed loss function demonstrates statistically significant improvements over existing methods in capturing the true data distribution, significantly enhancing the quality of synthetic data generated with GANs. The benchmarking framework shows that the enhanced synthetic data quality leads to improved performance in downstream machine learning (ML) tasks, ultimately paving the way for easier data sharing.
Synthesizing multi-log grasp poses
Mar 18, 2024Abstract:Multi-object grasping is a challenging task. It is important for energy and cost-efficient operation of industrial crane manipulators, such as those used to collect tree logs off the forest floor and onto forest machines. In this work, we used synthetic data from physics simulations to explore how data-driven modeling can be used to infer multi-object grasp poses from images. We showed that convolutional neural networks can be trained specifically for synthesizing multi-object grasps. Using RGB-Depth images and instance segmentation masks as input, a U-Net model outputs grasp maps with corresponding grapple orientation and opening width. Given an observation of a pile of logs, the model can be used to synthesize and rate the possible grasp poses and select the most suitable one, with the possibility to respect changing operational constraints such as lift capacity and reach. When tested on previously unseen data, the proposed model found successful grasp poses with an accuracy of 95%.
A Systematic Approach to Robustness Modelling for Deep Convolutional Neural Networks
Jan 24, 2024
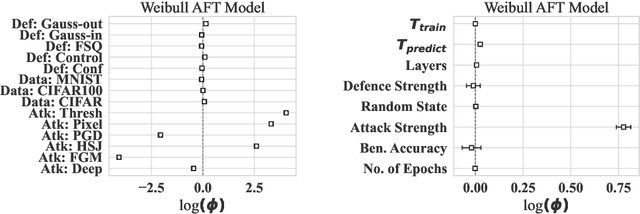
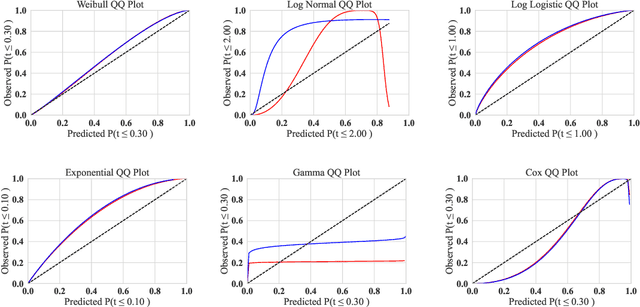


Abstract:Convolutional neural networks have shown to be widely applicable to a large number of fields when large amounts of labelled data are available. The recent trend has been to use models with increasingly larger sets of tunable parameters to increase model accuracy, reduce model loss, or create more adversarially robust models -- goals that are often at odds with one another. In particular, recent theoretical work raises questions about the ability for even larger models to generalize to data outside of the controlled train and test sets. As such, we examine the role of the number of hidden layers in the ResNet model, demonstrated on the MNIST, CIFAR10, CIFAR100 datasets. We test a variety of parameters including the size of the model, the floating point precision, and the noise level of both the training data and the model output. To encapsulate the model's predictive power and computational cost, we provide a method that uses induced failures to model the probability of failure as a function of time and relate that to a novel metric that allows us to quickly determine whether or not the cost of training a model outweighs the cost of attacking it. Using this approach, we are able to approximate the expected failure rate using a small number of specially crafted samples rather than increasingly larger benchmark datasets. We demonstrate the efficacy of this technique on both the MNIST and CIFAR10 datasets using 8-, 16-, 32-, and 64-bit floating-point numbers, various data pre-processing techniques, and several attacks on five configurations of the ResNet model. Then, using empirical measurements, we examine the various trade-offs between cost, robustness, latency, and reliability to find that larger models do not significantly aid in adversarial robustness despite costing significantly more to train.
LatentAugment: Data Augmentation via Guided Manipulation of GAN's Latent Space
Jul 21, 2023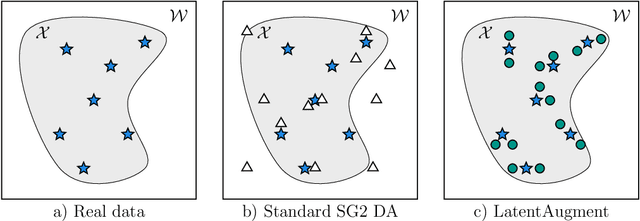
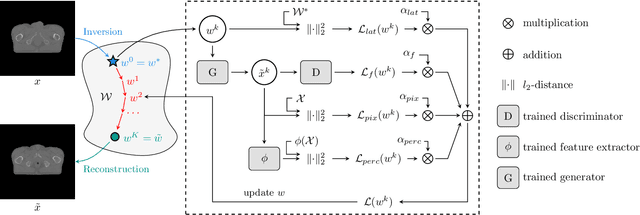
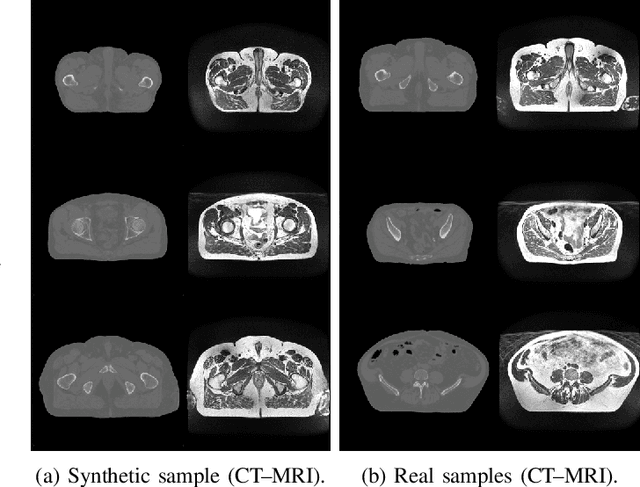
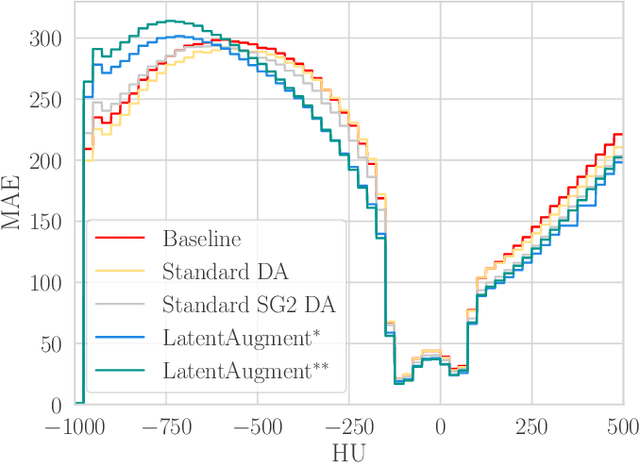
Abstract:Data Augmentation (DA) is a technique to increase the quantity and diversity of the training data, and by that alleviate overfitting and improve generalisation. However, standard DA produces synthetic data for augmentation with limited diversity. Generative Adversarial Networks (GANs) may unlock additional information in a dataset by generating synthetic samples having the appearance of real images. However, these models struggle to simultaneously address three key requirements: fidelity and high-quality samples; diversity and mode coverage; and fast sampling. Indeed, GANs generate high-quality samples rapidly, but have poor mode coverage, limiting their adoption in DA applications. We propose LatentAugment, a DA strategy that overcomes the low diversity of GANs, opening up for use in DA applications. Without external supervision, LatentAugment modifies latent vectors and moves them into latent space regions to maximise the synthetic images' diversity and fidelity. It is also agnostic to the dataset and the downstream task. A wide set of experiments shows that LatentAugment improves the generalisation of a deep model translating from MRI-to-CT beating both standard DA as well GAN-based sampling. Moreover, still in comparison with GAN-based sampling, LatentAugment synthetic samples show superior mode coverage and diversity. Code is available at: https://github.com/ltronchin/LatentAugment.
QU-BraTS: MICCAI BraTS 2020 Challenge on Quantifying Uncertainty in Brain Tumor Segmentation -- Analysis of Ranking Metrics and Benchmarking Results
Dec 19, 2021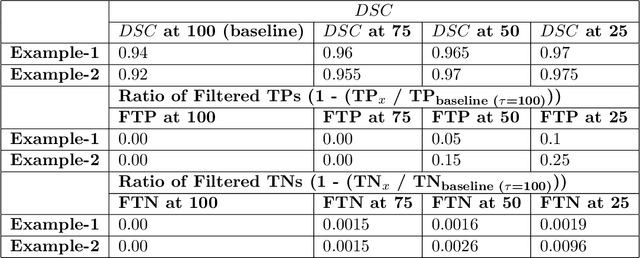
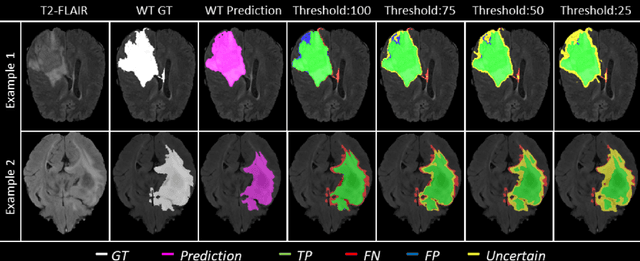
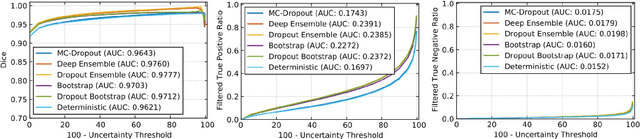
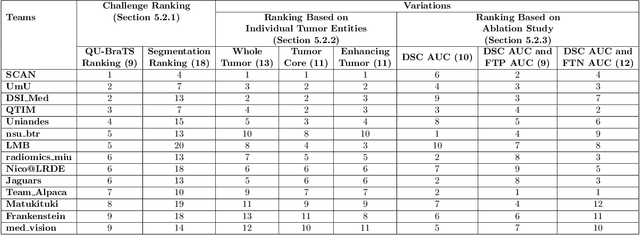
Abstract:Deep learning (DL) models have provided the state-of-the-art performance in a wide variety of medical imaging benchmarking challenges, including the Brain Tumor Segmentation (BraTS) challenges. However, the task of focal pathology multi-compartment segmentation (e.g., tumor and lesion sub-regions) is particularly challenging, and potential errors hinder the translation of DL models into clinical workflows. Quantifying the reliability of DL model predictions in the form of uncertainties, could enable clinical review of the most uncertain regions, thereby building trust and paving the way towards clinical translation. Recently, a number of uncertainty estimation methods have been introduced for DL medical image segmentation tasks. Developing metrics to evaluate and compare the performance of uncertainty measures will assist the end-user in making more informed decisions. In this study, we explore and evaluate a metric developed during the BraTS 2019-2020 task on uncertainty quantification (QU-BraTS), and designed to assess and rank uncertainty estimates for brain tumor multi-compartment segmentation. This metric (1) rewards uncertainty estimates that produce high confidence in correct assertions, and those that assign low confidence levels at incorrect assertions, and (2) penalizes uncertainty measures that lead to a higher percentages of under-confident correct assertions. We further benchmark the segmentation uncertainties generated by 14 independent participating teams of QU-BraTS 2020, all of which also participated in the main BraTS segmentation task. Overall, our findings confirm the importance and complementary value that uncertainty estimates provide to segmentation algorithms, and hence highlight the need for uncertainty quantification in medical image analyses. Our evaluation code is made publicly available at https://github.com/RagMeh11/QU-BraTS.
A Data-Adaptive Loss Function for Incomplete Data and Incremental Learning in Semantic Image Segmentation
Apr 22, 2021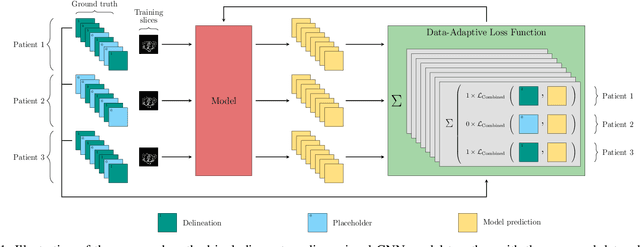
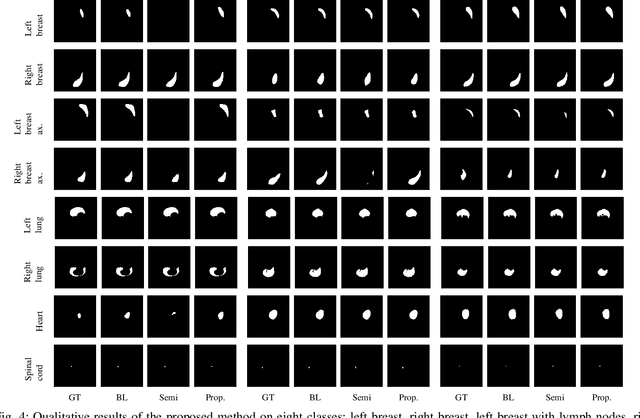
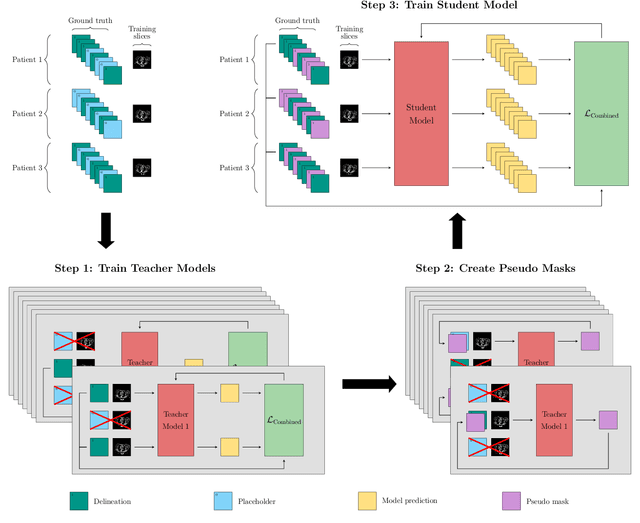
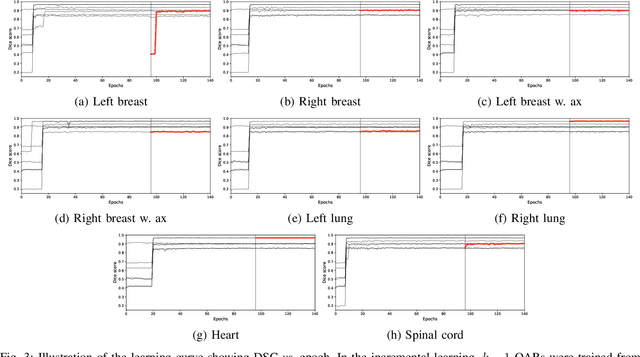

Abstract:In the last years, deep learning has dramatically improved the performances in a variety of medical image analysis applications. Among different types of deep learning models, convolutional neural networks have been among the most successful and they have been used in many applications in medical imaging. Training deep convolutional neural networks often requires large amounts of image data to generalize well to new unseen images. It is often time-consuming and expensive to collect large amounts of data in the medical image domain due to expensive imaging systems, and the need for experts to manually make ground truth annotations. A potential problem arises if new structures are added when a decision support system is already deployed and in use. Since the field of radiation therapy is constantly developing, the new structures would also have to be covered by the decision support system. In the present work, we propose a novel loss function, that adapts to the available data in order to utilize all available data, even when some have missing annotations. We demonstrate that the proposed loss function also works well in an incremental learning setting, where it can automatically incorporate new structures as they appear. Experiments on a large in-house data set show that the proposed method performs on par with baseline models, while greatly reducing the training time.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge