Richard L. J. Qiu
Systematic Review and Meta-analysis of AI-driven MRI Motion Artifact Detection and Correction
Sep 05, 2025Abstract:Background: To systematically review and perform a meta-analysis of artificial intelligence (AI)-driven methods for detecting and correcting magnetic resonance imaging (MRI) motion artifacts, assessing current developments, effectiveness, challenges, and future research directions. Methods: A comprehensive systematic review and meta-analysis were conducted, focusing on deep learning (DL) approaches, particularly generative models, for the detection and correction of MRI motion artifacts. Quantitative data were extracted regarding utilized datasets, DL architectures, and performance metrics. Results: DL, particularly generative models, show promise for reducing motion artifacts and improving image quality; however, limited generalizability, reliance on paired training data, and risk of visual distortions remain key challenges that motivate standardized datasets and reporting. Conclusions: AI-driven methods, particularly DL generative models, show significant potential for improving MRI image quality by effectively addressing motion artifacts. However, critical challenges must be addressed, including the need for comprehensive public datasets, standardized reporting protocols for artifact levels, and more advanced, adaptable DL techniques to reduce reliance on extensive paired datasets. Addressing these aspects could substantially enhance MRI diagnostic accuracy, reduce healthcare costs, and improve patient care outcomes.
MRI motion correction via efficient residual-guided denoising diffusion probabilistic models
May 06, 2025Abstract:Purpose: Motion artifacts in magnetic resonance imaging (MRI) significantly degrade image quality and impair quantitative analysis. Conventional mitigation strategies, such as repeated acquisitions or motion tracking, are costly and workflow-intensive. This study introduces Res-MoCoDiff, an efficient denoising diffusion probabilistic model tailored for MRI motion artifact correction. Methods: Res-MoCoDiff incorporates a novel residual error shifting mechanism in the forward diffusion process, aligning the noise distribution with motion-corrupted data and enabling an efficient four-step reverse diffusion. A U-net backbone enhanced with Swin-Transformer blocks conventional attention layers, improving adaptability across resolutions. Training employs a combined l1+l2 loss, which promotes image sharpness and reduces pixel-level errors. Res-MoCoDiff was evaluated on synthetic dataset generated using a realistic motion simulation framework and on an in-vivo dataset. Comparative analyses were conducted against established methods, including CycleGAN, Pix2pix, and MT-DDPM using quantitative metrics such as peak signal-to-noise ratio (PSNR), structural similarity index measure (SSIM), and normalized mean squared error (NMSE). Results: The proposed method demonstrated superior performance in removing motion artifacts across all motion severity levels. Res-MoCoDiff consistently achieved the highest SSIM and the lowest NMSE values, with a PSNR of up to 41.91+-2.94 dB for minor distortions. Notably, the average sampling time was reduced to 0.37 seconds per batch of two image slices, compared with 101.74 seconds for conventional approaches.
RoMedFormer: A Rotary-Embedding Transformer Foundation Model for 3D Genito-Pelvic Structure Segmentation in MRI and CT
Mar 18, 2025



Abstract:Deep learning-based segmentation of genito-pelvic structures in MRI and CT is crucial for applications such as radiation therapy, surgical planning, and disease diagnosis. However, existing segmentation models often struggle with generalizability across imaging modalities, and anatomical variations. In this work, we propose RoMedFormer, a rotary-embedding transformer-based foundation model designed for 3D female genito-pelvic structure segmentation in both MRI and CT. RoMedFormer leverages self-supervised learning and rotary positional embeddings to enhance spatial feature representation and capture long-range dependencies in 3D medical data. We pre-train our model using a diverse dataset of 3D MRI and CT scans and fine-tune it for downstream segmentation tasks. Experimental results demonstrate that RoMedFormer achieves superior performance segmenting genito-pelvic organs. Our findings highlight the potential of transformer-based architectures in medical image segmentation and pave the way for more transferable segmentation frameworks.
Advancing MRI Reconstruction: A Systematic Review of Deep Learning and Compressed Sensing Integration
Jan 24, 2025
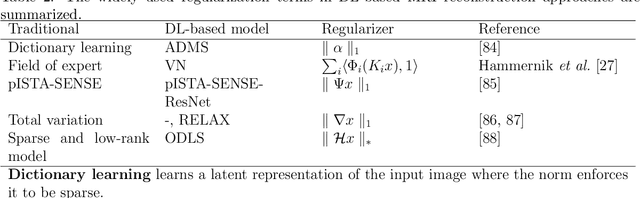
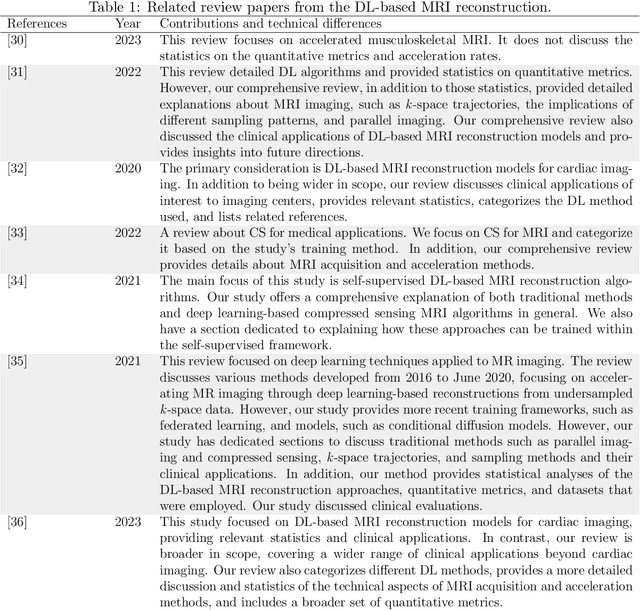
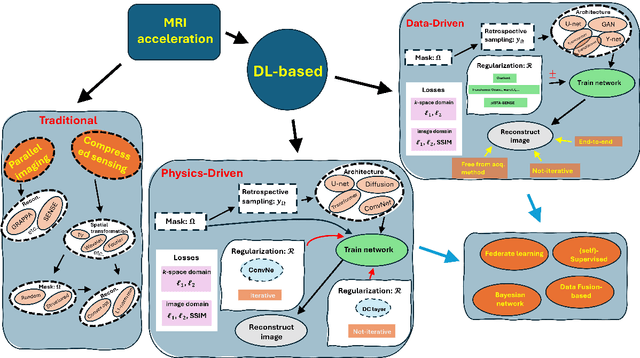

Abstract:Magnetic resonance imaging (MRI) is a non-invasive imaging modality and provides comprehensive anatomical and functional insights into the human body. However, its long acquisition times can lead to patient discomfort, motion artifacts, and limiting real-time applications. To address these challenges, strategies such as parallel imaging have been applied, which utilize multiple receiver coils to speed up the data acquisition process. Additionally, compressed sensing (CS) is a method that facilitates image reconstruction from sparse data, significantly reducing image acquisition time by minimizing the amount of data collection needed. Recently, deep learning (DL) has emerged as a powerful tool for improving MRI reconstruction. It has been integrated with parallel imaging and CS principles to achieve faster and more accurate MRI reconstructions. This review comprehensively examines DL-based techniques for MRI reconstruction. We categorize and discuss various DL-based methods, including end-to-end approaches, unrolled optimization, and federated learning, highlighting their potential benefits. Our systematic review highlights significant contributions and underscores the potential of DL in MRI reconstruction. Additionally, we summarize key results and trends in DL-based MRI reconstruction, including quantitative metrics, the dataset, acceleration factors, and the progress of and research interest in DL techniques over time. Finally, we discuss potential future directions and the importance of DL-based MRI reconstruction in advancing medical imaging. To facilitate further research in this area, we provide a GitHub repository that includes up-to-date DL-based MRI reconstruction publications and public datasets-https://github.com/mosaf/Awesome-DL-based-CS-MRI.
T1-contrast Enhanced MRI Generation from Multi-parametric MRI for Glioma Patients with Latent Tumor Conditioning
Sep 03, 2024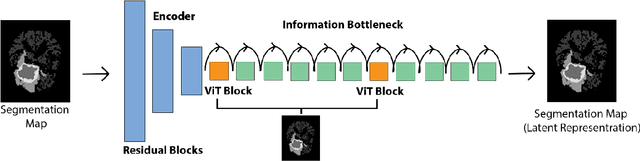
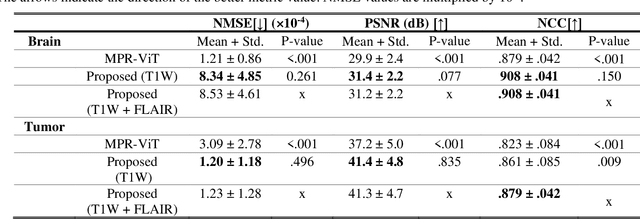
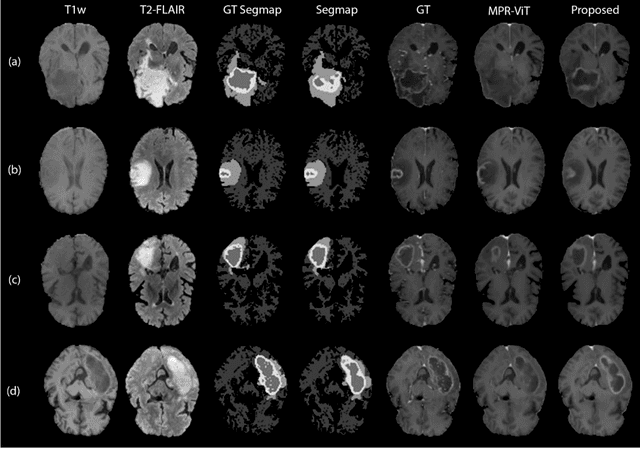
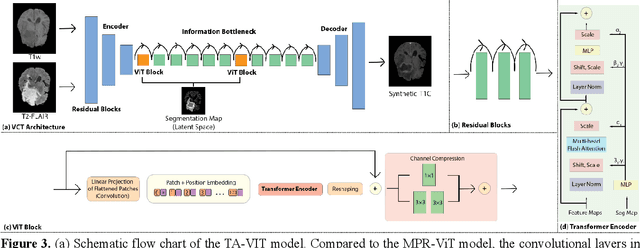
Abstract:Objective: Gadolinium-based contrast agents (GBCAs) are commonly used in MRI scans of patients with gliomas to enhance brain tumor characterization using T1-weighted (T1W) MRI. However, there is growing concern about GBCA toxicity. This study develops a deep-learning framework to generate T1-postcontrast (T1C) from pre-contrast multiparametric MRI. Approach: We propose the tumor-aware vision transformer (TA-ViT) model that predicts high-quality T1C images. The predicted tumor region is significantly improved (P < .001) by conditioning the transformer layers from predicted segmentation maps through adaptive layer norm zero mechanism. The predicted segmentation maps were generated with the multi-parametric residual (MPR) ViT model and transformed into a latent space to produce compressed, feature-rich representations. The TA-ViT model predicted T1C MRI images of 501 glioma cases. Selected patients were split into training (N=400), validation (N=50), and test (N=51) sets. Main Results: Both qualitative and quantitative results demonstrate that the TA-ViT model performs superior against the benchmark MRP-ViT model. Our method produces synthetic T1C MRI with high soft tissue contrast and more accurately reconstructs both the tumor and whole brain volumes. The synthesized T1C images achieved remarkable improvements in both tumor and healthy tissue regions compared to the MRP-ViT model. For healthy tissue and tumor regions, the results were as follows: NMSE: 8.53 +/- 4.61E-4; PSNR: 31.2 +/- 2.2; NCC: 0.908 +/- .041 and NMSE: 1.22 +/- 1.27E-4, PSNR: 41.3 +/- 4.7, and NCC: 0.879 +/- 0.042, respectively. Significance: The proposed method generates synthetic T1C images that closely resemble real T1C images. Future development and application of this approach may enable contrast-agent-free MRI for brain tumor patients, eliminating the risk of GBCA toxicity and simplifying the MRI scan protocol.
Deep Learning Based Apparent Diffusion Coefficient Map Generation1 from Multi-parametric MR Images for Patients with Diffuse Gliomas
Jul 02, 2024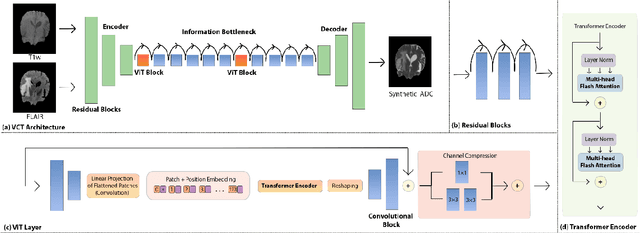
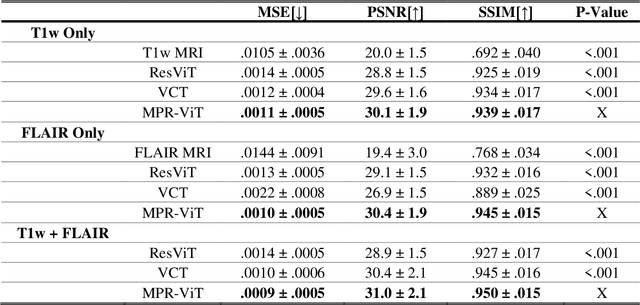
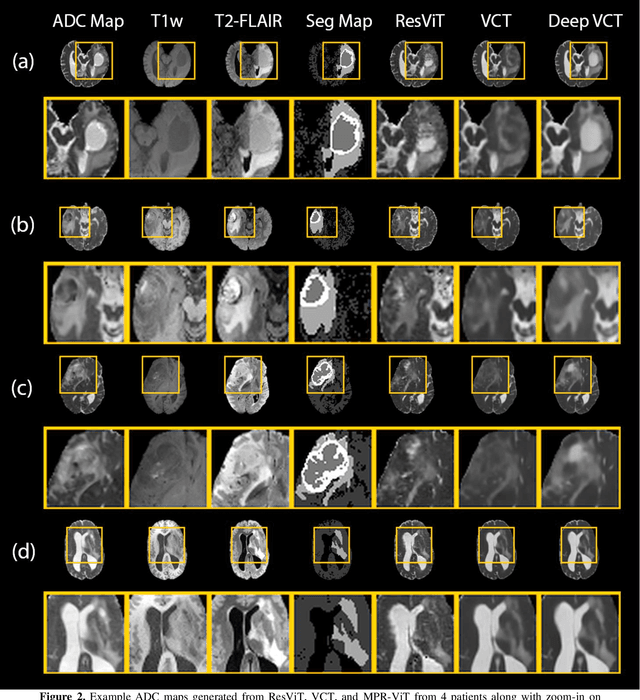
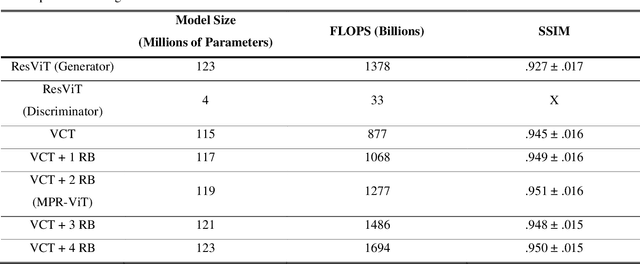
Abstract:Purpose: Apparent diffusion coefficient (ADC) maps derived from diffusion weighted (DWI) MRI provides functional measurements about the water molecules in tissues. However, DWI is time consuming and very susceptible to image artifacts, leading to inaccurate ADC measurements. This study aims to develop a deep learning framework to synthesize ADC maps from multi-parametric MR images. Methods: We proposed the multiparametric residual vision transformer model (MPR-ViT) that leverages the long-range context of ViT layers along with the precision of convolutional operators. Residual blocks throughout the network significantly increasing the representational power of the model. The MPR-ViT model was applied to T1w and T2- fluid attenuated inversion recovery images of 501 glioma cases from a publicly available dataset including preprocessed ADC maps. Selected patients were divided into training (N=400), validation (N=50) and test (N=51) sets, respectively. Using the preprocessed ADC maps as ground truth, model performance was evaluated and compared against the Vision Convolutional Transformer (VCT) and residual vision transformer (ResViT) models. Results: The results are as follows using T1w + T2-FLAIR MRI as inputs: MPR-ViT - PSNR: 31.0 +/- 2.1, MSE: 0.009 +/- 0.0005, SSIM: 0.950 +/- 0.015. In addition, ablation studies showed the relative impact on performance of each input sequence. Both qualitative and quantitative results indicate that the proposed MR- ViT model performs favorably against the ground truth data. Conclusion: We show that high-quality ADC maps can be synthesized from structural MRI using a MPR- VCT model. Our predicted images show better conformality to the ground truth volume than ResViT and VCT predictions. These high-quality synthetic ADC maps would be particularly useful for disease diagnosis and intervention, especially when ADC maps have artifacts or are unavailable.
Adaptive Self-Supervised Consistency-Guided Diffusion Model for Accelerated MRI Reconstruction
Jun 21, 2024Abstract:Purpose: To propose a self-supervised deep learning-based compressed sensing MRI (DL-based CS-MRI) method named "Adaptive Self-Supervised Consistency Guided Diffusion Model (ASSCGD)" to accelerate data acquisition without requiring fully sampled datasets. Materials and Methods: We used the fastMRI multi-coil brain axial T2-weighted (T2-w) dataset from 1,376 cases and single-coil brain quantitative magnetization prepared 2 rapid acquisition gradient echoes (MP2RAGE) T1 maps from 318 cases to train and test our model. Robustness against domain shift was evaluated using two out-of-distribution (OOD) datasets: multi-coil brain axial postcontrast T1 -weighted (T1c) dataset from 50 cases and axial T1-weighted (T1-w) dataset from 50 patients. Data were retrospectively subsampled at acceleration rates R in {2x, 4x, 8x}. ASSCGD partitions a random sampling pattern into two disjoint sets, ensuring data consistency during training. We compared our method with ReconFormer Transformer and SS-MRI, assessing performance using normalized mean squared error (NMSE), peak signal-to-noise ratio (PSNR), and structural similarity index (SSIM). Statistical tests included one-way analysis of variance (ANOVA) and multi-comparison Tukey's Honesty Significant Difference (HSD) tests. Results: ASSCGD preserved fine structures and brain abnormalities visually better than comparative methods at R = 8x for both multi-coil and single-coil datasets. It achieved the lowest NMSE at R in {4x, 8x}, and the highest PSNR and SSIM values at all acceleration rates for the multi-coil dataset. Similar trends were observed for the single-coil dataset, though SSIM values were comparable to ReconFormer at R in {2x, 8x}. These results were further confirmed by the voxel-wise correlation scatter plots. OOD results showed significant (p << 10^-5 ) improvements in undersampled image quality after reconstruction.
Mammo-CLIP: Leveraging Contrastive Language-Image Pre-training (CLIP) for Enhanced Breast Cancer Diagnosis with Multi-view Mammography
Apr 24, 2024Abstract:Although fusion of information from multiple views of mammograms plays an important role to increase accuracy of breast cancer detection, developing multi-view mammograms-based computer-aided diagnosis (CAD) schemes still faces challenges and no such CAD schemes have been used in clinical practice. To overcome the challenges, we investigate a new approach based on Contrastive Language-Image Pre-training (CLIP), which has sparked interest across various medical imaging tasks. By solving the challenges in (1) effectively adapting the single-view CLIP for multi-view feature fusion and (2) efficiently fine-tuning this parameter-dense model with limited samples and computational resources, we introduce Mammo-CLIP, the first multi-modal framework to process multi-view mammograms and corresponding simple texts. Mammo-CLIP uses an early feature fusion strategy to learn multi-view relationships in four mammograms acquired from the CC and MLO views of the left and right breasts. To enhance learning efficiency, plug-and-play adapters are added into CLIP image and text encoders for fine-tuning parameters and limiting updates to about 1% of the parameters. For framework evaluation, we assembled two datasets retrospectively. The first dataset, comprising 470 malignant and 479 benign cases, was used for few-shot fine-tuning and internal evaluation of the proposed Mammo-CLIP via 5-fold cross-validation. The second dataset, including 60 malignant and 294 benign cases, was used to test generalizability of Mammo-CLIP. Study results show that Mammo-CLIP outperforms the state-of-art cross-view transformer in AUC (0.841 vs. 0.817, 0.837 vs. 0.807) on both datasets. It also surpasses previous two CLIP-based methods by 20.3% and 14.3%. This study highlights the potential of applying the finetuned vision-language models for developing next-generation, image-text-based CAD schemes of breast cancer.
Image-Domain Material Decomposition for Dual-energy CT using Unsupervised Learning with Data-fidelity Loss
Nov 17, 2023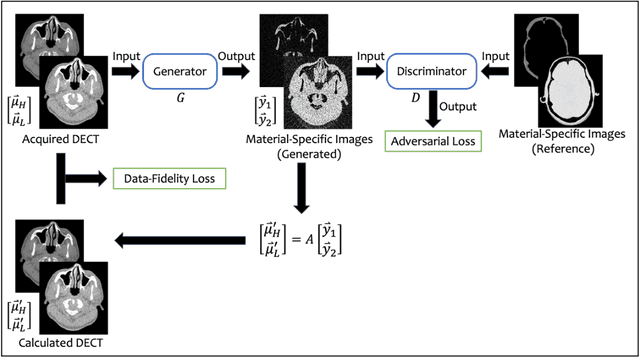
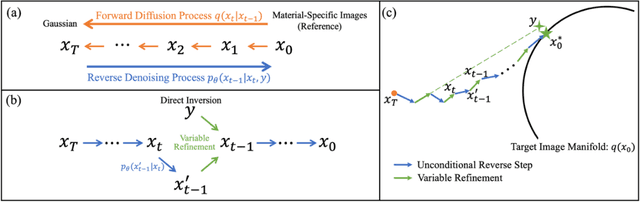
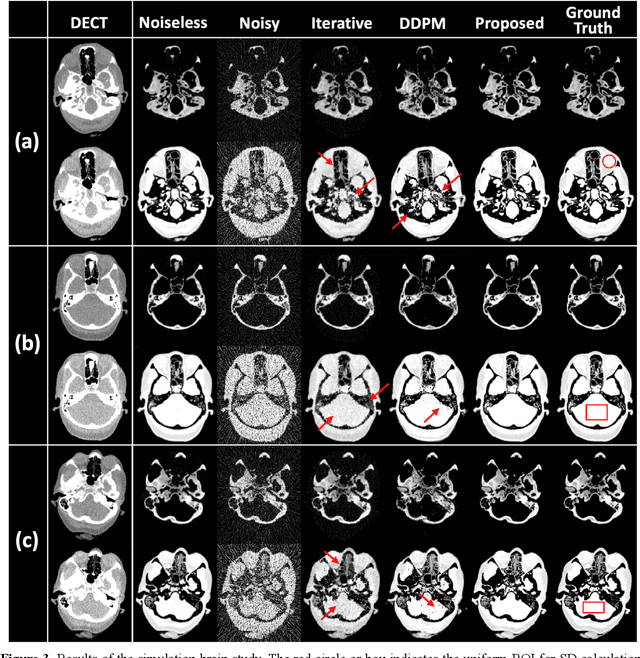
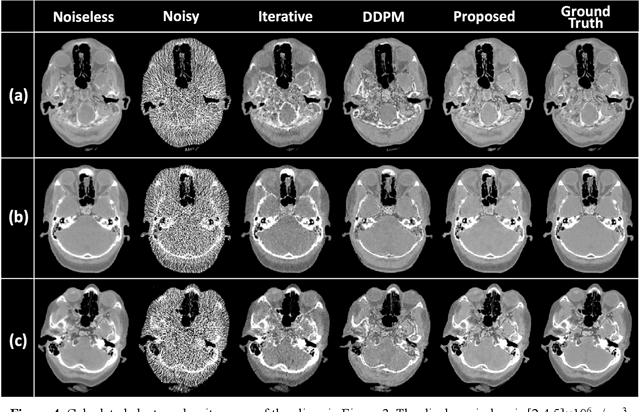
Abstract:Background: Dual-energy CT (DECT) and material decomposition play vital roles in quantitative medical imaging. However, the decomposition process may suffer from significant noise amplification, leading to severely degraded image signal-to-noise ratios (SNRs). While existing iterative algorithms perform noise suppression using different image priors, these heuristic image priors cannot accurately represent the features of the target image manifold. Although deep learning-based decomposition methods have been reported, these methods are in the supervised-learning framework requiring paired data for training, which is not readily available in clinical settings. Purpose: This work aims to develop an unsupervised-learning framework with data-measurement consistency for image-domain material decomposition in DECT.
Synthetic CT Generation from MRI using 3D Transformer-based Denoising Diffusion Model
May 31, 2023


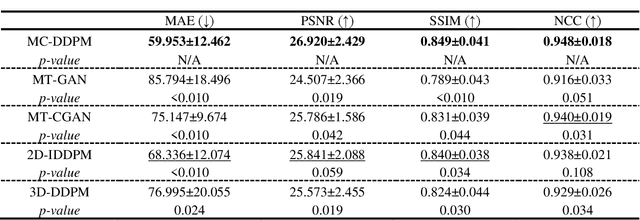
Abstract:Magnetic resonance imaging (MRI)-based synthetic computed tomography (sCT) simplifies radiation therapy treatment planning by eliminating the need for CT simulation and error-prone image registration, ultimately reducing patient radiation dose and setup uncertainty. We propose an MRI-to-CT transformer-based denoising diffusion probabilistic model (MC-DDPM) to transform MRI into high-quality sCT to facilitate radiation treatment planning. MC-DDPM implements diffusion processes with a shifted-window transformer network to generate sCT from MRI. The proposed model consists of two processes: a forward process which adds Gaussian noise to real CT scans, and a reverse process in which a shifted-window transformer V-net (Swin-Vnet) denoises the noisy CT scans conditioned on the MRI from the same patient to produce noise-free CT scans. With an optimally trained Swin-Vnet, the reverse diffusion process was used to generate sCT scans matching MRI anatomy. We evaluated the proposed method by generating sCT from MRI on a brain dataset and a prostate dataset. Qualitative evaluation was performed using the mean absolute error (MAE) of Hounsfield unit (HU), peak signal to noise ratio (PSNR), multi-scale Structure Similarity index (MS-SSIM) and normalized cross correlation (NCC) indexes between ground truth CTs and sCTs. MC-DDPM generated brain sCTs with state-of-the-art quantitative results with MAE 43.317 HU, PSNR 27.046 dB, SSIM 0.965, and NCC 0.983. For the prostate dataset, MC-DDPM achieved MAE 59.953 HU, PSNR 26.920 dB, SSIM 0.849, and NCC 0.948. In conclusion, we have developed and validated a novel approach for generating CT images from routine MRIs using a transformer-based DDPM. This model effectively captures the complex relationship between CT and MRI images, allowing for robust and high-quality synthetic CT (sCT) images to be generated in minutes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge