Amey Chaware
Detecting immune cells with label-free two-photon autofluorescence and deep learning
Jun 17, 2025Abstract:Label-free imaging has gained broad interest because of its potential to omit elaborate staining procedures which is especially relevant for in vivo use. Label-free multiphoton microscopy (MPM), for instance, exploits two-photon excitation of natural autofluorescence (AF) from native, metabolic proteins, making it ideal for in vivo endomicroscopy. Deep learning (DL) models have been widely used in other optical imaging technologies to predict specific target annotations and thereby digitally augment the specificity of these label-free images. However, this computational specificity has only rarely been implemented for MPM. In this work, we used a data set of label-free MPM images from a series of different immune cell types (5,075 individual cells for binary classification in mixed samples and 3,424 cells for a multi-class classification task) and trained a convolutional neural network (CNN) to classify cell types based on this label-free AF as input. A low-complexity squeezeNet architecture was able to achieve reliable immune cell classification results (0.89 ROC-AUC, 0.95 PR-AUC, for binary classification in mixed samples; 0.689 F1 score, 0.697 precision, 0.748 recall, and 0.683 MCC for six-class classification in isolated samples). Perturbation tests confirmed that the model is not confused by extracellular environment and that both input AF channels (NADH and FAD) are about equally important to the classification. In the future, such predictive DL models could directly detect specific immune cells in unstained images and thus, computationally improve the specificity of label-free MPM which would have great potential for in vivo endomicroscopy.
Tensorial tomographic Fourier Ptychography with applications to muscle tissue imaging
May 14, 2023Abstract:We report Tensorial tomographic Fourier Ptychography (ToFu), a new non-scanning label-free tomographic microscopy method for simultaneous imaging of quantitative phase and anisotropic specimen information in 3D. Built upon Fourier Ptychography, a quantitative phase imaging technique, ToFu additionally highlights the vectorial nature of light. The imaging setup consists of a standard microscope equipped with an LED matrix, a polarization generator, and a polarization-sensitive camera. Permittivity tensors of anisotropic samples are computationally recovered from polarized intensity measurements across three dimensions. We demonstrate ToFu's efficiency through volumetric reconstructions of refractive index, birefringence, and orientation for various validation samples, as well as tissue samples from muscle fibers and diseased heart tissue. Our reconstructions of muscle fibers resolve their 3D fine-filament structure and yield consistent morphological measurements compared to gold-standard second harmonic generation scanning confocal microscope images found in the literature. Additionally, we demonstrate reconstructions of a heart tissue sample that carries important polarization information for detecting cardiac amyloidosis.
Digital staining in optical microscopy using deep learning -- a review
Mar 14, 2023Abstract:Until recently, conventional biochemical staining had the undisputed status as well-established benchmark for most biomedical problems related to clinical diagnostics, fundamental research and biotechnology. Despite this role as gold-standard, staining protocols face several challenges, such as a need for extensive, manual processing of samples, substantial time delays, altered tissue homeostasis, limited choice of contrast agents for a given sample, 2D imaging instead of 3D tomography and many more. Label-free optical technologies, on the other hand, do not rely on exogenous and artificial markers, by exploiting intrinsic optical contrast mechanisms, where the specificity is typically less obvious to the human observer. Over the past few years, digital staining has emerged as a promising concept to use modern deep learning for the translation from optical contrast to established biochemical contrast of actual stainings. In this review article, we provide an in-depth analysis of the current state-of-the-art in this field, suggest methods of good practice, identify pitfalls and challenges and postulate promising advances towards potential future implementations and applications.
Multi-scale gigapixel microscopy using a multi-camera array microscope
Nov 30, 2022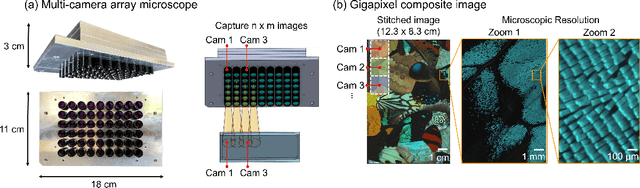
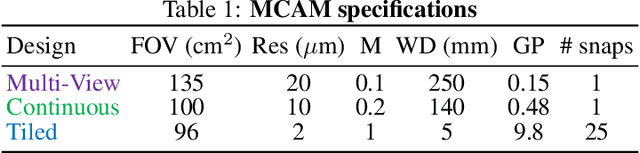
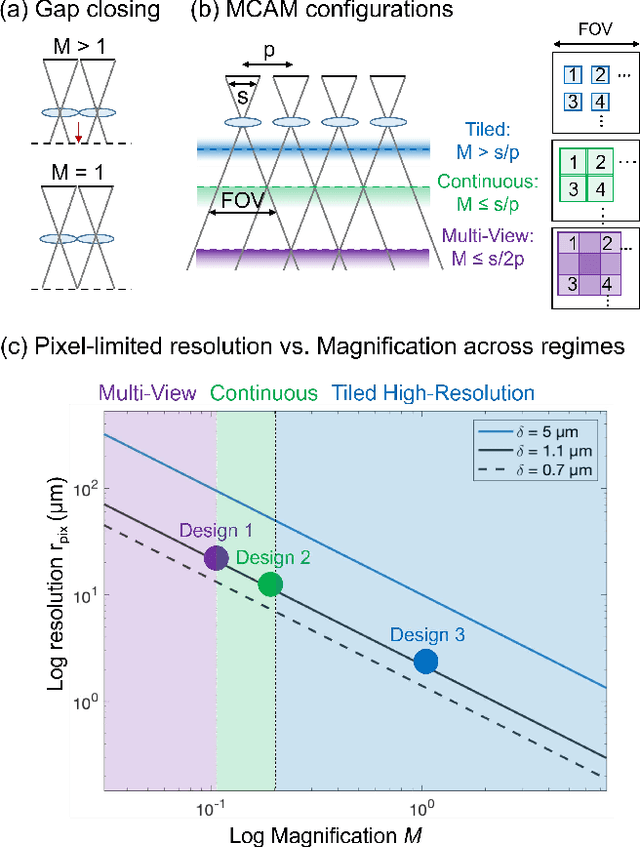
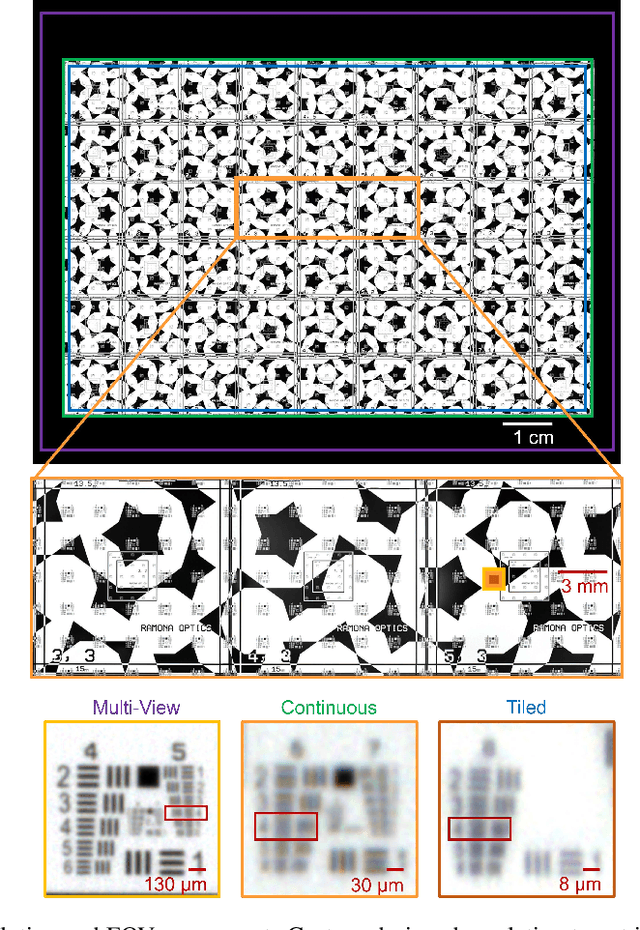
Abstract:This article experimentally examines different configurations of a novel multi-camera array microscope (MCAM) imaging technology. The MCAM is based upon a densely packed array of "micro-cameras" to jointly image across a large field-of-view at high resolution. Each micro-camera within the array images a unique area of a sample of interest, and then all acquired data with 54 micro-cameras are digitally combined into composite frames, whose total pixel counts significantly exceed the pixel counts of standard microscope systems. We present results from three unique MCAM configurations for different use cases. First, we demonstrate a configuration that simultaneously images and estimates the 3D object depth across a 100 x 135 mm^2 field-of-view (FOV) at approximately 20 um resolution, which results in 0.15 gigapixels (GP) per snapshot. Second, we demonstrate an MCAM configuration that records video across a continuous 83 x 123 mm^2 FOV with two-fold increased resolution (0.48 GP per frame). Finally, we report a third high-resolution configuration (2 um resolution) that can rapidly produce 9.8 GP composites of large histopathology specimens.
Increasing a microscope's effective field of view via overlapped imaging and machine learning
Oct 10, 2021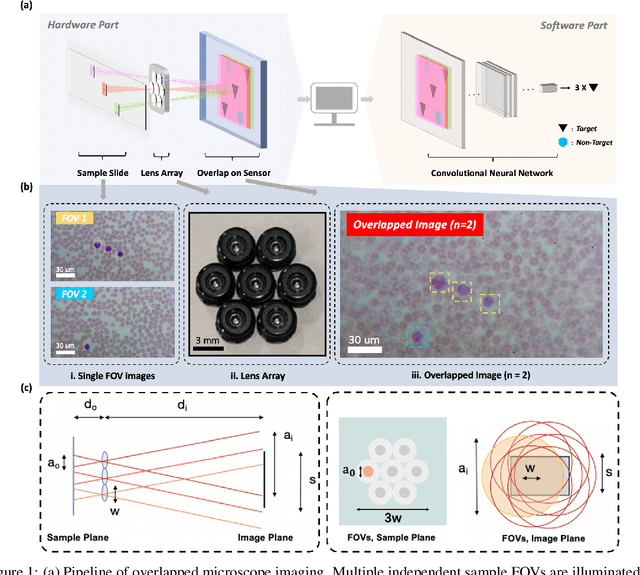
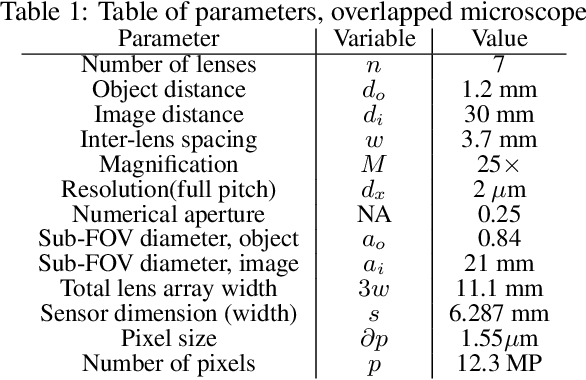
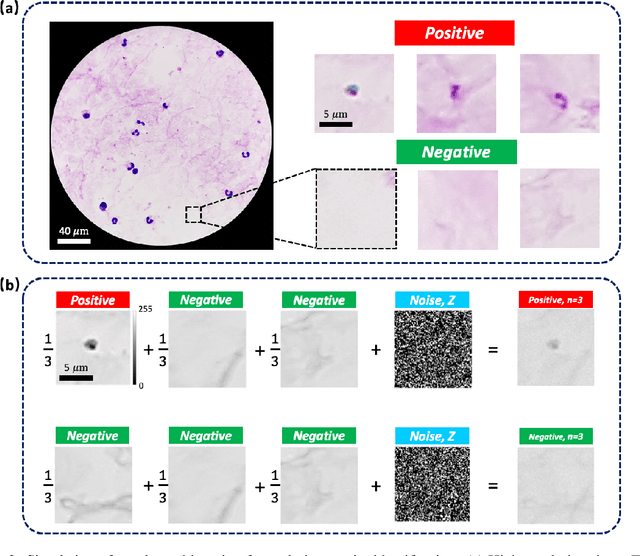
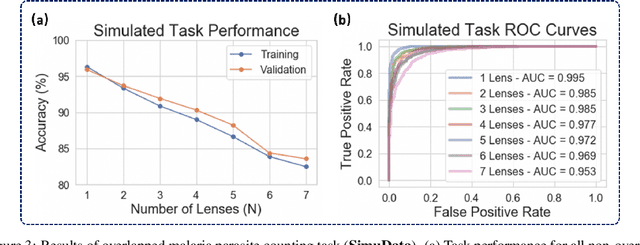
Abstract:This work demonstrates a multi-lens microscopic imaging system that overlaps multiple independent fields of view on a single sensor for high-efficiency automated specimen analysis. Automatic detection, classification and counting of various morphological features of interest is now a crucial component of both biomedical research and disease diagnosis. While convolutional neural networks (CNNs) have dramatically improved the accuracy of counting cells and sub-cellular features from acquired digital image data, the overall throughput is still typically hindered by the limited space-bandwidth product (SBP) of conventional microscopes. Here, we show both in simulation and experiment that overlapped imaging and co-designed analysis software can achieve accurate detection of diagnostically-relevant features for several applications, including counting of white blood cells and the malaria parasite, leading to multi-fold increase in detection and processing throughput with minimal reduction in accuracy.
Physics-enhanced machine learning for virtual fluorescence microscopy
Apr 21, 2020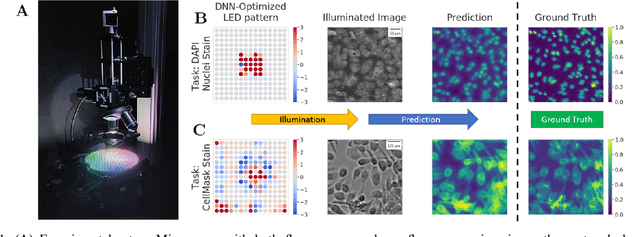
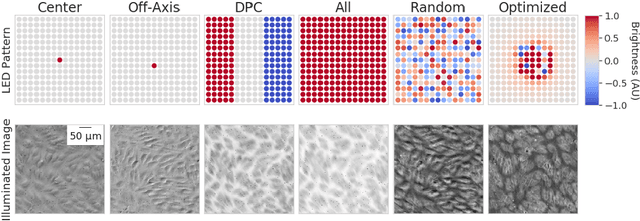
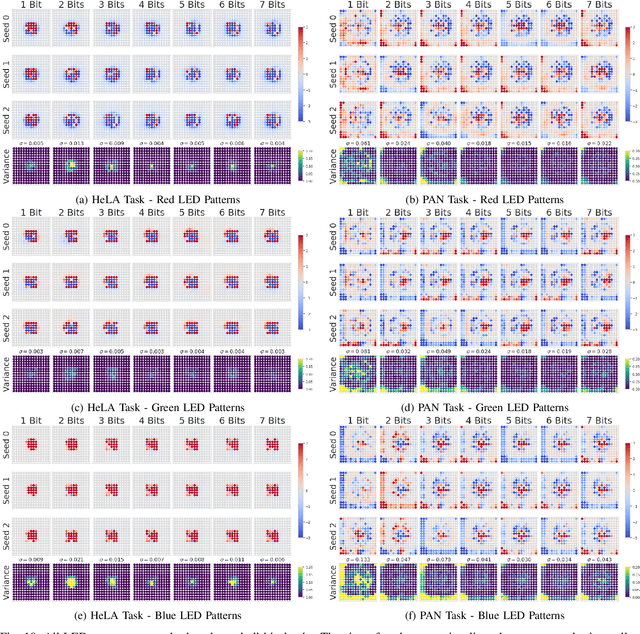
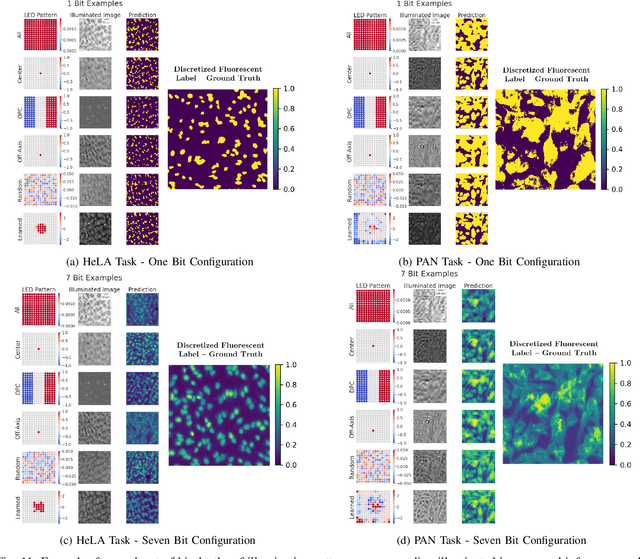

Abstract:This paper introduces a new method of data-driven microscope design for virtual fluorescence microscopy. Our results show that by including a model of illumination within the first layers of a deep convolutional neural network, it is possible to learn task-specific LED patterns that substantially improve the ability to infer fluorescence image information from unstained transmission microscopy images. We validated our method on two different experimental setups, with different magnifications and different sample types, to show a consistent improvement in performance as compared to conventional illumination methods. Additionally, to understand the importance of learned illumination on inference task, we varied the dynamic range of the fluorescent image targets (from one to seven bits), and showed that the margin of improvement for learned patterns increased with the information content of the target. This work demonstrates the power of programmable optical elements at enabling better machine learning algorithm performance and at providing physical insight into next generation of machine-controlled imaging systems.
Towards an Intelligent Microscope: adaptively learned illumination for optimal sample classification
Oct 22, 2019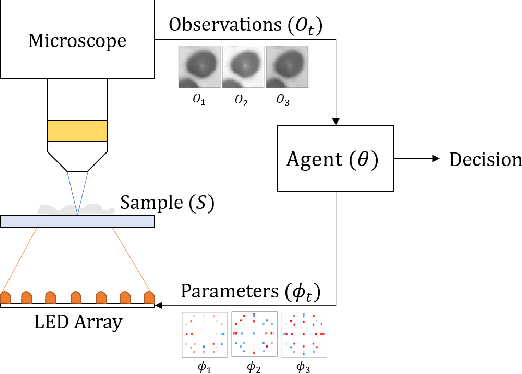
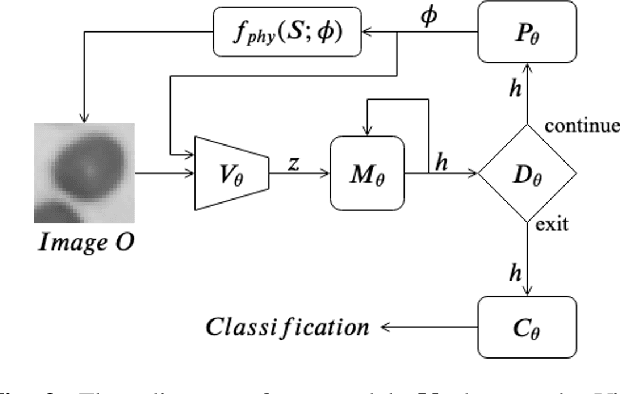
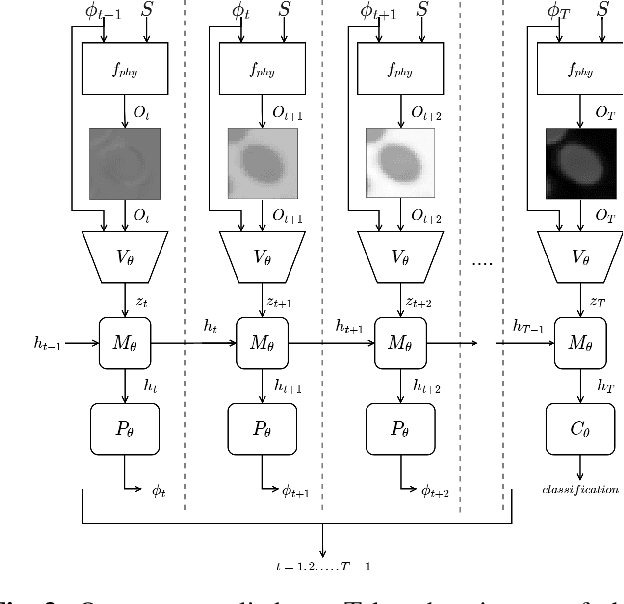
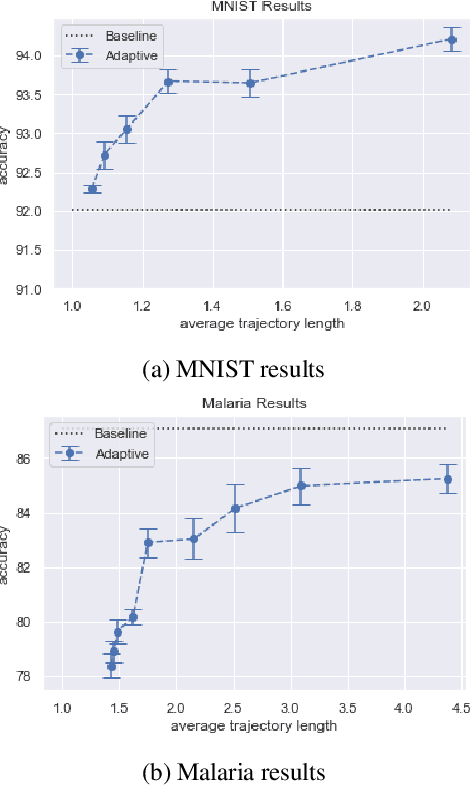
Abstract:Recent machine learning techniques have dramatically changed how we process digital images. However, the way in which we capture images is still largely driven by human intuition and experience. This restriction is in part due to the many available degrees of freedom that alter the image acquisition process (lens focus, exposure, filtering, etc). Here we focus on one such degree of freedom - illumination within a microscope - which can drastically alter information captured by the image sensor. We present a reinforcement learning system that adaptively explores optimal patterns to illuminate specimens for immediate classification. The agent uses a recurrent latent space to encode a large set of variably-illuminated samples and illumination patterns. We train our agent using a reward that balances classification confidence with image acquisition cost. By synthesizing knowledge over multiple snapshots, the agent can classify on the basis of all previous images with higher accuracy than from naively illuminated images, thus demonstrating a smarter way to physically capture task-specific information.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge