Roarke Horstmeyer
Detecting immune cells with label-free two-photon autofluorescence and deep learning
Jun 17, 2025Abstract:Label-free imaging has gained broad interest because of its potential to omit elaborate staining procedures which is especially relevant for in vivo use. Label-free multiphoton microscopy (MPM), for instance, exploits two-photon excitation of natural autofluorescence (AF) from native, metabolic proteins, making it ideal for in vivo endomicroscopy. Deep learning (DL) models have been widely used in other optical imaging technologies to predict specific target annotations and thereby digitally augment the specificity of these label-free images. However, this computational specificity has only rarely been implemented for MPM. In this work, we used a data set of label-free MPM images from a series of different immune cell types (5,075 individual cells for binary classification in mixed samples and 3,424 cells for a multi-class classification task) and trained a convolutional neural network (CNN) to classify cell types based on this label-free AF as input. A low-complexity squeezeNet architecture was able to achieve reliable immune cell classification results (0.89 ROC-AUC, 0.95 PR-AUC, for binary classification in mixed samples; 0.689 F1 score, 0.697 precision, 0.748 recall, and 0.683 MCC for six-class classification in isolated samples). Perturbation tests confirmed that the model is not confused by extracellular environment and that both input AF channels (NADH and FAD) are about equally important to the classification. In the future, such predictive DL models could directly detect specific immune cells in unstained images and thus, computationally improve the specificity of label-free MPM which would have great potential for in vivo endomicroscopy.
Computational 3D topographic microscopy from terabytes of data per sample
Jun 05, 2023



Abstract:We present a large-scale computational 3D topographic microscope that enables 6-gigapixel profilometric 3D imaging at micron-scale resolution across $>$110 cm$^2$ areas over multi-millimeter axial ranges. Our computational microscope, termed STARCAM (Scanning Topographic All-in-focus Reconstruction with a Computational Array Microscope), features a parallelized, 54-camera architecture with 3-axis translation to capture, for each sample of interest, a multi-dimensional, 2.1-terabyte (TB) dataset, consisting of a total of 224,640 9.4-megapixel images. We developed a self-supervised neural network-based algorithm for 3D reconstruction and stitching that jointly estimates an all-in-focus photometric composite and 3D height map across the entire field of view, using multi-view stereo information and image sharpness as a focal metric. The memory-efficient, compressed differentiable representation offered by the neural network effectively enables joint participation of the entire multi-TB dataset during the reconstruction process. To demonstrate the broad utility of our new computational microscope, we applied STARCAM to a variety of decimeter-scale objects, with applications ranging from cultural heritage to industrial inspection.
Tensorial tomographic Fourier Ptychography with applications to muscle tissue imaging
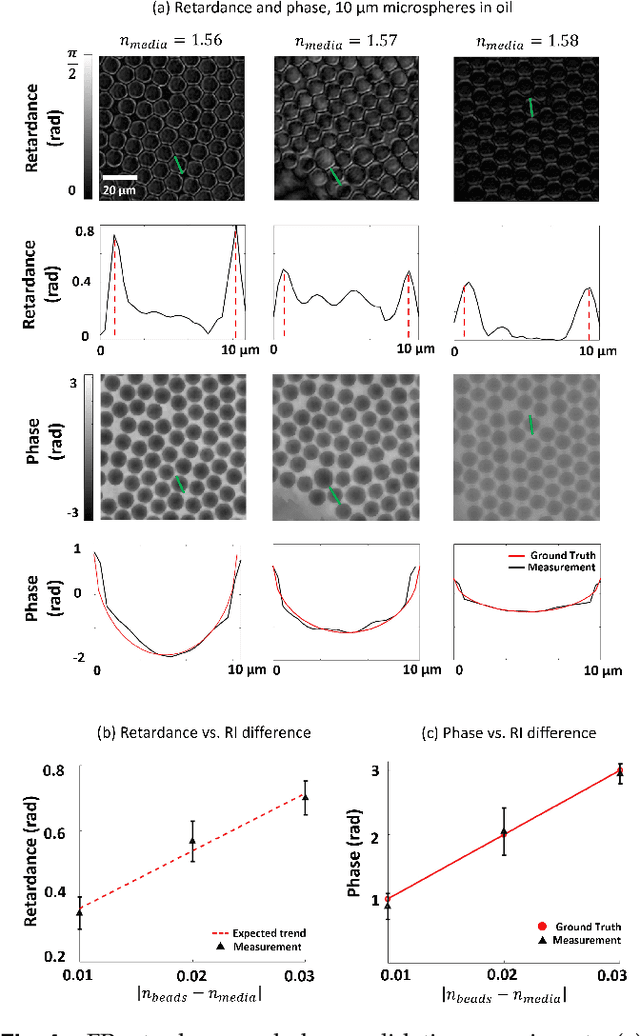
May 14, 2023Abstract:We report Tensorial tomographic Fourier Ptychography (ToFu), a new non-scanning label-free tomographic microscopy method for simultaneous imaging of quantitative phase and anisotropic specimen information in 3D. Built upon Fourier Ptychography, a quantitative phase imaging technique, ToFu additionally highlights the vectorial nature of light. The imaging setup consists of a standard microscope equipped with an LED matrix, a polarization generator, and a polarization-sensitive camera. Permittivity tensors of anisotropic samples are computationally recovered from polarized intensity measurements across three dimensions. We demonstrate ToFu's efficiency through volumetric reconstructions of refractive index, birefringence, and orientation for various validation samples, as well as tissue samples from muscle fibers and diseased heart tissue. Our reconstructions of muscle fibers resolve their 3D fine-filament structure and yield consistent morphological measurements compared to gold-standard second harmonic generation scanning confocal microscope images found in the literature. Additionally, we demonstrate reconstructions of a heart tissue sample that carries important polarization information for detecting cardiac amyloidosis.
Digital staining in optical microscopy using deep learning -- a review
Mar 14, 2023Abstract:Until recently, conventional biochemical staining had the undisputed status as well-established benchmark for most biomedical problems related to clinical diagnostics, fundamental research and biotechnology. Despite this role as gold-standard, staining protocols face several challenges, such as a need for extensive, manual processing of samples, substantial time delays, altered tissue homeostasis, limited choice of contrast agents for a given sample, 2D imaging instead of 3D tomography and many more. Label-free optical technologies, on the other hand, do not rely on exogenous and artificial markers, by exploiting intrinsic optical contrast mechanisms, where the specificity is typically less obvious to the human observer. Over the past few years, digital staining has emerged as a promising concept to use modern deep learning for the translation from optical contrast to established biochemical contrast of actual stainings. In this review article, we provide an in-depth analysis of the current state-of-the-art in this field, suggest methods of good practice, identify pitfalls and challenges and postulate promising advances towards potential future implementations and applications.
Parallelized computational 3D video microscopy of freely moving organisms at multiple gigapixels per second
Jan 19, 2023Abstract:To study the behavior of freely moving model organisms such as zebrafish (Danio rerio) and fruit flies (Drosophila) across multiple spatial scales, it would be ideal to use a light microscope that can resolve 3D information over a wide field of view (FOV) at high speed and high spatial resolution. However, it is challenging to design an optical instrument to achieve all of these properties simultaneously. Existing techniques for large-FOV microscopic imaging and for 3D image measurement typically require many sequential image snapshots, thus compromising speed and throughput. Here, we present 3D-RAPID, a computational microscope based on a synchronized array of 54 cameras that can capture high-speed 3D topographic videos over a 135-cm^2 area, achieving up to 230 frames per second at throughputs exceeding 5 gigapixels (GPs) per second. 3D-RAPID features a 3D reconstruction algorithm that, for each synchronized temporal snapshot, simultaneously fuses all 54 images seamlessly into a globally-consistent composite that includes a coregistered 3D height map. The self-supervised 3D reconstruction algorithm itself trains a spatiotemporally-compressed convolutional neural network (CNN) that maps raw photometric images to 3D topography, using stereo overlap redundancy and ray-propagation physics as the only supervision mechanism. As a result, our end-to-end 3D reconstruction algorithm is robust to generalization errors and scales to arbitrarily long videos from arbitrarily sized camera arrays. The scalable hardware and software design of 3D-RAPID addresses a longstanding problem in the field of behavioral imaging, enabling parallelized 3D observation of large collections of freely moving organisms at high spatiotemporal throughputs, which we demonstrate in ants (Pogonomyrmex barbatus), fruit flies, and zebrafish larvae.
Large-scale single-photon imaging
Dec 28, 2022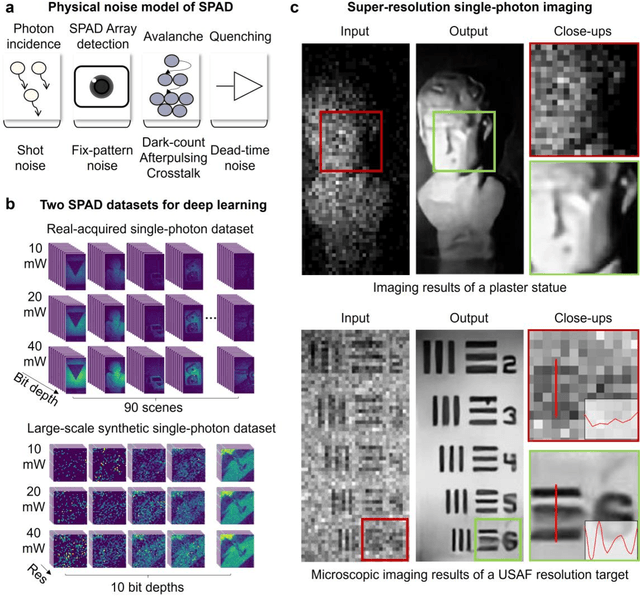
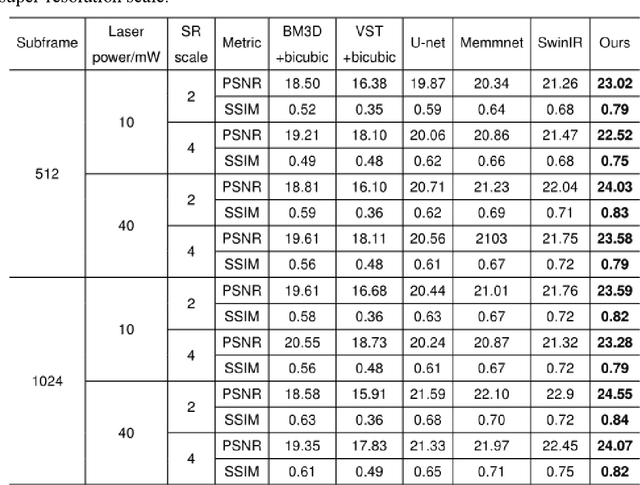

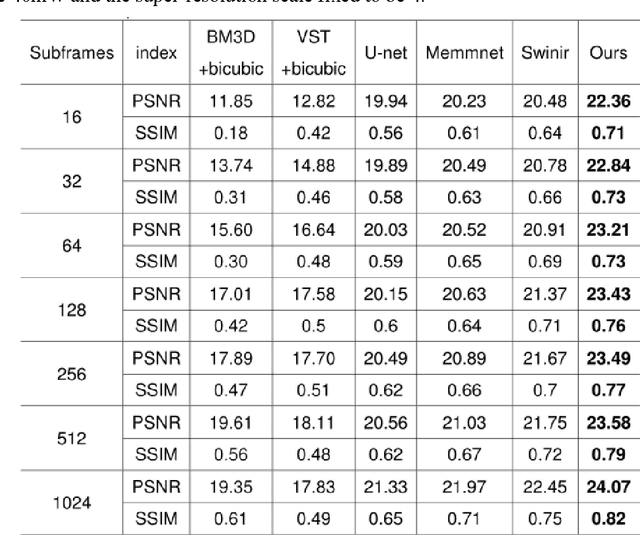
Abstract:Benefiting from its single-photon sensitivity, single-photon avalanche diode (SPAD) array has been widely applied in various fields such as fluorescence lifetime imaging and quantum computing. However, large-scale high-fidelity single-photon imaging remains a big challenge, due to the complex hardware manufacture craft and heavy noise disturbance of SPAD arrays. In this work, we introduce deep learning into SPAD, enabling super-resolution single-photon imaging over an order of magnitude, with significant enhancement of bit depth and imaging quality. We first studied the complex photon flow model of SPAD electronics to accurately characterize multiple physical noise sources, and collected a real SPAD image dataset (64 $\times$ 32 pixels, 90 scenes, 10 different bit depth, 3 different illumination flux, 2790 images in total) to calibrate noise model parameters. With this real-world physical noise model, we for the first time synthesized a large-scale realistic single-photon image dataset (image pairs of 5 different resolutions with maximum megapixels, 17250 scenes, 10 different bit depth, 3 different illumination flux, 2.6 million images in total) for subsequent network training. To tackle the severe super-resolution challenge of SPAD inputs with low bit depth, low resolution, and heavy noise, we further built a deep transformer network with a content-adaptive self-attention mechanism and gated fusion modules, which can dig global contextual features to remove multi-source noise and extract full-frequency details. We applied the technique on a series of experiments including macroscopic and microscopic imaging, microfluidic inspection, and Fourier ptychography. The experiments validate the technique's state-of-the-art super-resolution SPAD imaging performance, with more than 5 dB superiority on PSNR compared to the existing methods.
Multi-scale gigapixel microscopy using a multi-camera array microscope
Nov 30, 2022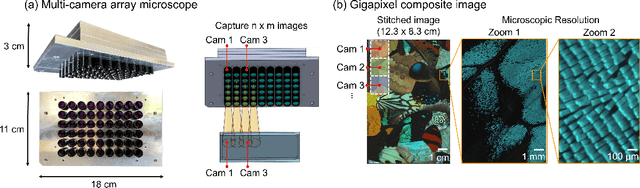
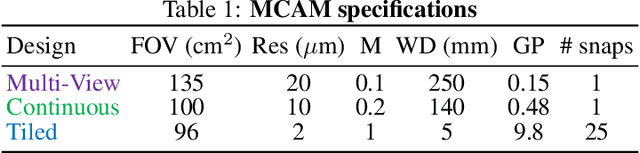
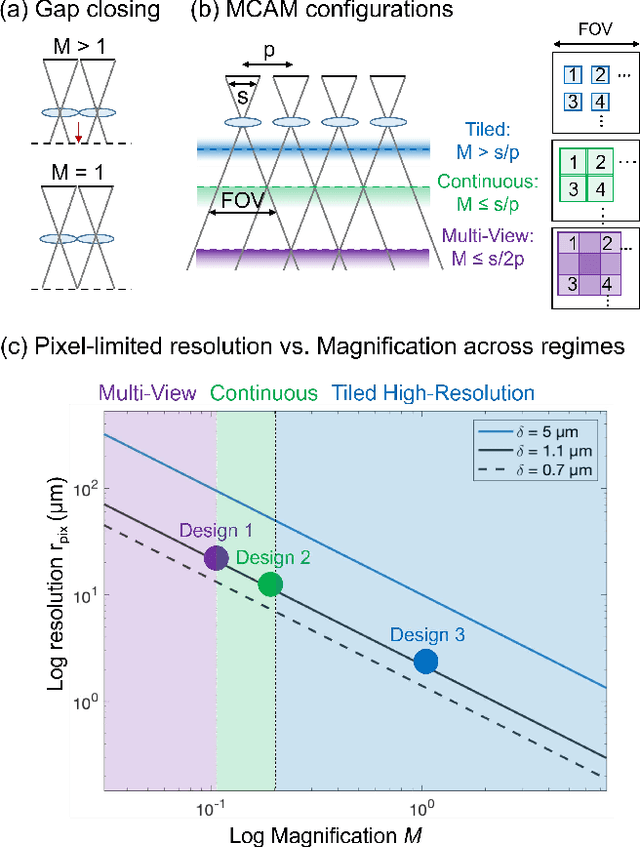
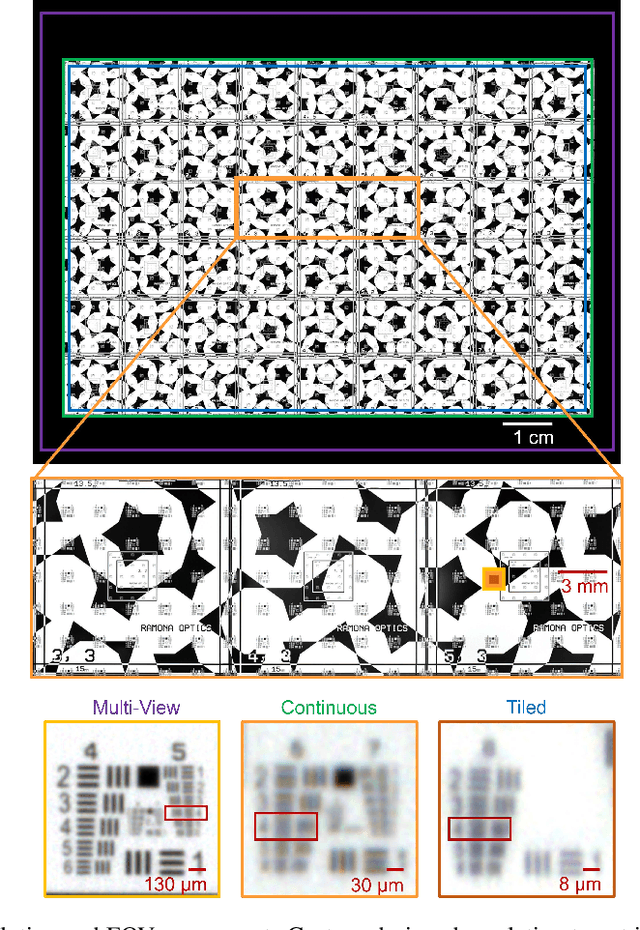
Abstract:This article experimentally examines different configurations of a novel multi-camera array microscope (MCAM) imaging technology. The MCAM is based upon a densely packed array of "micro-cameras" to jointly image across a large field-of-view at high resolution. Each micro-camera within the array images a unique area of a sample of interest, and then all acquired data with 54 micro-cameras are digitally combined into composite frames, whose total pixel counts significantly exceed the pixel counts of standard microscope systems. We present results from three unique MCAM configurations for different use cases. First, we demonstrate a configuration that simultaneously images and estimates the 3D object depth across a 100 x 135 mm^2 field-of-view (FOV) at approximately 20 um resolution, which results in 0.15 gigapixels (GP) per snapshot. Second, we demonstrate an MCAM configuration that records video across a continuous 83 x 123 mm^2 FOV with two-fold increased resolution (0.48 GP per frame). Finally, we report a third high-resolution configuration (2 um resolution) that can rapidly produce 9.8 GP composites of large histopathology specimens.
Tensorial tomographic differential phase-contrast microscopy
Apr 25, 2022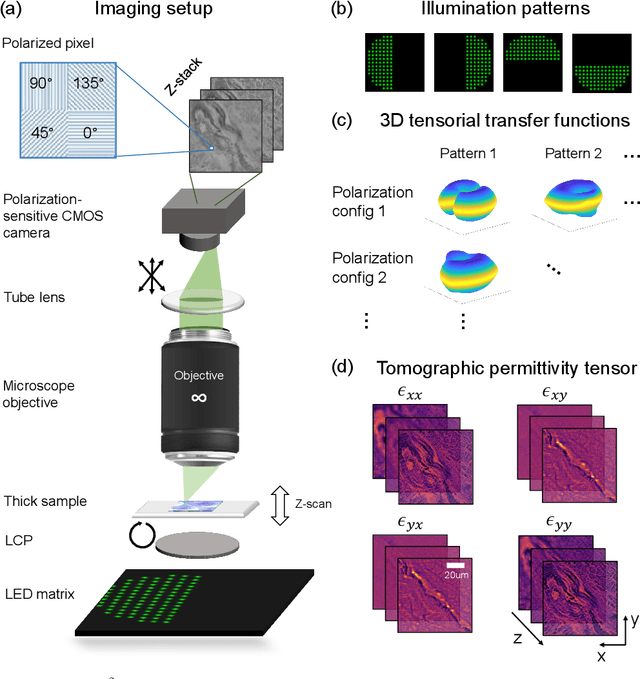
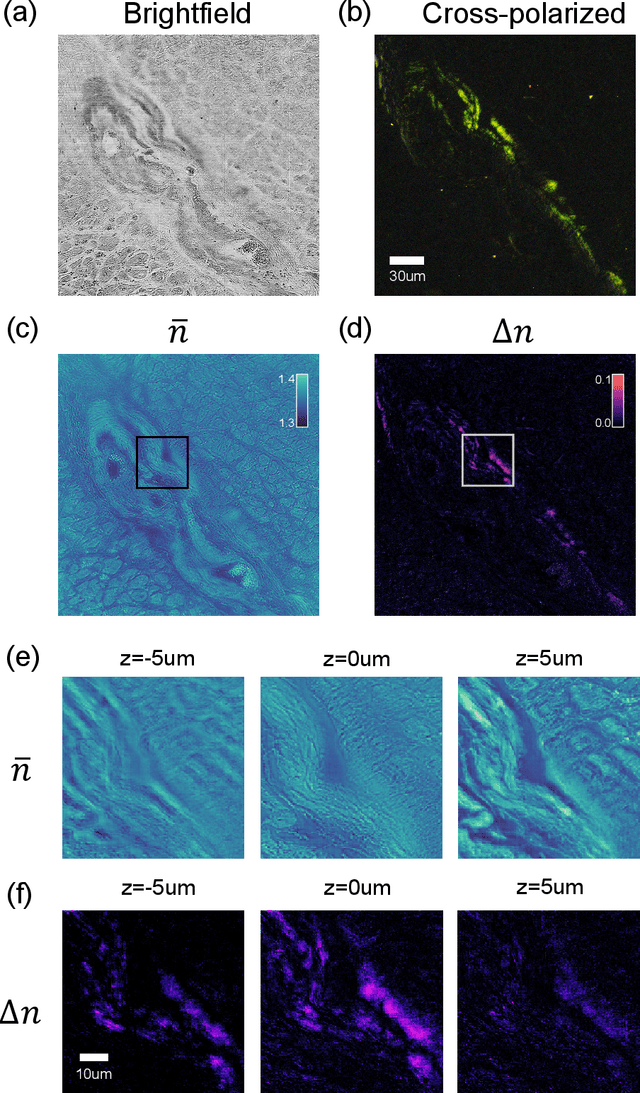
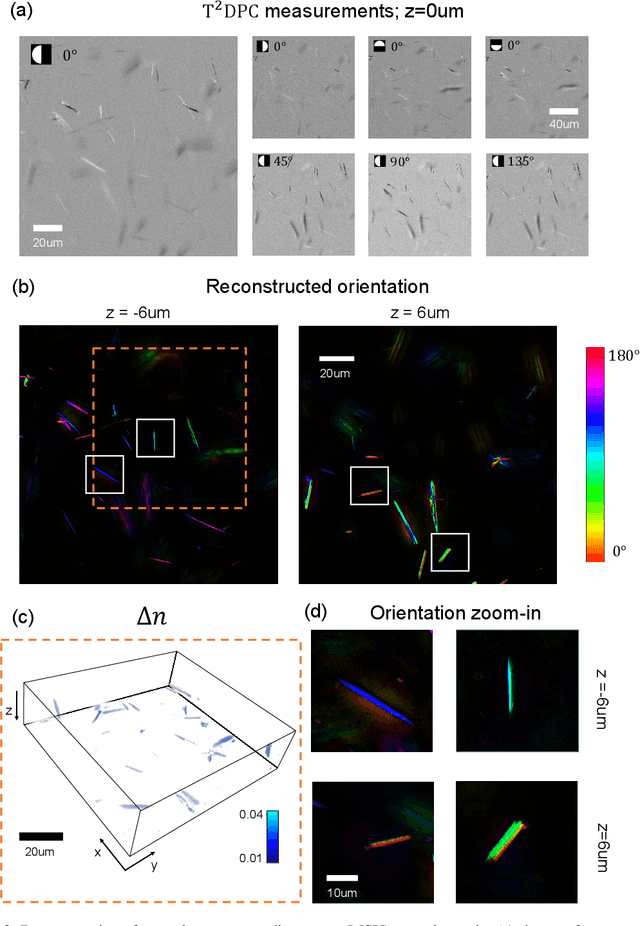
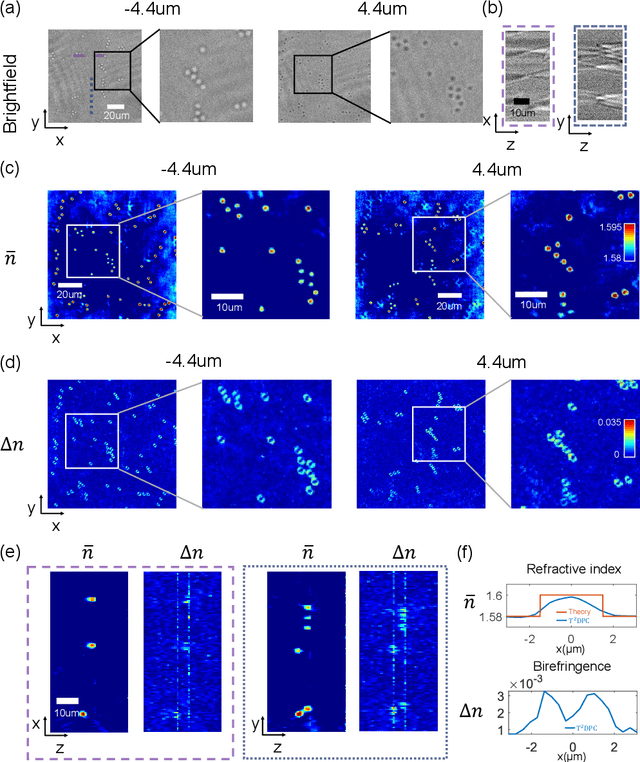

Abstract:We report Tensorial Tomographic Differential Phase-Contrast microscopy (T2DPC), a quantitative label-free tomographic imaging method for simultaneous measurement of phase and anisotropy. T2DPC extends differential phase-contrast microscopy, a quantitative phase imaging technique, to highlight the vectorial nature of light. The method solves for permittivity tensor of anisotropic samples from intensity measurements acquired with a standard microscope equipped with an LED matrix, a circular polarizer, and a polarization-sensitive camera. We demonstrate accurate volumetric reconstructions of refractive index, birefringence, and orientation for various validation samples, and show that the reconstructed polarization structures of a biological specimen are predictive of pathology.
Transient motion classification through turbid volumes via parallelized single-photon detection and deep contrastive embedding
Apr 04, 2022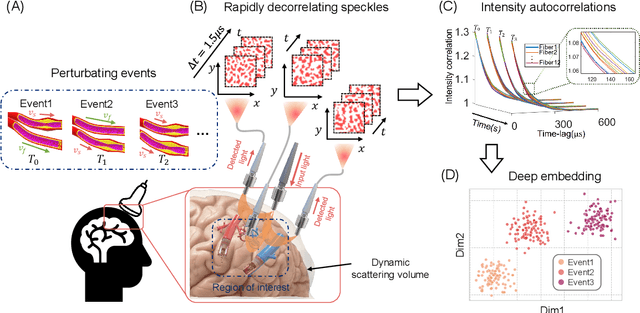
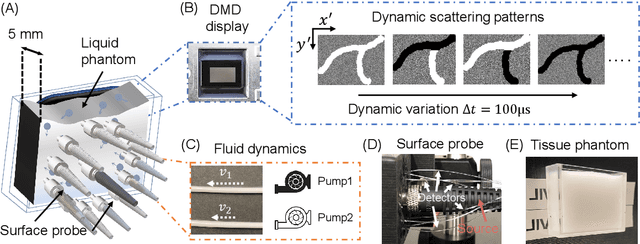
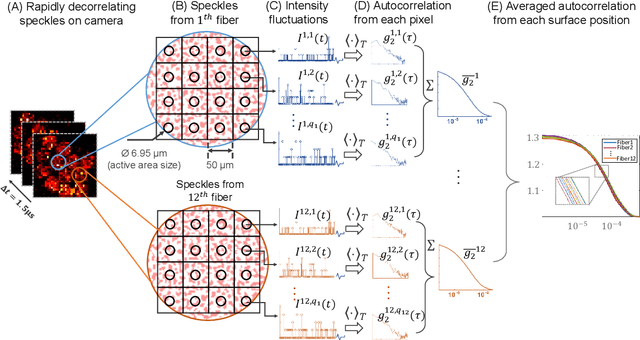
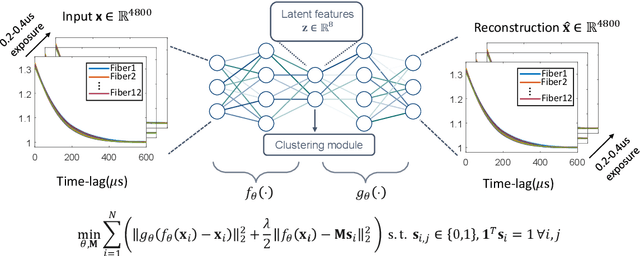
Abstract:Fast noninvasive probing of spatially varying decorrelating events, such as cerebral blood flow beneath the human skull, is an essential task in various scientific and clinical settings. One of the primary optical techniques used is diffuse correlation spectroscopy (DCS), whose classical implementation uses a single or few single-photon detectors, resulting in poor spatial localization accuracy and relatively low temporal resolution. Here, we propose a technique termed Classifying Rapid decorrelation Events via Parallelized single photon dEtection (CREPE)}, a new form of DCS that can probe and classify different decorrelating movements hidden underneath turbid volume with high sensitivity using parallelized speckle detection from a $32\times32$ pixel SPAD array. We evaluate our setup by classifying different spatiotemporal-decorrelating patterns hidden beneath a 5mm tissue-like phantom made with rapidly decorrelating dynamic scattering media. Twelve multi-mode fibers are used to collect scattered light from different positions on the surface of the tissue phantom. To validate our setup, we generate perturbed decorrelation patterns by both a digital micromirror device (DMD) modulated at multi-kilo-hertz rates, as well as a vessel phantom containing flowing fluid. Along with a deep contrastive learning algorithm that outperforms classic unsupervised learning methods, we demonstrate our approach can accurately detect and classify different transient decorrelation events (happening in 0.1-0.4s) underneath turbid scattering media, without any data labeling. This has the potential to be applied to noninvasively monitor deep tissue motion patterns, for example identifying normal or abnormal cerebral blood flow events, at multi-Hertz rates within a compact and static detection probe.
Quantitative Jones matrix imaging using vectorial Fourier ptychography
Oct 15, 2021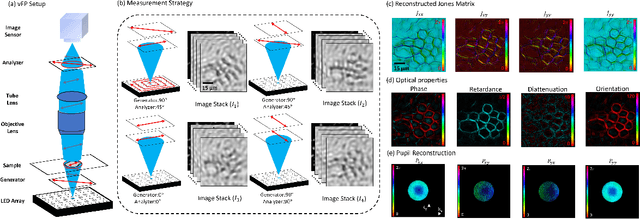
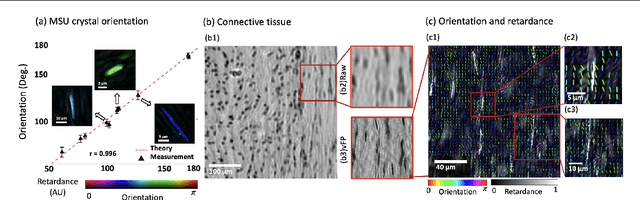
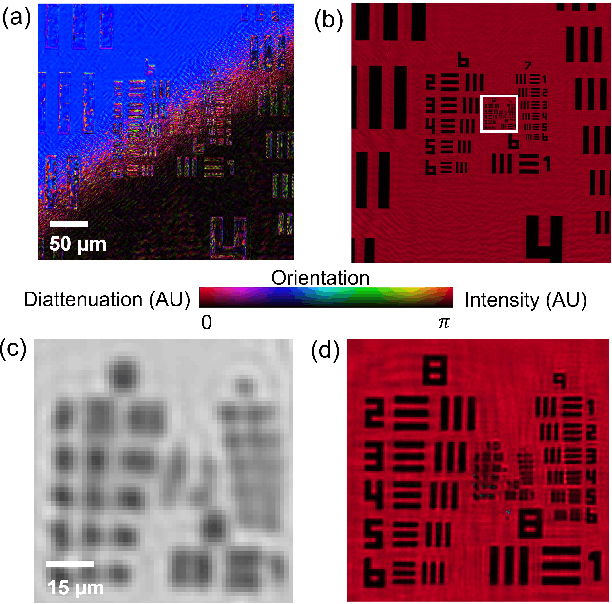

Abstract:This paper presents a microscopic imaging technique that uses variable-angle illumination to recover the complex polarimetric properties of a specimen at high resolution and over a large field-of-view. The approach extends Fourier ptychography, which is a synthetic aperture-based imaging approach to improve resolution with phaseless measurements, to additionally account for the vectorial nature of light. After images are acquired using a standard microscope outfitted with an LED illumination array and two polarizers, our vectorial Fourier Ptychography (vFP) algorithm solves for the complex 2x2 Jones matrix of the anisotropic specimen of interest at each resolved spatial location. We introduce a new sequential Gauss-Newton-based solver that additionally jointly estimates and removes polarization-dependent imaging system aberrations. We demonstrate effective vFP performance by generating large-area (29 mm$^2$), high-resolution (1.24 $\mu$m full-pitch) reconstructions of sample absorption, phase, orientation, diattenuation, and retardance for a variety of calibration samples and biological specimens.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge