Yaqian Chen
BreastSegNet: Multi-label Segmentation of Breast MRI
Jul 18, 2025Abstract:Breast MRI provides high-resolution imaging critical for breast cancer screening and preoperative staging. However, existing segmentation methods for breast MRI remain limited in scope, often focusing on only a few anatomical structures, such as fibroglandular tissue or tumors, and do not cover the full range of tissues seen in scans. This narrows their utility for quantitative analysis. In this study, we present BreastSegNet, a multi-label segmentation algorithm for breast MRI that covers nine anatomical labels: fibroglandular tissue (FGT), vessel, muscle, bone, lesion, lymph node, heart, liver, and implant. We manually annotated a large set of 1123 MRI slices capturing these structures with detailed review and correction from an expert radiologist. Additionally, we benchmark nine segmentation models, including U-Net, SwinUNet, UNet++, SAM, MedSAM, and nnU-Net with multiple ResNet-based encoders. Among them, nnU-Net ResEncM achieves the highest average Dice scores of 0.694 across all labels. It performs especially well on heart, liver, muscle, FGT, and bone, with Dice scores exceeding 0.73, and approaching 0.90 for heart and liver. All model code and weights are publicly available, and we plan to release the data at a later date.
Improving Surgical Risk Prediction Through Integrating Automated Body Composition Analysis: a Retrospective Trial on Colectomy Surgery
Jun 16, 2025Abstract:Objective: To evaluate whether preoperative body composition metrics automatically extracted from CT scans can predict postoperative outcomes after colectomy, either alone or combined with clinical variables or existing risk predictors. Main outcomes and measures: The primary outcome was the predictive performance for 1-year all-cause mortality following colectomy. A Cox proportional hazards model with 1-year follow-up was used, and performance was evaluated using the concordance index (C-index) and Integrated Brier Score (IBS). Secondary outcomes included postoperative complications, unplanned readmission, blood transfusion, and severe infection, assessed using AUC and Brier Score from logistic regression. Odds ratios (OR) described associations between individual CT-derived body composition metrics and outcomes. Over 300 features were extracted from preoperative CTs across multiple vertebral levels, including skeletal muscle area, density, fat areas, and inter-tissue metrics. NSQIP scores were available for all surgeries after 2012.
MRI-CORE: A Foundation Model for Magnetic Resonance Imaging
Jun 13, 2025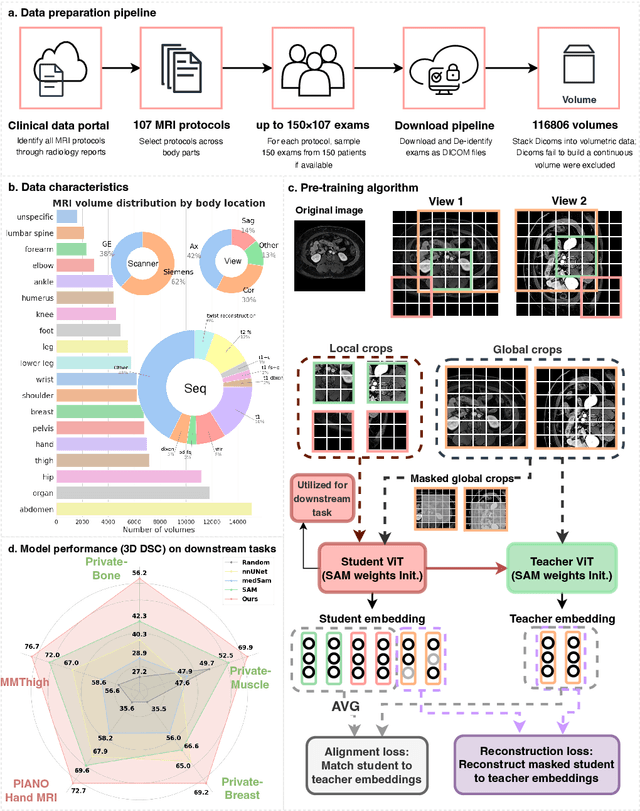
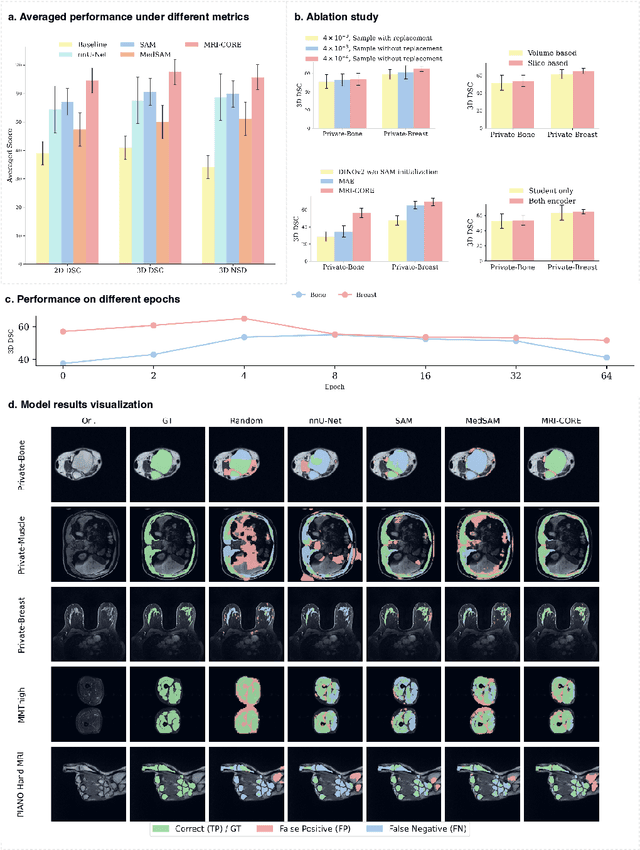
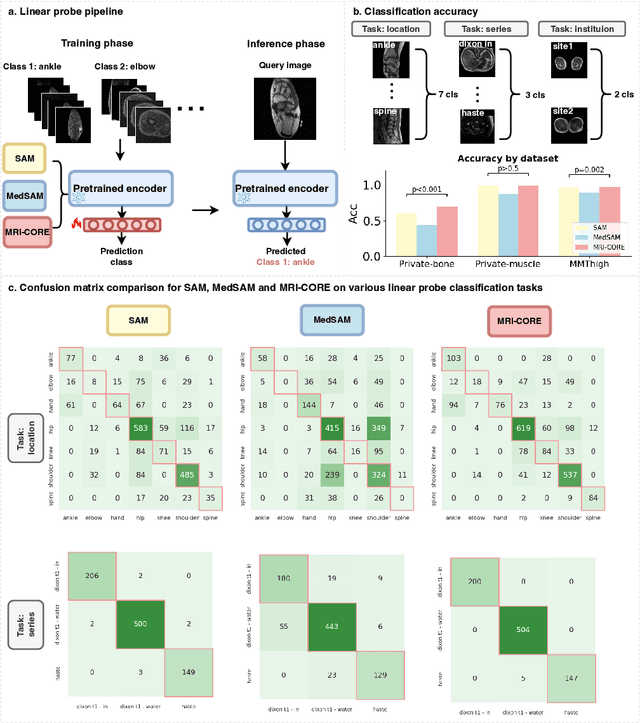
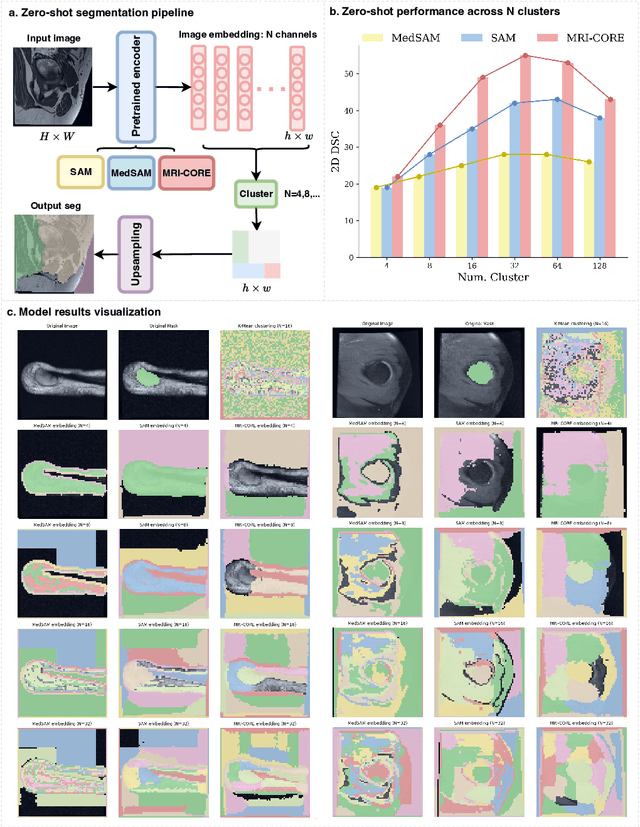
Abstract:The widespread use of Magnetic Resonance Imaging (MRI) and the rise of deep learning have enabled the development of powerful predictive models for a wide range of diagnostic tasks in MRI, such as image classification or object segmentation. However, training models for specific new tasks often requires large amounts of labeled data, which is difficult to obtain due to high annotation costs and data privacy concerns. To circumvent this issue, we introduce MRI-CORE (MRI COmprehensive Representation Encoder), a vision foundation model pre-trained using more than 6 million slices from over 110,000 MRI volumes across 18 main body locations. Experiments on five diverse object segmentation tasks in MRI demonstrate that MRI-CORE can significantly improve segmentation performance in realistic scenarios with limited labeled data availability, achieving an average gain of 6.97% 3D Dice Coefficient using only 10 annotated slices per task. We further demonstrate new model capabilities in MRI such as classification of image properties including body location, sequence type and institution, and zero-shot segmentation. These results highlight the value of MRI-CORE as a generalist vision foundation model for MRI, potentially lowering the data annotation resource barriers for many applications.
GuidedMorph: Two-Stage Deformable Registration for Breast MRI
May 19, 2025
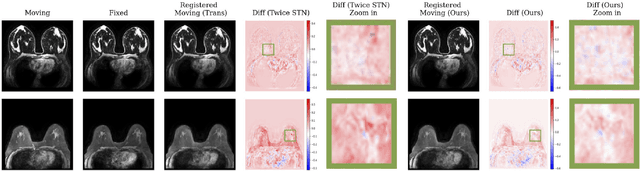
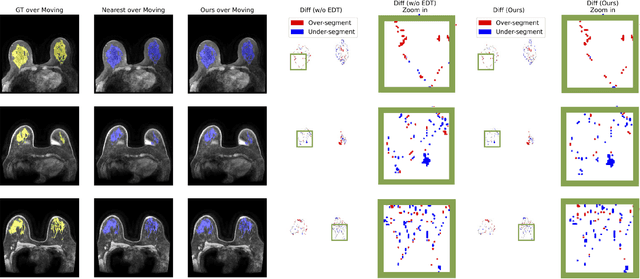
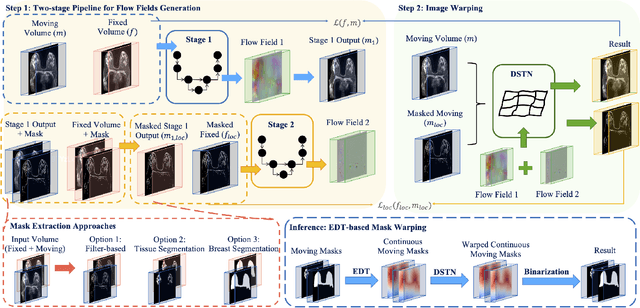
Abstract:Accurately registering breast MR images from different time points enables the alignment of anatomical structures and tracking of tumor progression, supporting more effective breast cancer detection, diagnosis, and treatment planning. However, the complexity of dense tissue and its highly non-rigid nature pose challenges for conventional registration methods, which primarily focus on aligning general structures while overlooking intricate internal details. To address this, we propose \textbf{GuidedMorph}, a novel two-stage registration framework designed to better align dense tissue. In addition to a single-scale network for global structure alignment, we introduce a framework that utilizes dense tissue information to track breast movement. The learned transformation fields are fused by introducing the Dual Spatial Transformer Network (DSTN), improving overall alignment accuracy. A novel warping method based on the Euclidean distance transform (EDT) is also proposed to accurately warp the registered dense tissue and breast masks, preserving fine structural details during deformation. The framework supports paradigms that require external segmentation models and with image data only. It also operates effectively with the VoxelMorph and TransMorph backbones, offering a versatile solution for breast registration. We validate our method on ISPY2 and internal dataset, demonstrating superior performance in dense tissue, overall breast alignment, and breast structural similarity index measure (SSIM), with notable improvements by over 13.01% in dense tissue Dice, 3.13% in breast Dice, and 1.21% in breast SSIM compared to the best learning-based baseline.
Accelerating Volumetric Medical Image Annotation via Short-Long Memory SAM 2
May 03, 2025Abstract:Manual annotation of volumetric medical images, such as magnetic resonance imaging (MRI) and computed tomography (CT), is a labor-intensive and time-consuming process. Recent advancements in foundation models for video object segmentation, such as Segment Anything Model 2 (SAM 2), offer a potential opportunity to significantly speed up the annotation process by manually annotating one or a few slices and then propagating target masks across the entire volume. However, the performance of SAM 2 in this context varies. Our experiments show that relying on a single memory bank and attention module is prone to error propagation, particularly at boundary regions where the target is present in the previous slice but absent in the current one. To address this problem, we propose Short-Long Memory SAM 2 (SLM-SAM 2), a novel architecture that integrates distinct short-term and long-term memory banks with separate attention modules to improve segmentation accuracy. We evaluate SLM-SAM 2 on three public datasets covering organs, bones, and muscles across MRI and CT modalities. We show that the proposed method markedly outperforms the default SAM 2, achieving average Dice Similarity Coefficient improvement of 0.14 and 0.11 in the scenarios when 5 volumes and 1 volume are available for the initial adaptation, respectively. SLM-SAM 2 also exhibits stronger resistance to over-propagation, making a notable step toward more accurate automated annotation of medical images for segmentation model development.
Breast density in MRI: an AI-based quantification and relationship to assessment in mammography
Apr 21, 2025Abstract:Mammographic breast density is a well-established risk factor for breast cancer. Recently there has been interest in breast MRI as an adjunct to mammography, as this modality provides an orthogonal and highly quantitative assessment of breast tissue. However, its 3D nature poses analytic challenges related to delineating and aggregating complex structures across slices. Here, we applied an in-house machine-learning algorithm to assess breast density on normal breasts in three MRI datasets. Breast density was consistent across different datasets (0.104 - 0.114). Analysis across different age groups also demonstrated strong consistency across datasets and confirmed a trend of decreasing density with age as reported in previous studies. MR breast density was correlated with mammographic breast density, although some notable differences suggest that certain breast density components are captured only on MRI. Future work will determine how to integrate MR breast density with current tools to improve future breast cancer risk prediction.
RaD: A Metric for Medical Image Distribution Comparison in Out-of-Domain Detection and Other Applications
Dec 02, 2024Abstract:Determining whether two sets of images belong to the same or different domain is a crucial task in modern medical image analysis and deep learning, where domain shift is a common problem that commonly results in decreased model performance. This determination is also important to evaluate the output quality of generative models, e.g., image-to-image translation models used to mitigate domain shift. Current metrics for this either rely on the (potentially biased) choice of some downstream task such as segmentation, or adopt task-independent perceptual metrics (e.g., FID) from natural imaging which insufficiently capture anatomical consistency and realism in medical images. We introduce a new perceptual metric tailored for medical images: Radiomic Feature Distance (RaD), which utilizes standardized, clinically meaningful and interpretable image features. We show that RaD is superior to other metrics for out-of-domain (OOD) detection in a variety of experiments. Furthermore, RaD outperforms previous perceptual metrics (FID, KID, etc.) for image-to-image translation by correlating more strongly with downstream task performance as well as anatomical consistency and realism, and shows similar utility for evaluating unconditional image generation. RaD also offers additional benefits such as interpretability, as well as stability and computational efficiency at low sample sizes. Our results are supported by broad experiments spanning four multi-domain medical image datasets, nine downstream tasks, six image translation models, and other factors, highlighting the broad potential of RaD for medical image analysis.
Segment anything model 2: an application to 2D and 3D medical images
Aug 06, 2024Abstract:Segment Anything Model (SAM) has gained significant attention because of its ability to segment varous objects in images given a prompt. The recently developed SAM 2 has extended this ability to video inputs. This opens an opportunity to apply SAM to 3D images, one of the fundamental tasks in the medical imaging field. In this paper, we extensively evaluate SAM 2's ability to segment both 2D and 3D medical images by first collecting 18 medical imaging datasets, including common 3D modalities such as computed tomography (CT), magnetic resonance imaging (MRI), and positron emission tomography (PET) as well as 2D modalities such as X-ray and ultrasound. Two evaluation pipelines of SAM 2 are considered: (1) multi-frame 3D segmentation, where prompts are provided to one or multiple slice(s) selected from the volume, and (2) single-frame 2D segmentation, where prompts are provided to each slice. The former is only applicable to 3D modalities, while the latter applies to both 2D and 3D modalities. Our results show that SAM 2 exhibits similar performance as SAM under single-frame 2D segmentation, and has variable performance under multi-frame 3D segmentation depending on the choices of slices to annotate, the direction of the propagation, the predictions utilized during the propagation, etc.
Realtime Robust Shape Estimation of Deformable Linear Object
Mar 24, 2024



Abstract:Realtime shape estimation of continuum objects and manipulators is essential for developing accurate planning and control paradigms. The existing methods that create dense point clouds from camera images, and/or use distinguishable markers on a deformable body have limitations in realtime tracking of large continuum objects/manipulators. The physical occlusion of markers can often compromise accurate shape estimation. We propose a robust method to estimate the shape of linear deformable objects in realtime using scattered and unordered key points. By utilizing a robust probability-based labeling algorithm, our approach identifies the true order of the detected key points and then reconstructs the shape using piecewise spline interpolation. The approach only relies on knowing the number of the key points and the interval between two neighboring points. We demonstrate the robustness of the method when key points are partially occluded. The proposed method is also integrated into a simulation in Unity for tracking the shape of a cable with a length of 1m and a radius of 5mm. The simulation results show that our proposed approach achieves an average length error of 1.07% over the continuum's centerline and an average cross-section error of 2.11mm. The real-world experiments of tracking and estimating a heavy-load cable prove that the proposed approach is robust under occlusion and complex entanglement scenarios.
ContourDiff: Unpaired Image Translation with Contour-Guided Diffusion Models
Mar 16, 2024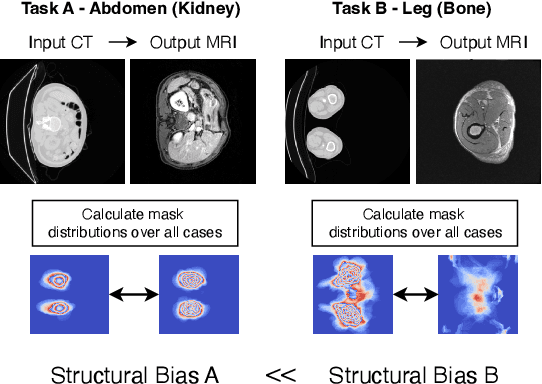
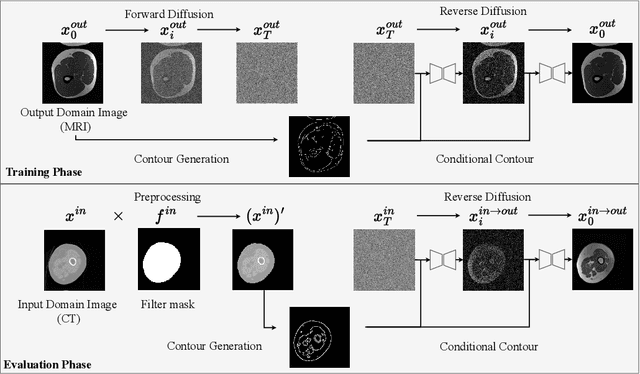
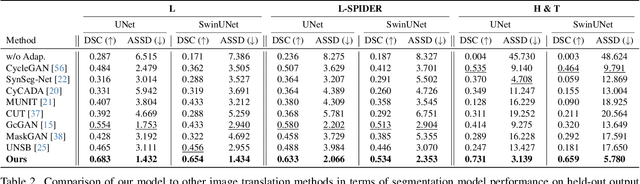
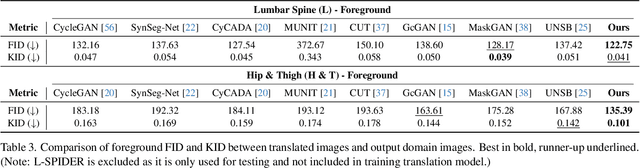
Abstract:Accurately translating medical images across different modalities (e.g., CT to MRI) has numerous downstream clinical and machine learning applications. While several methods have been proposed to achieve this, they often prioritize perceptual quality with respect to output domain features over preserving anatomical fidelity. However, maintaining anatomy during translation is essential for many tasks, e.g., when leveraging masks from the input domain to develop a segmentation model with images translated to the output domain. To address these challenges, we propose ContourDiff, a novel framework that leverages domain-invariant anatomical contour representations of images. These representations are simple to extract from images, yet form precise spatial constraints on their anatomical content. We introduce a diffusion model that converts contour representations of images from arbitrary input domains into images in the output domain of interest. By applying the contour as a constraint at every diffusion sampling step, we ensure the preservation of anatomical content. We evaluate our method by training a segmentation model on images translated from CT to MRI with their original CT masks and testing its performance on real MRIs. Our method outperforms other unpaired image translation methods by a significant margin, furthermore without the need to access any input domain information during training.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge