Roy Colglazier
Improving Surgical Risk Prediction Through Integrating Automated Body Composition Analysis: a Retrospective Trial on Colectomy Surgery
Jun 16, 2025Abstract:Objective: To evaluate whether preoperative body composition metrics automatically extracted from CT scans can predict postoperative outcomes after colectomy, either alone or combined with clinical variables or existing risk predictors. Main outcomes and measures: The primary outcome was the predictive performance for 1-year all-cause mortality following colectomy. A Cox proportional hazards model with 1-year follow-up was used, and performance was evaluated using the concordance index (C-index) and Integrated Brier Score (IBS). Secondary outcomes included postoperative complications, unplanned readmission, blood transfusion, and severe infection, assessed using AUC and Brier Score from logistic regression. Odds ratios (OR) described associations between individual CT-derived body composition metrics and outcomes. Over 300 features were extracted from preoperative CTs across multiple vertebral levels, including skeletal muscle area, density, fat areas, and inter-tissue metrics. NSQIP scores were available for all surgeries after 2012.
Deep learning automates Cobb angle measurement compared with multi-expert observers
Mar 18, 2024Abstract:Scoliosis, a prevalent condition characterized by abnormal spinal curvature leading to deformity, requires precise assessment methods for effective diagnosis and management. The Cobb angle is a widely used scoliosis quantification method that measures the degree of curvature between the tilted vertebrae. Yet, manual measuring of Cobb angles is time-consuming and labor-intensive, fraught with significant interobserver and intraobserver variability. To address these challenges and the lack of interpretability found in certain existing automated methods, we have created fully automated software that not only precisely measures the Cobb angle but also provides clear visualizations of these measurements. This software integrates deep neural network-based spine region detection and segmentation, spine centerline identification, pinpointing the most significantly tilted vertebrae, and direct visualization of Cobb angles on the original images. Upon comparison with the assessments of 7 expert readers, our algorithm exhibited a mean deviation in Cobb angle measurements of 4.17 degrees, notably surpassing the manual approach's average intra-reader discrepancy of 5.16 degrees. The algorithm also achieved intra-class correlation coefficients (ICC) exceeding 0.96 and Pearson correlation coefficients above 0.944, reflecting robust agreement with expert assessments and superior measurement reliability. Through the comprehensive reader study and statistical analysis, we believe this algorithm not only ensures a higher consensus with expert readers but also enhances interpretability and reproducibility during assessments. It holds significant promise for clinical application, potentially aiding physicians in more accurate scoliosis assessment and diagnosis, thereby improving patient care.
SegmentAnyBone: A Universal Model that Segments Any Bone at Any Location on MRI
Jan 23, 2024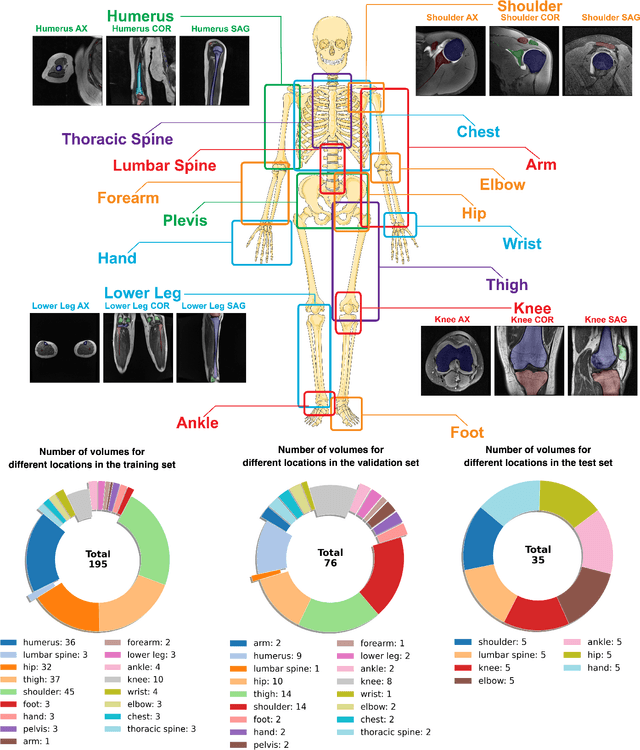
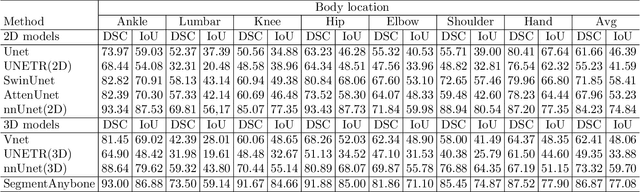
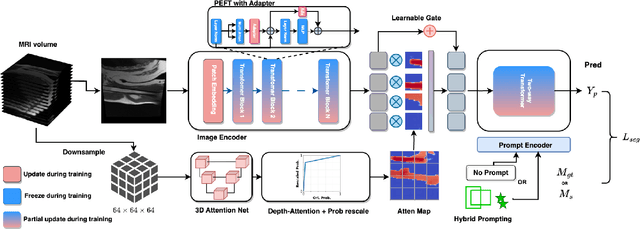

Abstract:Magnetic Resonance Imaging (MRI) is pivotal in radiology, offering non-invasive and high-quality insights into the human body. Precise segmentation of MRIs into different organs and tissues would be highly beneficial since it would allow for a higher level of understanding of the image content and enable important measurements, which are essential for accurate diagnosis and effective treatment planning. Specifically, segmenting bones in MRI would allow for more quantitative assessments of musculoskeletal conditions, while such assessments are largely absent in current radiological practice. The difficulty of bone MRI segmentation is illustrated by the fact that limited algorithms are publicly available for use, and those contained in the literature typically address a specific anatomic area. In our study, we propose a versatile, publicly available deep-learning model for bone segmentation in MRI across multiple standard MRI locations. The proposed model can operate in two modes: fully automated segmentation and prompt-based segmentation. Our contributions include (1) collecting and annotating a new MRI dataset across various MRI protocols, encompassing over 300 annotated volumes and 8485 annotated slices across diverse anatomic regions; (2) investigating several standard network architectures and strategies for automated segmentation; (3) introducing SegmentAnyBone, an innovative foundational model-based approach that extends Segment Anything Model (SAM); (4) comparative analysis of our algorithm and previous approaches; and (5) generalization analysis of our algorithm across different anatomical locations and MRI sequences, as well as an external dataset. We publicly release our model at https://github.com/mazurowski-lab/SegmentAnyBone.
Improving Image Classification of Knee Radiographs: An Automated Image Labeling Approach
Sep 06, 2023Abstract:Large numbers of radiographic images are available in knee radiology practices which could be used for training of deep learning models for diagnosis of knee abnormalities. However, those images do not typically contain readily available labels due to limitations of human annotations. The purpose of our study was to develop an automated labeling approach that improves the image classification model to distinguish normal knee images from those with abnormalities or prior arthroplasty. The automated labeler was trained on a small set of labeled data to automatically label a much larger set of unlabeled data, further improving the image classification performance for knee radiographic diagnosis. We developed our approach using 7,382 patients and validated it on a separate set of 637 patients. The final image classification model, trained using both manually labeled and pseudo-labeled data, had the higher weighted average AUC (WAUC: 0.903) value and higher AUC-ROC values among all classes (normal AUC-ROC: 0.894; abnormal AUC-ROC: 0.896, arthroplasty AUC-ROC: 0.990) compared to the baseline model (WAUC=0.857; normal AUC-ROC: 0.842; abnormal AUC-ROC: 0.848, arthroplasty AUC-ROC: 0.987), trained using only manually labeled data. DeLong tests show that the improvement is significant on normal (p-value<0.002) and abnormal (p-value<0.001) images. Our findings demonstrated that the proposed automated labeling approach significantly improves the performance of image classification for radiographic knee diagnosis, allowing for facilitating patient care and curation of large knee datasets.
SuperMask: Generating High-resolution object masks from multi-view, unaligned low-resolution MRIs
Mar 13, 2023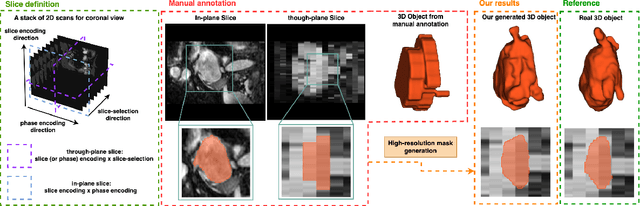

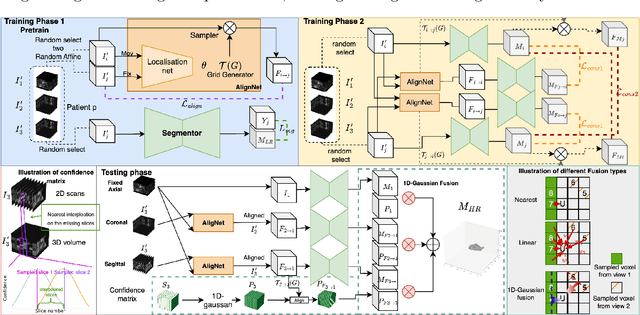
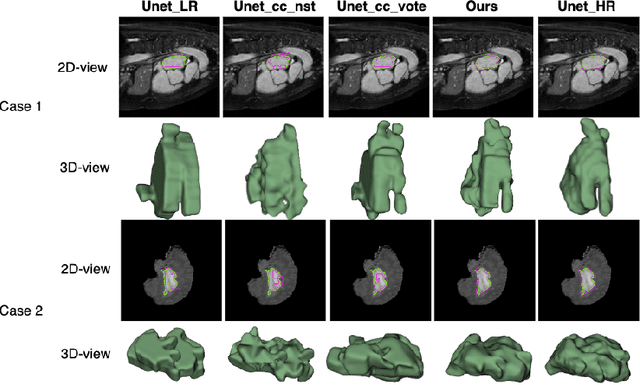
Abstract:Three-dimensional segmentation in magnetic resonance images (MRI), which reflects the true shape of the objects, is challenging since high-resolution isotropic MRIs are rare and typical MRIs are anisotropic, with the out-of-plane dimension having a much lower resolution. A potential remedy to this issue lies in the fact that often multiple sequences are acquired on different planes. However, in practice, these sequences are not orthogonal to each other, limiting the applicability of many previous solutions to reconstruct higher-resolution images from multiple lower-resolution ones. We propose a weakly-supervised deep learning-based solution to generating high-resolution masks from multiple low-resolution images. Our method combines segmentation and unsupervised registration networks by introducing two new regularizations to make registration and segmentation reinforce each other. Finally, we introduce a multi-view fusion method to generate high-resolution target object masks. The experimental results on two datasets show the superiority of our methods. Importantly, the advantage of not using high-resolution images in the training process makes our method applicable to a wide variety of MRI segmentation tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge