Paolo Soda
Learning from Limited and Incomplete Data: A Multimodal Framework for Predicting Pathological Response in NSCLC
Mar 16, 2026Abstract:Major pathological response (pR) following neoadjuvant therapy is a clinically meaningful endpoint in non-small cell lung cancer, strongly associated with improved survival. However, accurate preoperative prediction of pR remains challenging, particularly in real-world clinical settings characterized by limited data availability and incomplete clinical profiles. In this study, we propose a multimodal deep learning framework designed to address these constraints by integrating foundation model-based CT feature extraction with a missing-aware architecture for clinical variables. This approach enables robust learning from small cohorts while explicitly modeling missing clinical information, without relying on conventional imputation strategies. A weighted fusion mechanism is employed to leverage the complementary contributions of imaging and clinical modalities, yielding a multimodal model that consistently outperforms both unimodal imaging and clinical baselines. These findings underscore the added value of integrating heterogeneous data sources and highlight the potential of multimodal, missing-aware systems to support pR prediction under realistic clinical conditions.
A Systematic Benchmark of GAN Architectures for MRI-to-CT Synthesis
Mar 13, 2026Abstract:The translation from Magnetic resonance imaging (MRI) to Computed tomography (CT) has been proposed as an effective solution to facilitate MRI-only clinical workflows while limiting exposure to ionizing radiation. Although numerous Generative Adversarial Network (GAN) architectures have been proposed for MRI-to-CT translation, systematic and fair comparisons across heterogeneous models remain limited. We present a comprehensive benchmark of ten GAN architectures evaluated on the SynthRAD2025 dataset across three anatomical districts (abdomen, thorax, head-and-neck). All models were trained under a unified validation protocol with identical preprocessing and optimization settings. Performance was assessed using complementary metrics capturing voxel-wise accuracy, structural fidelity, perceptual quality, and distribution-level realism, alongside an analysis of computational complexity. Supervised Paired models consistently outperformed Unpaired approaches, confirming the importance of voxel-wise supervision. Pix2Pix achieved the most balanced performance across districts while maintaining a favorable quality-to-complexity trade-off. Multi-district training improved structural robustness, whereas intra-district training maximized voxel-wise fidelity. This benchmark provides quantitative and computational guidance for model selection in MRI-only radiotherapy workflows and establishes a reproducible framework for future comparative studies. To ensure the reproducibility of our experiments we make our code public, together with the overall results, at the following link:https://github.com/arco-group/MRI_TO_CT.git
Retrieval-Augmented Anatomical Guidance for Text-to-CT Generation
Mar 09, 2026Abstract:Text-conditioned generative models for volumetric medical imaging provide semantic control but lack explicit anatomical guidance, often resulting in outputs that are spatially ambiguous or anatomically inconsistent. In contrast, structure-driven methods ensure strong anatomical consistency but typically assume access to ground-truth annotations, which are unavailable when the target image is to be synthesized. We propose a retrieval-augmented approach for Text-to-CT generation that integrates semantic and anatomical information under a realistic inference setting. Given a radiology report, our method retrieves a semantically related clinical case using a 3D vision-language encoder and leverages its associated anatomical annotation as a structural proxy. This proxy is injected into a text-conditioned latent diffusion model via a ControlNet branch, providing coarse anatomical guidance while maintaining semantic flexibility. Experiments on the CT-RATE dataset show that retrieval-augmented generation improves image fidelity and clinical consistency compared to text-only baselines, while additionally enabling explicit spatial controllability, a capability inherently absent in such approaches. Further analysis highlights the importance of retrieval quality, with semantically aligned proxies yielding consistent gains across all evaluation axes. This work introduces a principled and scalable mechanism to bridge semantic conditioning and anatomical plausibility in volumetric medical image synthesis. Code will be released.
Hybrid Quantum Neural Network for Multivariate Clinical Time Series Forecasting
Mar 09, 2026Abstract:Forecasting physiological signals can support proactive monitoring and timely clinical intervention by anticipating critical changes in patient status. In this work, we address multivariate multi-horizon forecasting of physiological time series by jointly predicting heart rate, oxygen saturation, pulse rate, and respiratory rate at forecasting horizons of 15, 30, and 60 seconds. We propose a hybrid quantum-classical architecture that integrates a Variational Quantum Circuit (VQC) within a recurrent neural backbone. A GRU encoder summarizes the historical observation window into a latent representation, which is then projected into quantum angles used to parameterize the VQC. The quantum layer acts as a learnable non-linear feature mixer, modeling cross-variable interactions before the final prediction stage. We evaluate the proposed approach on the BIDMC PPG and Respiration dataset under a Leave-One-Patient-Out protocol. The results show competitive accuracy compared with classical and deep learning baselines, together with greater robustness to noise and missing inputs. These findings suggest that hybrid quantum layers can provide useful inductive biases for physiological time series forecasting in small-cohort clinical settings.
Concept-Enhanced Multimodal RAG: Towards Interpretable and Accurate Radiology Report Generation
Feb 17, 2026Abstract:Radiology Report Generation (RRG) through Vision-Language Models (VLMs) promises to reduce documentation burden, improve reporting consistency, and accelerate clinical workflows. However, their clinical adoption remains limited by the lack of interpretability and the tendency to hallucinate findings misaligned with imaging evidence. Existing research typically treats interpretability and accuracy as separate objectives, with concept-based explainability techniques focusing primarily on transparency, while Retrieval-Augmented Generation (RAG) methods targeting factual grounding through external retrieval. We present Concept-Enhanced Multimodal RAG (CEMRAG), a unified framework that decomposes visual representations into interpretable clinical concepts and integrates them with multimodal RAG. This approach exploits enriched contextual prompts for RRG, improving both interpretability and factual accuracy. Experiments on MIMIC-CXR and IU X-Ray across multiple VLM architectures, training regimes, and retrieval configurations demonstrate consistent improvements over both conventional RAG and concept-only baselines on clinical accuracy metrics and standard NLP measures. These results challenge the assumed trade-off between interpretability and performance, showing that transparent visual concepts can enhance rather than compromise diagnostic accuracy in medical VLMs. Our modular design decomposes interpretability into visual transparency and structured language model conditioning, providing a principled pathway toward clinically trustworthy AI-assisted radiology.
Handling Missing Modalities in Multimodal Survival Prediction for Non-Small Cell Lung Cancer
Jan 15, 2026Abstract:Accurate survival prediction in Non-Small Cell Lung Cancer (NSCLC) requires the integration of heterogeneous clinical, radiological, and histopathological information. While Multimodal Deep Learning (MDL) offers a promises for precision prognosis and survival prediction, its clinical applicability is severely limited by small cohort sizes and the presence of missing modalities, often forcing complete-case filtering or aggressive imputation. In this work, we present a missing-aware multimodal survival framework that integrates Computed Tomography (CT), Whole-Slide Histopathology (WSI) Images, and structured clinical variables for overall survival modeling in unresectable stage II-III NSCLC. By leveraging Foundation Models (FM) for modality-specific feature extraction and a missing-aware encoding strategy, the proposed approach enables intermediate multimodal fusion under naturally incomplete modality profiles. The proposed architecture is resilient to missing modalities by design, allowing the model to utilize all available data without being forced to drop patients during training or inference. Experimental results demonstrate that intermediate fusion consistently outperforms unimodal baselines as well as early and late fusion strategies, with the strongest performance achieved by the fusion of WSI and clinical modalities (73.30 C-index). Further analyses of modality importance reveal an adaptive behavior in which less informative modalities, i.e., CT modality, are automatically down-weighted and contribute less to the final survival prediction.
Context-Gated Cross-Modal Perception with Visual Mamba for PET-CT Lung Tumor Segmentation
Oct 31, 2025Abstract:Accurate lung tumor segmentation is vital for improving diagnosis and treatment planning, and effectively combining anatomical and functional information from PET and CT remains a major challenge. In this study, we propose vMambaX, a lightweight multimodal framework integrating PET and CT scan images through a Context-Gated Cross-Modal Perception Module (CGM). Built on the Visual Mamba architecture, vMambaX adaptively enhances inter-modality feature interaction, emphasizing informative regions while suppressing noise. Evaluated on the PCLT20K dataset, the model outperforms baseline models while maintaining lower computational complexity. These results highlight the effectiveness of adaptive cross-modal gating for multimodal tumor segmentation and demonstrate the potential of vMambaX as an efficient and scalable framework for advanced lung cancer analysis. The code is available at https://github.com/arco-group/vMambaX.
SPARSE Data, Rich Results: Few-Shot Semi-Supervised Learning via Class-Conditioned Image Translation
Aug 08, 2025Abstract:Deep learning has revolutionized medical imaging, but its effectiveness is severely limited by insufficient labeled training data. This paper introduces a novel GAN-based semi-supervised learning framework specifically designed for low labeled-data regimes, evaluated across settings with 5 to 50 labeled samples per class. Our approach integrates three specialized neural networks -- a generator for class-conditioned image translation, a discriminator for authenticity assessment and classification, and a dedicated classifier -- within a three-phase training framework. The method alternates between supervised training on limited labeled data and unsupervised learning that leverages abundant unlabeled images through image-to-image translation rather than generation from noise. We employ ensemble-based pseudo-labeling that combines confidence-weighted predictions from the discriminator and classifier with temporal consistency through exponential moving averaging, enabling reliable label estimation for unlabeled data. Comprehensive evaluation across eleven MedMNIST datasets demonstrates that our approach achieves statistically significant improvements over six state-of-the-art GAN-based semi-supervised methods, with particularly strong performance in the extreme 5-shot setting where the scarcity of labeled data is most challenging. The framework maintains its superiority across all evaluated settings (5, 10, 20, and 50 shots per class). Our approach offers a practical solution for medical imaging applications where annotation costs are prohibitive, enabling robust classification performance even with minimal labeled data. Code is available at https://github.com/GuidoManni/SPARSE.
Sample-Aware Test-Time Adaptation for Medical Image-to-Image Translation
Aug 01, 2025Abstract:Image-to-image translation has emerged as a powerful technique in medical imaging, enabling tasks such as image denoising and cross-modality conversion. However, it suffers from limitations in handling out-of-distribution samples without causing performance degradation. To address this limitation, we propose a novel Test-Time Adaptation (TTA) framework that dynamically adjusts the translation process based on the characteristics of each test sample. Our method introduces a Reconstruction Module to quantify the domain shift and a Dynamic Adaptation Block that selectively modifies the internal features of a pretrained translation model to mitigate the shift without compromising the performance on in-distribution samples that do not require adaptation. We evaluate our approach on two medical image-to-image translation tasks: low-dose CT denoising and T1 to T2 MRI translation, showing consistent improvements over both the baseline translation model without TTA and prior TTA methods. Our analysis highlights the limitations of the state-of-the-art that uniformly apply the adaptation to both out-of-distribution and in-distribution samples, demonstrating that dynamic, sample-specific adjustment offers a promising path to improve model resilience in real-world scenarios. The code is available at: https://github.com/cosbidev/Sample-Aware_TTA.
Lesion-Aware Generative Artificial Intelligence for Virtual Contrast-Enhanced Mammography in Breast Cancer
May 05, 2025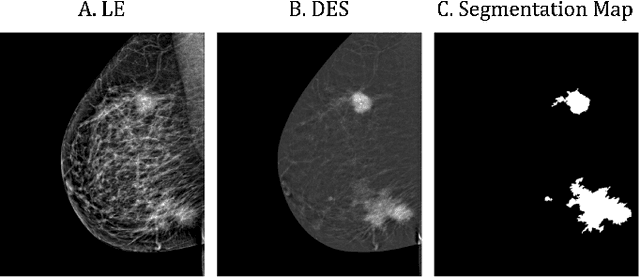
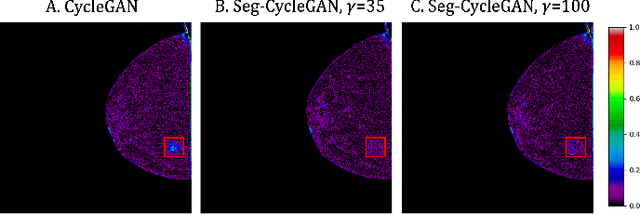
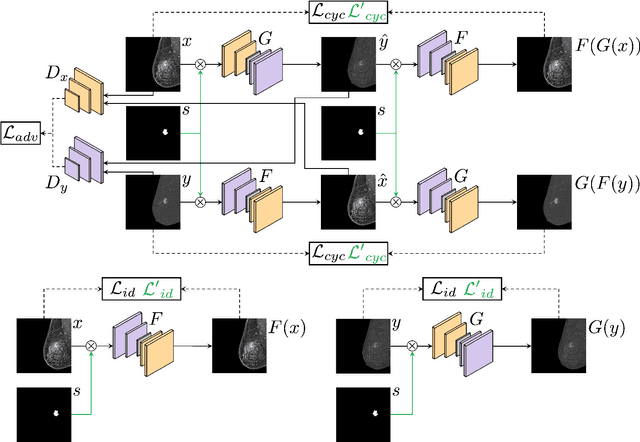
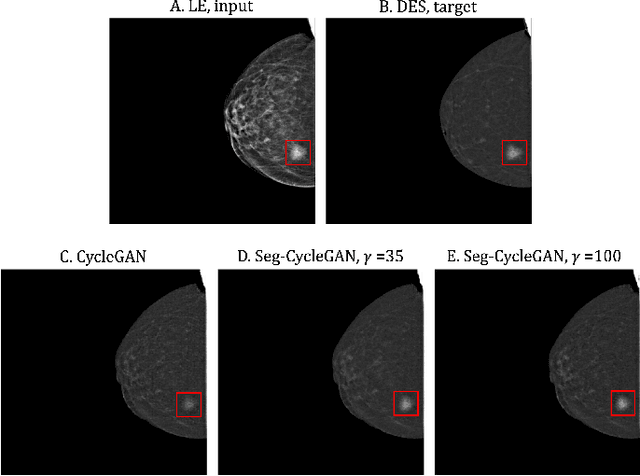

Abstract:Contrast-Enhanced Spectral Mammography (CESM) is a dual-energy mammographic technique that improves lesion visibility through the administration of an iodinated contrast agent. It acquires both a low-energy image, comparable to standard mammography, and a high-energy image, which are then combined to produce a dual-energy subtracted image highlighting lesion contrast enhancement. While CESM offers superior diagnostic accuracy compared to standard mammography, its use entails higher radiation exposure and potential side effects associated with the contrast medium. To address these limitations, we propose Seg-CycleGAN, a generative deep learning framework for Virtual Contrast Enhancement in CESM. The model synthesizes high-fidelity dual-energy subtracted images from low-energy images, leveraging lesion segmentation maps to guide the generative process and improve lesion reconstruction. Building upon the standard CycleGAN architecture, Seg-CycleGAN introduces localized loss terms focused on lesion areas, enhancing the synthesis of diagnostically relevant regions. Experiments on the CESM@UCBM dataset demonstrate that Seg-CycleGAN outperforms the baseline in terms of PSNR and SSIM, while maintaining competitive MSE and VIF. Qualitative evaluations further confirm improved lesion fidelity in the generated images. These results suggest that segmentation-aware generative models offer a viable pathway toward contrast-free CESM alternatives.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge