Francesco Di Feola
Sample-Aware Test-Time Adaptation for Medical Image-to-Image Translation
Aug 01, 2025Abstract:Image-to-image translation has emerged as a powerful technique in medical imaging, enabling tasks such as image denoising and cross-modality conversion. However, it suffers from limitations in handling out-of-distribution samples without causing performance degradation. To address this limitation, we propose a novel Test-Time Adaptation (TTA) framework that dynamically adjusts the translation process based on the characteristics of each test sample. Our method introduces a Reconstruction Module to quantify the domain shift and a Dynamic Adaptation Block that selectively modifies the internal features of a pretrained translation model to mitigate the shift without compromising the performance on in-distribution samples that do not require adaptation. We evaluate our approach on two medical image-to-image translation tasks: low-dose CT denoising and T1 to T2 MRI translation, showing consistent improvements over both the baseline translation model without TTA and prior TTA methods. Our analysis highlights the limitations of the state-of-the-art that uniformly apply the adaptation to both out-of-distribution and in-distribution samples, demonstrating that dynamic, sample-specific adjustment offers a promising path to improve model resilience in real-world scenarios. The code is available at: https://github.com/cosbidev/Sample-Aware_TTA.
Texture-Aware StarGAN for CT data harmonisation
Mar 19, 2025Abstract:Computed Tomography (CT) plays a pivotal role in medical diagnosis; however, variability across reconstruction kernels hinders data-driven approaches, such as deep learning models, from achieving reliable and generalized performance. To this end, CT data harmonization has emerged as a promising solution to minimize such non-biological variances by standardizing data across different sources or conditions. In this context, Generative Adversarial Networks (GANs) have proved to be a powerful framework for harmonization, framing it as a style-transfer problem. However, GAN-based approaches still face limitations in capturing complex relationships within the images, which are essential for effective harmonization. In this work, we propose a novel texture-aware StarGAN for CT data harmonization, enabling one-to-many translations across different reconstruction kernels. Although the StarGAN model has been successfully applied in other domains, its potential for CT data harmonization remains unexplored. Furthermore, our approach introduces a multi-scale texture loss function that embeds texture information across different spatial and angular scales into the harmonization process, effectively addressing kernel-induced texture variations. We conducted extensive experimentation on a publicly available dataset, utilizing a total of 48667 chest CT slices from 197 patients distributed over three different reconstruction kernels, demonstrating the superiority of our method over the baseline StarGAN.
Whole-Body Image-to-Image Translation for a Virtual Scanner in a Healthcare Digital Twin
Mar 18, 2025



Abstract:Generating positron emission tomography (PET) images from computed tomography (CT) scans via deep learning offers a promising pathway to reduce radiation exposure and costs associated with PET imaging, improving patient care and accessibility to functional imaging. Whole-body image translation presents challenges due to anatomical heterogeneity, often limiting generalized models. We propose a framework that segments whole-body CT images into four regions-head, trunk, arms, and legs-and uses district-specific Generative Adversarial Networks (GANs) for tailored CT-to-PET translation. Synthetic PET images from each region are stitched together to reconstruct the whole-body scan. Comparisons with a baseline non-segmented GAN and experiments with Pix2Pix and CycleGAN architectures tested paired and unpaired scenarios. Quantitative evaluations at district, whole-body, and lesion levels demonstrated significant improvements with our district-specific GANs. Pix2Pix yielded superior metrics, ensuring precise, high-quality image synthesis. By addressing anatomical heterogeneity, this approach achieves state-of-the-art results in whole-body CT-to-PET translation. This methodology supports healthcare Digital Twins by enabling accurate virtual PET scans from CT data, creating virtual imaging representations to monitor, predict, and optimize health outcomes.
MedCoDi-M: A Multi-Prompt Foundation Model for Multimodal Medical Data Generation
Jan 08, 2025



Abstract:Artificial Intelligence is revolutionizing medical practice, enhancing diagnostic accuracy and healthcare delivery. However, its adaptation in medical settings still faces significant challenges, related to data availability and privacy constraints. Synthetic data has emerged as a promising solution to mitigate these issues, addressing data scarcity while preserving privacy. Recently, Latent Diffusion Models have emerged as a powerful tool for generating high-quality synthetic data. Meanwhile, the integration of different modalities has gained interest, emphasizing the need of models capable of handle multimodal medical data.Existing approaches struggle to integrate complementary information and lack the ability to generate modalities simultaneously. To address this challenge, we present MedCoDi-M, a 6.77-billion-parameter model, designed for multimodal medical data generation, that, following Foundation Model paradigm, exploits contrastive learning and large quantity of data to build a shared latent space which capture the relationships between different data modalities. Further, we introduce the Multi-Prompt training technique, which significantly boosts MedCoDi-M's generation under different settings. We extensively validate MedCoDi-M: first we benchmark it against five competitors on the MIMIC-CXR dataset, a state-of-the-art dataset for Chest X-ray and radiological report generation. Secondly, we perform a Visual Turing Test with expert radiologists to assess the realism and clinical relevance of the generated data, ensuring alignment with real-world scenarios. Finally, we assess the utility of MedCoDi-M in addressing key challenges in the medical field, such as anonymization, data scarcity and imbalance learning. The results are promising, demonstrating the applicability of MedCoDi-M in medical contexts. Project page is at https://cosbidev.github.io/MedCoDi-M/.
A Systematic Review of Intermediate Fusion in Multimodal Deep Learning for Biomedical Applications
Aug 02, 2024Abstract:Deep learning has revolutionized biomedical research by providing sophisticated methods to handle complex, high-dimensional data. Multimodal deep learning (MDL) further enhances this capability by integrating diverse data types such as imaging, textual data, and genetic information, leading to more robust and accurate predictive models. In MDL, differently from early and late fusion methods, intermediate fusion stands out for its ability to effectively combine modality-specific features during the learning process. This systematic review aims to comprehensively analyze and formalize current intermediate fusion methods in biomedical applications. We investigate the techniques employed, the challenges faced, and potential future directions for advancing intermediate fusion methods. Additionally, we introduce a structured notation to enhance the understanding and application of these methods beyond the biomedical domain. Our findings are intended to support researchers, healthcare professionals, and the broader deep learning community in developing more sophisticated and insightful multimodal models. Through this review, we aim to provide a foundational framework for future research and practical applications in the dynamic field of MDL.
Multi-Scale Texture Loss for CT denoising with GANs
Mar 25, 2024

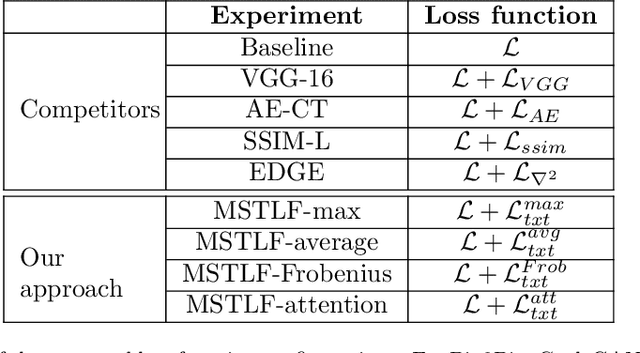

Abstract:Generative Adversarial Networks (GANs) have proved as a powerful framework for denoising applications in medical imaging. However, GAN-based denoising algorithms still suffer from limitations in capturing complex relationships within the images. In this regard, the loss function plays a crucial role in guiding the image generation process, encompassing how much a synthetic image differs from a real image. To grasp highly complex and non-linear textural relationships in the training process, this work presents a loss function that leverages the intrinsic multi-scale nature of the Gray-Level-Co-occurrence Matrix (GLCM). Although the recent advances in deep learning have demonstrated superior performance in classification and detection tasks, we hypothesize that its information content can be valuable when integrated into GANs' training. To this end, we propose a differentiable implementation of the GLCM suited for gradient-based optimization. Our approach also introduces a self-attention layer that dynamically aggregates the multi-scale texture information extracted from the images. We validate our approach by carrying out extensive experiments in the context of low-dose CT denoising, a challenging application that aims to enhance the quality of noisy CT scans. We utilize three publicly available datasets, including one simulated and two real datasets. The results are promising as compared to other well-established loss functions, being also consistent across three different GAN architectures. The code is available at: https://github.com/FrancescoDiFeola/DenoTextureLoss
A comparative study between paired and unpaired Image Quality Assessment in Low-Dose CT Denoising
Apr 11, 2023Abstract:The current deep learning approaches for low-dose CT denoising can be divided into paired and unpaired methods. The former involves the use of well-paired datasets, whilst the latter relaxes this constraint. The large availability of unpaired datasets has raised the interest in deepening unpaired denoising strategies that, in turn, need for robust evaluation techniques going beyond the qualitative evaluation. To this end, we can use quantitative image quality assessment scores that we divided into two categories, i.e., paired and unpaired measures. However, the interpretation of unpaired metrics is not straightforward, also because the consistency with paired metrics has not been fully investigated. To cope with this limitation, in this work we consider 15 paired and unpaired scores, which we applied to assess the performance of low-dose CT denoising. We perform an in-depth statistical analysis that not only studies the correlation between paired and unpaired metrics but also within each category. This brings out useful guidelines that can help researchers and practitioners select the right measure for their applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge