Jikai Zhang
Duke University, Durham NC, Duke Center for Artificial Intelligence in Radiology, Durham NC
Agentic Automation of BT-RADS Scoring: End-to-End Multi-Agent System for Standardized Brain Tumor Follow-up Assessment
Mar 23, 2026Abstract:The Brain Tumor Reporting and Data System (BT-RADS) standardizes post-treatment MRI response assessment in patients with diffuse gliomas but requires complex integration of imaging trends, medication effects, and radiation timing. This study evaluates an end-to-end multi-agent large language model (LLM) and convolutional neural network (CNN) system for automated BT-RADS classification. A multi-agent LLM system combined with automated CNN-based tumor segmentation was retrospectively evaluated on 509 consecutive post-treatment glioma MRI examinations from a single high-volume center. An extractor agent identified clinical variables (steroid status, bevacizumab status, radiation date) from unstructured clinical notes, while a scorer agent applied BT-RADS decision logic integrating extracted variables with volumetric measurements. Expert reference standard classifications were established by an independent board-certified neuroradiologist. Of 509 examinations, 492 met inclusion criteria. The system achieved 374/492 (76.0%; 95% CI, 72.1%-79.6%) accuracy versus 283/492 (57.5%; 95% CI, 53.1%-61.8%) for initial clinical assessments (+18.5 percentage points; P<.001). Context-dependent categories showed high sensitivity (BT-1b 100%, BT-1a 92.7%, BT-3a 87.5%), while threshold-dependent categories showed moderate sensitivity (BT-3c 74.8%, BT-2 69.2%, BT-4 69.3%, BT-3b 57.1%). For BT-4, positive predictive value was 92.9%. The multi-agent LLM system achieved higher BT-RADS classification agreement with expert reference standard compared to initial clinical scoring, with high accuracy for context-dependent scores and high positive predictive value for BT-4 detection.
Effectiveness of Automatically Curated Dataset in Thyroid Nodules Classification Algorithms Using Deep Learning
Feb 01, 2026Abstract:The diagnosis of thyroid nodule cancers commonly utilizes ultrasound images. Several studies showed that deep learning algorithms designed to classify benign and malignant thyroid nodules could match radiologists' performance. However, data availability for training deep learning models is often limited due to the significant effort required to curate such datasets. The previous study proposed a method to curate thyroid nodule datasets automatically. It was tested to have a 63% yield rate and 83% accuracy. However, the usefulness of the generated data for training deep learning models remains unknown. In this study, we conducted experiments to determine whether using a automatically-curated dataset improves deep learning algorithms' performance. We trained deep learning models on the manually annotated and automatically-curated datasets. We also trained with a smaller subset of the automatically-curated dataset that has higher accuracy to explore the optimum usage of such dataset. As a result, the deep learning model trained on the manually selected dataset has an AUC of 0.643 (95% confidence interval [CI]: 0.62, 0.66). It is significantly lower than the AUC of the 6automatically-curated dataset trained deep learning model, 0.694 (95% confidence interval [CI]: 0.67, 0.73, P < .001). The AUC of the accurate subset trained deep learning model is 0.689 (95% confidence interval [CI]: 0.66, 0.72, P > .43), which is insignificantly worse than the AUC of the full automatically-curated dataset. In conclusion, we showed that using a automatically-curated dataset can substantially increase the performance of deep learning algorithms, and it is suggested to use all the data rather than only using the accurate subset.
Language Models and Retrieval Augmented Generation for Automated Structured Data Extraction from Diagnostic Reports
Sep 18, 2024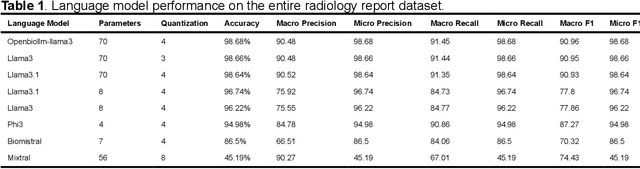

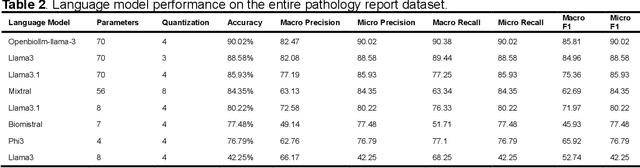

Abstract:Purpose: To develop and evaluate an automated system for extracting structured clinical information from unstructured radiology and pathology reports using open-weights large language models (LMs) and retrieval augmented generation (RAG), and to assess the effects of model configuration variables on extraction performance. Methods and Materials: The study utilized two datasets: 7,294 radiology reports annotated for Brain Tumor Reporting and Data System (BT-RADS) scores and 2,154 pathology reports annotated for isocitrate dehydrogenase (IDH) mutation status. An automated pipeline was developed to benchmark the performance of various LMs and RAG configurations. The impact of model size, quantization, prompting strategies, output formatting, and inference parameters was systematically evaluated. Results: The best performing models achieved over 98% accuracy in extracting BT-RADS scores from radiology reports and over 90% for IDH mutation status extraction from pathology reports. The top model being medical fine-tuned llama3. Larger, newer, and domain fine-tuned models consistently outperformed older and smaller models. Model quantization had minimal impact on performance. Few-shot prompting significantly improved accuracy. RAG improved performance for complex pathology reports but not for shorter radiology reports. Conclusions: Open LMs demonstrate significant potential for automated extraction of structured clinical data from unstructured clinical reports with local privacy-preserving application. Careful model selection, prompt engineering, and semi-automated optimization using annotated data are critical for optimal performance. These approaches could be reliable enough for practical use in research workflows, highlighting the potential for human-machine collaboration in healthcare data extraction.
The 2024 Brain Tumor Segmentation (BraTS) Challenge: Glioma Segmentation on Post-treatment MRI
May 28, 2024
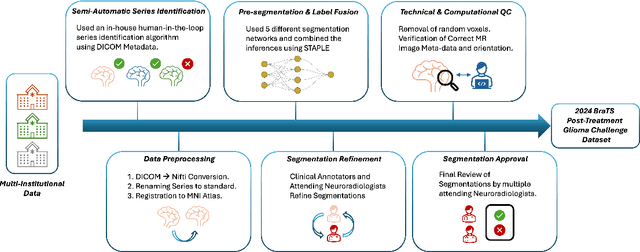
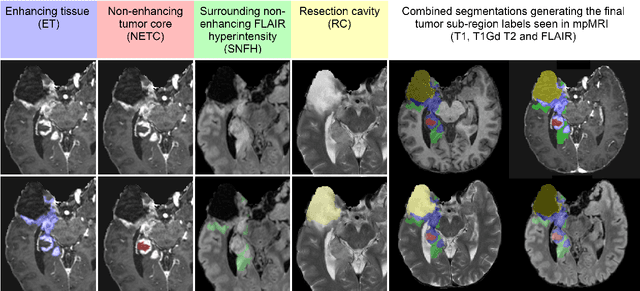
Abstract:Gliomas are the most common malignant primary brain tumors in adults and one of the deadliest types of cancer. There are many challenges in treatment and monitoring due to the genetic diversity and high intrinsic heterogeneity in appearance, shape, histology, and treatment response. Treatments include surgery, radiation, and systemic therapies, with magnetic resonance imaging (MRI) playing a key role in treatment planning and post-treatment longitudinal assessment. The 2024 Brain Tumor Segmentation (BraTS) challenge on post-treatment glioma MRI will provide a community standard and benchmark for state-of-the-art automated segmentation models based on the largest expert-annotated post-treatment glioma MRI dataset. Challenge competitors will develop automated segmentation models to predict four distinct tumor sub-regions consisting of enhancing tissue (ET), surrounding non-enhancing T2/fluid-attenuated inversion recovery (FLAIR) hyperintensity (SNFH), non-enhancing tumor core (NETC), and resection cavity (RC). Models will be evaluated on separate validation and test datasets using standardized performance metrics utilized across the BraTS 2024 cluster of challenges, including lesion-wise Dice Similarity Coefficient and Hausdorff Distance. Models developed during this challenge will advance the field of automated MRI segmentation and contribute to their integration into clinical practice, ultimately enhancing patient care.
Deep learning automates Cobb angle measurement compared with multi-expert observers
Mar 18, 2024Abstract:Scoliosis, a prevalent condition characterized by abnormal spinal curvature leading to deformity, requires precise assessment methods for effective diagnosis and management. The Cobb angle is a widely used scoliosis quantification method that measures the degree of curvature between the tilted vertebrae. Yet, manual measuring of Cobb angles is time-consuming and labor-intensive, fraught with significant interobserver and intraobserver variability. To address these challenges and the lack of interpretability found in certain existing automated methods, we have created fully automated software that not only precisely measures the Cobb angle but also provides clear visualizations of these measurements. This software integrates deep neural network-based spine region detection and segmentation, spine centerline identification, pinpointing the most significantly tilted vertebrae, and direct visualization of Cobb angles on the original images. Upon comparison with the assessments of 7 expert readers, our algorithm exhibited a mean deviation in Cobb angle measurements of 4.17 degrees, notably surpassing the manual approach's average intra-reader discrepancy of 5.16 degrees. The algorithm also achieved intra-class correlation coefficients (ICC) exceeding 0.96 and Pearson correlation coefficients above 0.944, reflecting robust agreement with expert assessments and superior measurement reliability. Through the comprehensive reader study and statistical analysis, we believe this algorithm not only ensures a higher consensus with expert readers but also enhances interpretability and reproducibility during assessments. It holds significant promise for clinical application, potentially aiding physicians in more accurate scoliosis assessment and diagnosis, thereby improving patient care.
SegmentAnyBone: A Universal Model that Segments Any Bone at Any Location on MRI
Jan 23, 2024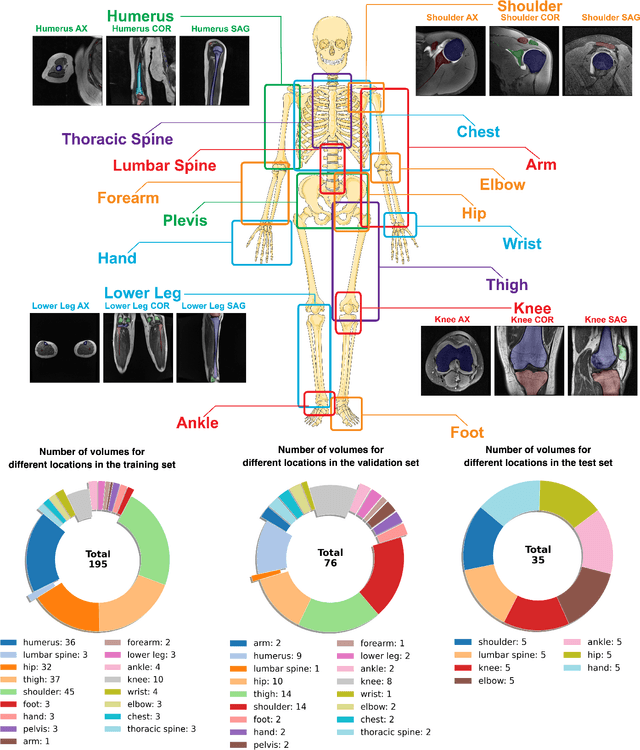
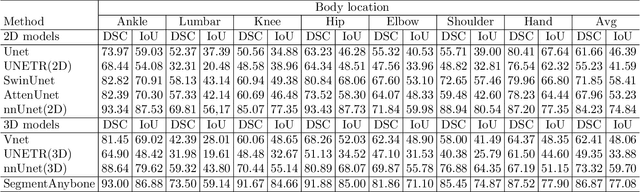
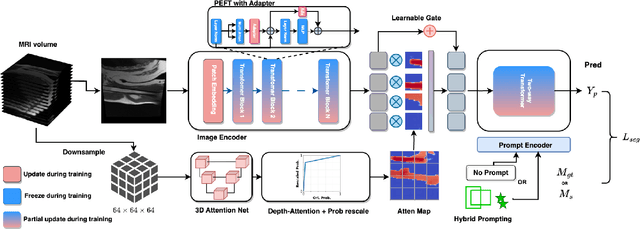

Abstract:Magnetic Resonance Imaging (MRI) is pivotal in radiology, offering non-invasive and high-quality insights into the human body. Precise segmentation of MRIs into different organs and tissues would be highly beneficial since it would allow for a higher level of understanding of the image content and enable important measurements, which are essential for accurate diagnosis and effective treatment planning. Specifically, segmenting bones in MRI would allow for more quantitative assessments of musculoskeletal conditions, while such assessments are largely absent in current radiological practice. The difficulty of bone MRI segmentation is illustrated by the fact that limited algorithms are publicly available for use, and those contained in the literature typically address a specific anatomic area. In our study, we propose a versatile, publicly available deep-learning model for bone segmentation in MRI across multiple standard MRI locations. The proposed model can operate in two modes: fully automated segmentation and prompt-based segmentation. Our contributions include (1) collecting and annotating a new MRI dataset across various MRI protocols, encompassing over 300 annotated volumes and 8485 annotated slices across diverse anatomic regions; (2) investigating several standard network architectures and strategies for automated segmentation; (3) introducing SegmentAnyBone, an innovative foundational model-based approach that extends Segment Anything Model (SAM); (4) comparative analysis of our algorithm and previous approaches; and (5) generalization analysis of our algorithm across different anatomical locations and MRI sequences, as well as an external dataset. We publicly release our model at https://github.com/mazurowski-lab/SegmentAnyBone.
Improving Image Classification of Knee Radiographs: An Automated Image Labeling Approach
Sep 06, 2023Abstract:Large numbers of radiographic images are available in knee radiology practices which could be used for training of deep learning models for diagnosis of knee abnormalities. However, those images do not typically contain readily available labels due to limitations of human annotations. The purpose of our study was to develop an automated labeling approach that improves the image classification model to distinguish normal knee images from those with abnormalities or prior arthroplasty. The automated labeler was trained on a small set of labeled data to automatically label a much larger set of unlabeled data, further improving the image classification performance for knee radiographic diagnosis. We developed our approach using 7,382 patients and validated it on a separate set of 637 patients. The final image classification model, trained using both manually labeled and pseudo-labeled data, had the higher weighted average AUC (WAUC: 0.903) value and higher AUC-ROC values among all classes (normal AUC-ROC: 0.894; abnormal AUC-ROC: 0.896, arthroplasty AUC-ROC: 0.990) compared to the baseline model (WAUC=0.857; normal AUC-ROC: 0.842; abnormal AUC-ROC: 0.848, arthroplasty AUC-ROC: 0.987), trained using only manually labeled data. DeLong tests show that the improvement is significant on normal (p-value<0.002) and abnormal (p-value<0.001) images. Our findings demonstrated that the proposed automated labeling approach significantly improves the performance of image classification for radiographic knee diagnosis, allowing for facilitating patient care and curation of large knee datasets.
Deep Learning for Classification of Thyroid Nodules on Ultrasound: Validation on an Independent Dataset
Jul 27, 2022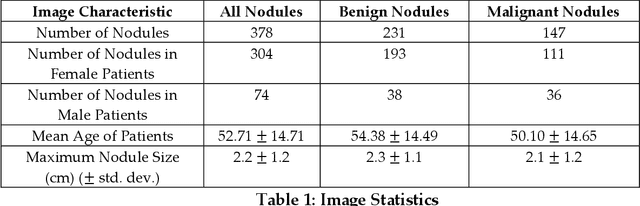
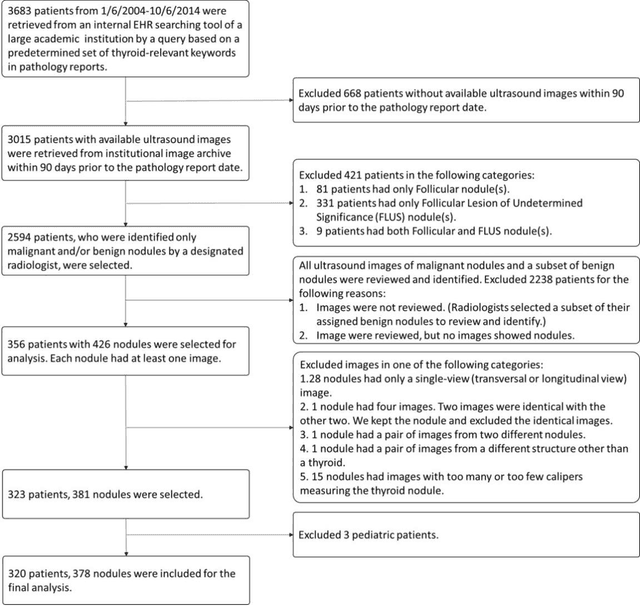
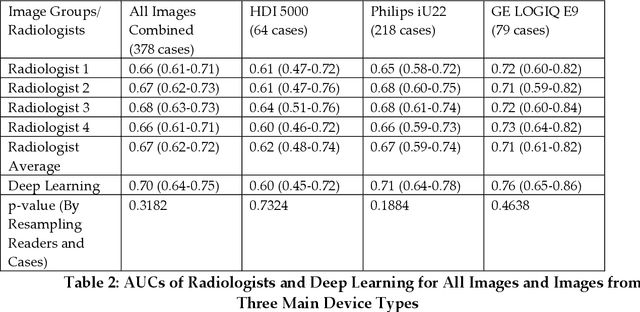
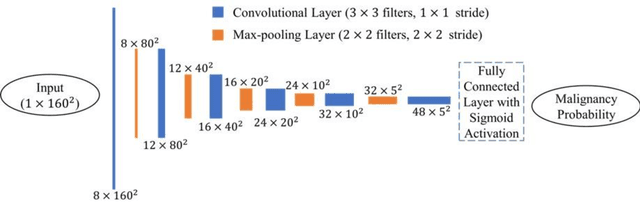
Abstract:Objectives: The purpose is to apply a previously validated deep learning algorithm to a new thyroid nodule ultrasound image dataset and compare its performances with radiologists. Methods: Prior study presented an algorithm which is able to detect thyroid nodules and then make malignancy classifications with two ultrasound images. A multi-task deep convolutional neural network was trained from 1278 nodules and originally tested with 99 separate nodules. The results were comparable with that of radiologists. The algorithm was further tested with 378 nodules imaged with ultrasound machines from different manufacturers and product types than the training cases. Four experienced radiologists were requested to evaluate the nodules for comparison with deep learning. Results: The Area Under Curve (AUC) of the deep learning algorithm and four radiologists were calculated with parametric, binormal estimation. For the deep learning algorithm, the AUC was 0.70 (95% CI: 0.64 - 0.75). The AUC of radiologists were 0.66 (95% CI: 0.61 - 0.71), 0.67 (95% CI:0.62 - 0.73), 0.68 (95% CI: 0.63 - 0.73), and 0.66 (95%CI: 0.61 - 0.71). Conclusion: In the new testing dataset, the deep learning algorithm achieved similar performances with all four radiologists.
Multistep Automated Data Labelling Procedure (MADLaP) for Thyroid Nodules on Ultrasound: An Artificial Intelligence Approach for Automating Image Annotation
Jun 28, 2022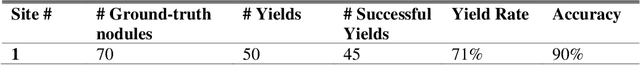
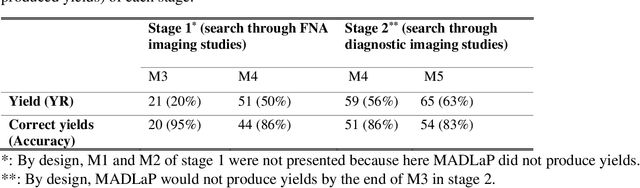
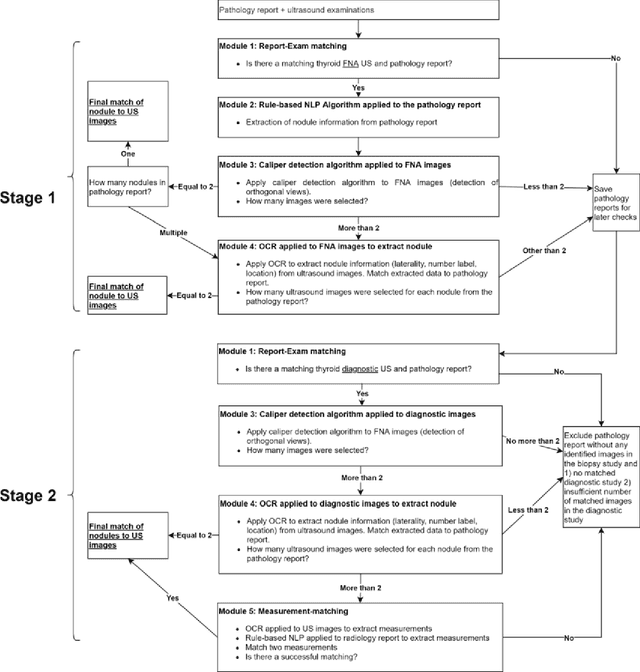


Abstract:Machine learning (ML) for diagnosis of thyroid nodules on ultrasound is an active area of research. However, ML tools require large, well-labelled datasets, the curation of which is time-consuming and labor-intensive. The purpose of our study was to develop and test a deep-learning-based tool to facilitate and automate the data annotation process for thyroid nodules; we named our tool Multistep Automated Data Labelling Procedure (MADLaP). MADLaP was designed to take multiple inputs included pathology reports, ultrasound images, and radiology reports. Using multiple step-wise modules including rule-based natural language processing, deep-learning-based imaging segmentation, and optical character recognition, MADLaP automatically identified images of a specific thyroid nodule and correctly assigned a pathology label. The model was developed using a training set of 378 patients across our health system and tested on a separate set of 93 patients. Ground truths for both sets were selected by an experienced radiologist. Performance metrics including yield (how many labeled images the model produced) and accuracy (percentage correct) were measured using the test set. MADLaP achieved a yield of 63% and an accuracy of 83%. The yield progressively increased as the input data moved through each module, while accuracy peaked part way through. Error analysis showed that inputs from certain examination sites had lower accuracy (40%) than the other sites (90%, 100%). MADLaP successfully created curated datasets of labeled ultrasound images of thyroid nodules. While accurate, the relatively suboptimal yield of MADLaP exposed some challenges when trying to automatically label radiology images from heterogeneous sources. The complex task of image curation and annotation could be automated, allowing for enrichment of larger datasets for use in machine learning development.
Automated Grading of Radiographic Knee Osteoarthritis Severity Combined with Joint Space Narrowing
Mar 16, 2022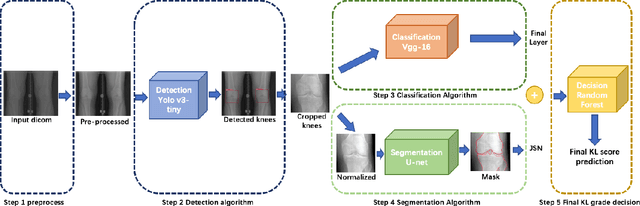
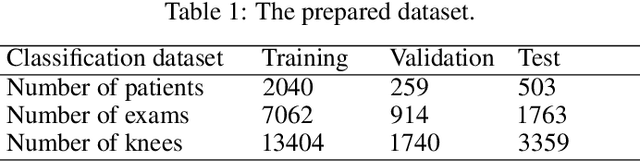
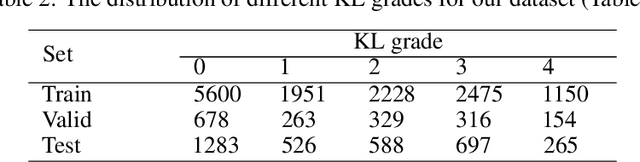
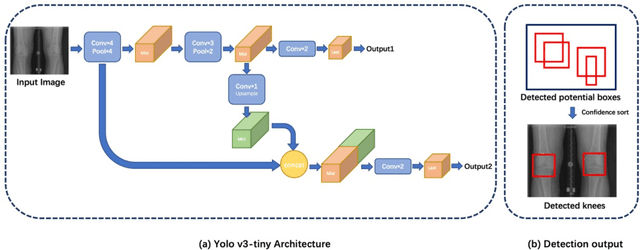
Abstract:The assessment of knee osteoarthritis (KOA) severity on knee X-rays is a central criteria for the use of total knee arthroplasty. However, this assessment suffers from imprecise standards and a remarkably high inter-reader variability. An algorithmic, automated assessment of KOA severity could improve overall outcomes of knee replacement procedures by increasing the appropriateness of its use. We propose a novel deep learning-based five-step algorithm to automatically grade KOA from posterior-anterior (PA) views of radiographs: (1) image preprocessing (2) localization of knees joints in the image using the YOLO v3-Tiny model, (3) initial assessment of the severity of osteoarthritis using a convolutional neural network-based classifier, (4) segmentation of the joints and calculation of the joint space narrowing (JSN), and (5), a combination of the JSN and the initial assessment to determine a final Kellgren-Lawrence (KL) score. Furthermore, by displaying the segmentation masks used to make the assessment, our algorithm demonstrates a higher degree of transparency compared to typical "black box" deep learning classifiers. We perform a comprehensive evaluation using two public datasets and one dataset from our institution, and show that our algorithm reaches state-of-the art performance. Moreover, we also collected ratings from multiple radiologists at our institution and showed that our algorithm performs at the radiologist level. The software has been made publicly available at https://github.com/MaciejMazurowski/osteoarthritis-classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge