Pretesh Patel
Efficient Vision Mamba for MRI Super-Resolution via Hybrid Selective Scanning
Dec 22, 2025
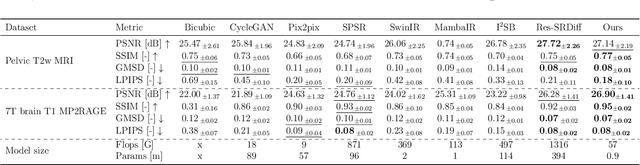
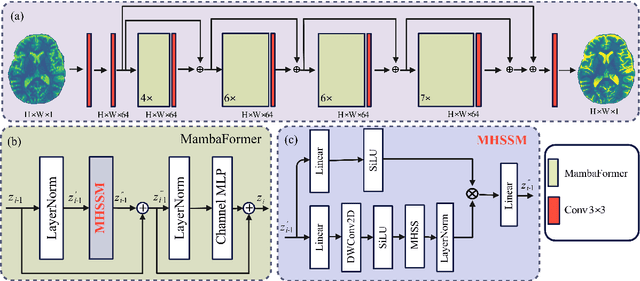

Abstract:Background: High-resolution MRI is critical for diagnosis, but long acquisition times limit clinical use. Super-resolution (SR) can enhance resolution post-scan, yet existing deep learning methods face fidelity-efficiency trade-offs. Purpose: To develop a computationally efficient and accurate deep learning framework for MRI SR that preserves anatomical detail for clinical integration. Materials and Methods: We propose a novel SR framework combining multi-head selective state-space models (MHSSM) with a lightweight channel MLP. The model uses 2D patch extraction with hybrid scanning to capture long-range dependencies. Each MambaFormer block integrates MHSSM, depthwise convolutions, and gated channel mixing. Evaluation used 7T brain T1 MP2RAGE maps (n=142) and 1.5T prostate T2w MRI (n=334). Comparisons included Bicubic interpolation, GANs (CycleGAN, Pix2pix, SPSR), transformers (SwinIR), Mamba (MambaIR), and diffusion models (I2SB, Res-SRDiff). Results: Our model achieved superior performance with exceptional efficiency. For 7T brain data: SSIM=0.951+-0.021, PSNR=26.90+-1.41 dB, LPIPS=0.076+-0.022, GMSD=0.083+-0.017, significantly outperforming all baselines (p<0.001). For prostate data: SSIM=0.770+-0.049, PSNR=27.15+-2.19 dB, LPIPS=0.190+-0.095, GMSD=0.087+-0.013. The framework used only 0.9M parameters and 57 GFLOPs, reducing parameters by 99.8% and computation by 97.5% versus Res-SRDiff, while outperforming SwinIR and MambaIR in accuracy and efficiency. Conclusion: The proposed framework provides an efficient, accurate MRI SR solution, delivering enhanced anatomical detail across datasets. Its low computational demand and state-of-the-art performance show strong potential for clinical translation.
Diffeomorphic Transformer-based Abdomen MRI-CT Deformable Image Registration
May 04, 2024
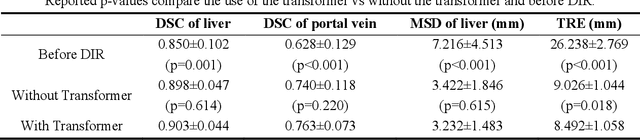



Abstract:This paper aims to create a deep learning framework that can estimate the deformation vector field (DVF) for directly registering abdominal MRI-CT images. The proposed method assumed a diffeomorphic deformation. By using topology-preserved deformation features extracted from the probabilistic diffeomorphic registration model, abdominal motion can be accurately obtained and utilized for DVF estimation. The model integrated Swin transformers, which have demonstrated superior performance in motion tracking, into the convolutional neural network (CNN) for deformation feature extraction. The model was optimized using a cross-modality image similarity loss and a surface matching loss. To compute the image loss, a modality-independent neighborhood descriptor (MIND) was used between the deformed MRI and CT images. The surface matching loss was determined by measuring the distance between the warped coordinates of the surfaces of contoured structures on the MRI and CT images. The deformed MRI image was assessed against the CT image using the target registration error (TRE), Dice similarity coefficient (DSC), and mean surface distance (MSD) between the deformed contours of the MRI image and manual contours of the CT image. When compared to only rigid registration, DIR with the proposed method resulted in an increase of the mean DSC values of the liver and portal vein from 0.850 and 0.628 to 0.903 and 0.763, a decrease of the mean MSD of the liver from 7.216 mm to 3.232 mm, and a decrease of the TRE from 26.238 mm to 8.492 mm. The proposed deformable image registration method based on a diffeomorphic transformer provides an effective and efficient way to generate an accurate DVF from an MRI-CT image pair of the abdomen. It could be utilized in the current treatment planning workflow for liver radiotherapy.
Synthetic CT Generation from MRI using 3D Transformer-based Denoising Diffusion Model
May 31, 2023


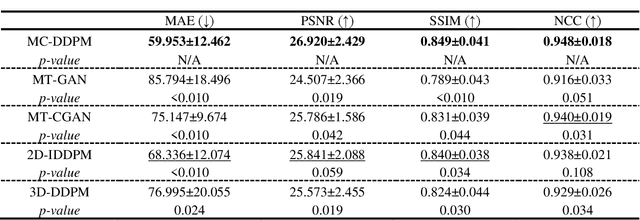
Abstract:Magnetic resonance imaging (MRI)-based synthetic computed tomography (sCT) simplifies radiation therapy treatment planning by eliminating the need for CT simulation and error-prone image registration, ultimately reducing patient radiation dose and setup uncertainty. We propose an MRI-to-CT transformer-based denoising diffusion probabilistic model (MC-DDPM) to transform MRI into high-quality sCT to facilitate radiation treatment planning. MC-DDPM implements diffusion processes with a shifted-window transformer network to generate sCT from MRI. The proposed model consists of two processes: a forward process which adds Gaussian noise to real CT scans, and a reverse process in which a shifted-window transformer V-net (Swin-Vnet) denoises the noisy CT scans conditioned on the MRI from the same patient to produce noise-free CT scans. With an optimally trained Swin-Vnet, the reverse diffusion process was used to generate sCT scans matching MRI anatomy. We evaluated the proposed method by generating sCT from MRI on a brain dataset and a prostate dataset. Qualitative evaluation was performed using the mean absolute error (MAE) of Hounsfield unit (HU), peak signal to noise ratio (PSNR), multi-scale Structure Similarity index (MS-SSIM) and normalized cross correlation (NCC) indexes between ground truth CTs and sCTs. MC-DDPM generated brain sCTs with state-of-the-art quantitative results with MAE 43.317 HU, PSNR 27.046 dB, SSIM 0.965, and NCC 0.983. For the prostate dataset, MC-DDPM achieved MAE 59.953 HU, PSNR 26.920 dB, SSIM 0.849, and NCC 0.948. In conclusion, we have developed and validated a novel approach for generating CT images from routine MRIs using a transformer-based DDPM. This model effectively captures the complex relationship between CT and MRI images, allowing for robust and high-quality synthetic CT (sCT) images to be generated in minutes.
Landmark Tracking in Liver US images Using Cascade Convolutional Neural Networks with Long Short-Term Memory
Sep 14, 2022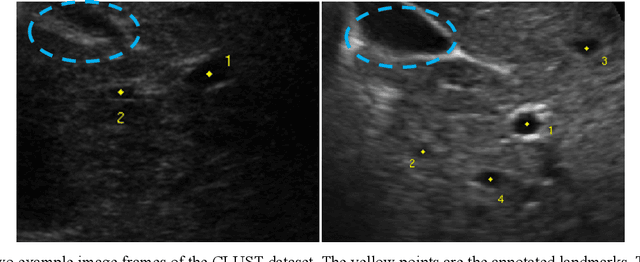
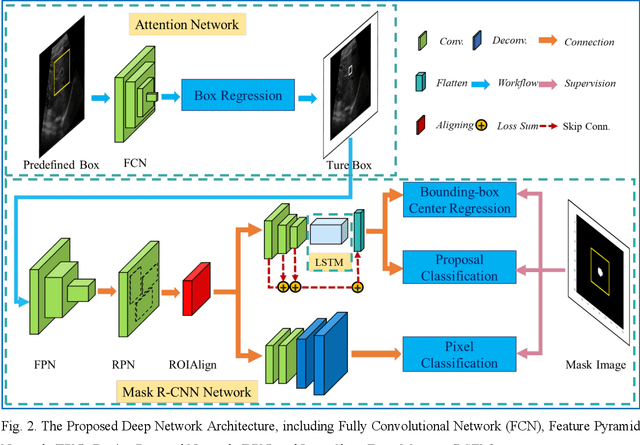
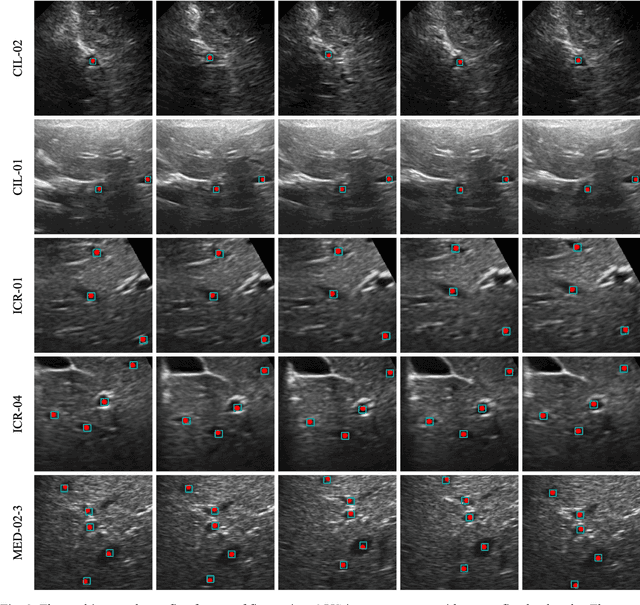
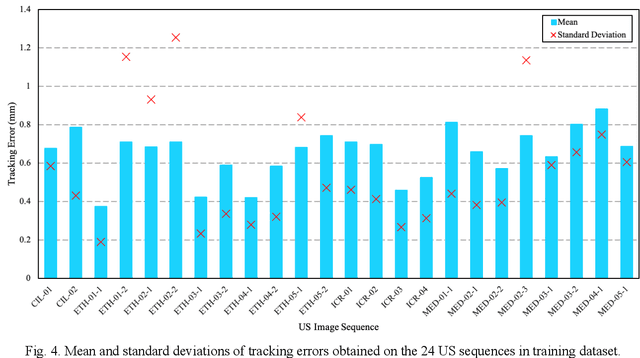
Abstract:This study proposed a deep learning-based tracking method for ultrasound (US) image-guided radiation therapy. The proposed cascade deep learning model is composed of an attention network, a mask region-based convolutional neural network (mask R-CNN), and a long short-term memory (LSTM) network. The attention network learns a mapping from a US image to a suspected area of landmark motion in order to reduce the search region. The mask R-CNN then produces multiple region-of-interest (ROI) proposals in the reduced region and identifies the proposed landmark via three network heads: bounding box regression, proposal classification, and landmark segmentation. The LSTM network models the temporal relationship among the successive image frames for bounding box regression and proposal classification. To consolidate the final proposal, a selection method is designed according to the similarities between sequential frames. The proposed method was tested on the liver US tracking datasets used in the Medical Image Computing and Computer Assisted Interventions (MICCAI) 2015 challenges, where the landmarks were annotated by three experienced observers to obtain their mean positions. Five-fold cross-validation on the 24 given US sequences with ground truths shows that the mean tracking error for all landmarks is 0.65+/-0.56 mm, and the errors of all landmarks are within 2 mm. We further tested the proposed model on 69 landmarks from the testing dataset that has a similar image pattern to the training pattern, resulting in a mean tracking error of 0.94+/-0.83 mm. Our experimental results have demonstrated the feasibility and accuracy of our proposed method in tracking liver anatomic landmarks using US images, providing a potential solution for real-time liver tracking for active motion management during radiation therapy.
Deformable Image Registration using Unsupervised Deep Learning for CBCT-guided Abdominal Radiotherapy
Aug 29, 2022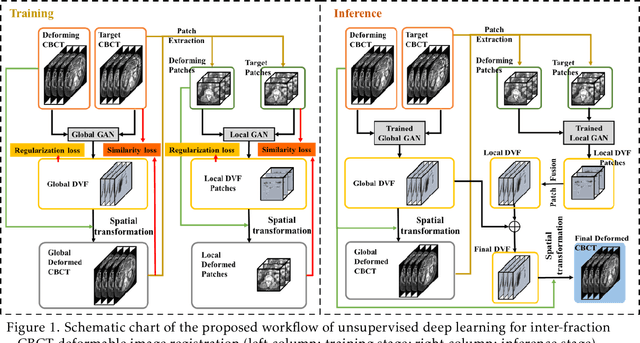
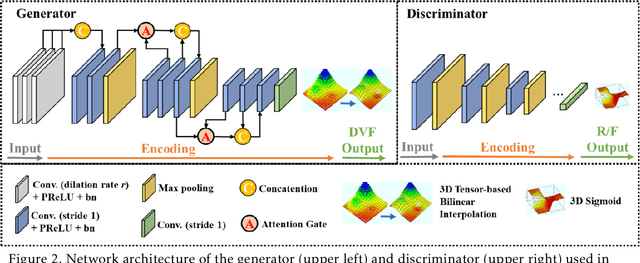
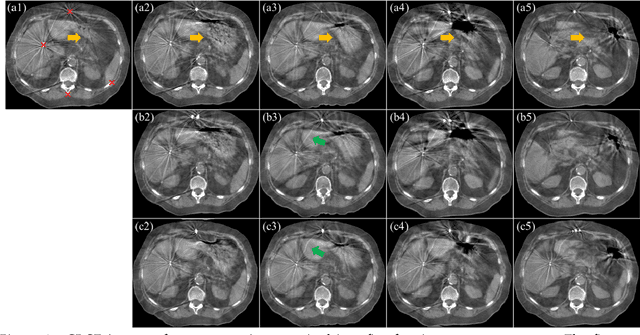
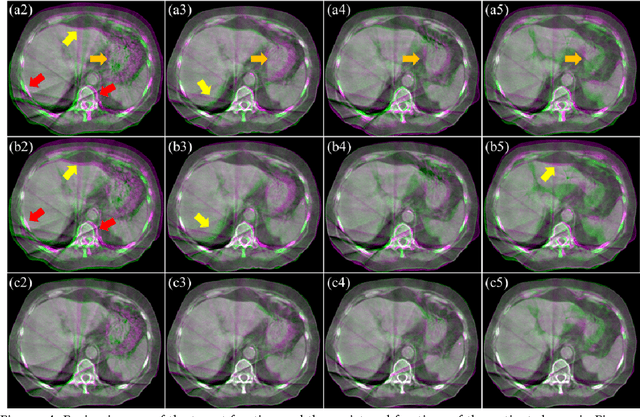
Abstract:CBCTs in image-guided radiotherapy provide crucial anatomy information for patient setup and plan evaluation. Longitudinal CBCT image registration could quantify the inter-fractional anatomic changes. The purpose of this study is to propose an unsupervised deep learning based CBCT-CBCT deformable image registration. The proposed deformable registration workflow consists of training and inference stages that share the same feed-forward path through a spatial transformation-based network (STN). The STN consists of a global generative adversarial network (GlobalGAN) and a local GAN (LocalGAN) to predict the coarse- and fine-scale motions, respectively. The network was trained by minimizing the image similarity loss and the deformable vector field (DVF) regularization loss without the supervision of ground truth DVFs. During the inference stage, patches of local DVF were predicted by the trained LocalGAN and fused to form a whole-image DVF. The local whole-image DVF was subsequently combined with the GlobalGAN generated DVF to obtain final DVF. The proposed method was evaluated using 100 fractional CBCTs from 20 abdominal cancer patients in the experiments and 105 fractional CBCTs from a cohort of 21 different abdominal cancer patients in a holdout test. Qualitatively, the registration results show great alignment between the deformed CBCT images and the target CBCT image. Quantitatively, the average target registration error (TRE) calculated on the fiducial markers and manually identified landmarks was 1.91+-1.11 mm. The average mean absolute error (MAE), normalized cross correlation (NCC) between the deformed CBCT and target CBCT were 33.42+-7.48 HU, 0.94+-0.04, respectively. This promising registration method could provide fast and accurate longitudinal CBCT alignment to facilitate inter-fractional anatomic changes analysis and prediction.
Fuzzy Inference Systems Optimization
Oct 15, 2011
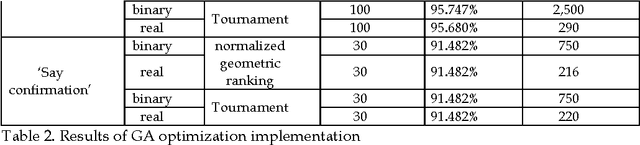


Abstract:This paper compares various optimization methods for fuzzy inference system optimization. The optimization methods compared are genetic algorithm, particle swarm optimization and simulated annealing. When these techniques were implemented it was observed that the performance of each technique within the fuzzy inference system classification was context dependent.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge