Jianying Hu
BMFM-RNA: An Open Framework for Building and Evaluating Transcriptomic Foundation Models
Jun 17, 2025



Abstract:Transcriptomic foundation models (TFMs) have recently emerged as powerful tools for analyzing gene expression in cells and tissues, supporting key tasks such as cell-type annotation, batch correction, and perturbation prediction. However, the diversity of model implementations and training strategies across recent TFMs, though promising, makes it challenging to isolate the contribution of individual design choices or evaluate their potential synergies. This hinders the field's ability to converge on best practices and limits the reproducibility of insights across studies. We present BMFM-RNA, an open-source, modular software package that unifies diverse TFM pretraining and fine-tuning objectives within a single framework. Leveraging this capability, we introduce a novel training objective, whole cell expression decoder (WCED), which captures global expression patterns using an autoencoder-like CLS bottleneck representation. In this paper, we describe the framework, supported input representations, and training objectives. We evaluated four model checkpoints pretrained on CELLxGENE using combinations of masked language modeling (MLM), WCED and multitask learning. Using the benchmarking capabilities of BMFM-RNA, we show that WCED-based models achieve performance that matches or exceeds state-of-the-art approaches like scGPT across more than a dozen datasets in both zero-shot and fine-tuning tasks. BMFM-RNA, available as part of the biomed-multi-omics project ( https://github.com/BiomedSciAI/biomed-multi-omic ), offers a reproducible foundation for systematic benchmarking and community-driven exploration of optimal TFM training strategies, enabling the development of more effective tools to leverage the latest advances in AI for understanding cell biology.
Multi-view biomedical foundation models for molecule-target and property prediction
Oct 25, 2024


Abstract:Foundation models applied to bio-molecular space hold promise to accelerate drug discovery. Molecular representation is key to building such models. Previous works have typically focused on a single representation or view of the molecules. Here, we develop a multi-view foundation model approach, that integrates molecular views of graph, image and text. Single-view foundation models are each pre-trained on a dataset of up to 200M molecules and then aggregated into combined representations. Our multi-view model is validated on a diverse set of 18 tasks, encompassing ligand-protein binding, molecular solubility, metabolism and toxicity. We show that the multi-view models perform robustly and are able to balance the strengths and weaknesses of specific views. We then apply this model to screen compounds against a large (>100 targets) set of G Protein-Coupled receptors (GPCRs). From this library of targets, we identify 33 that are related to Alzheimer's disease. On this subset, we employ our model to identify strong binders, which are validated through structure-based modeling and identification of key binding motifs.
An AI-Guided Data Centric Strategy to Detect and Mitigate Biases in Healthcare Datasets
Nov 06, 2023Abstract:The adoption of diagnosis and prognostic algorithms in healthcare has led to concerns about the perpetuation of bias against disadvantaged groups of individuals. Deep learning methods to detect and mitigate bias have revolved around modifying models, optimization strategies, and threshold calibration with varying levels of success. Here, we generate a data-centric, model-agnostic, task-agnostic approach to evaluate dataset bias by investigating the relationship between how easily different groups are learned at small sample sizes (AEquity). We then apply a systematic analysis of AEq values across subpopulations to identify and mitigate manifestations of racial bias in two known cases in healthcare - Chest X-rays diagnosis with deep convolutional neural networks and healthcare utilization prediction with multivariate logistic regression. AEq is a novel and broadly applicable metric that can be applied to advance equity by diagnosing and remediating bias in healthcare datasets.
Disease Progression Modeling Workbench 360
Jun 24, 2021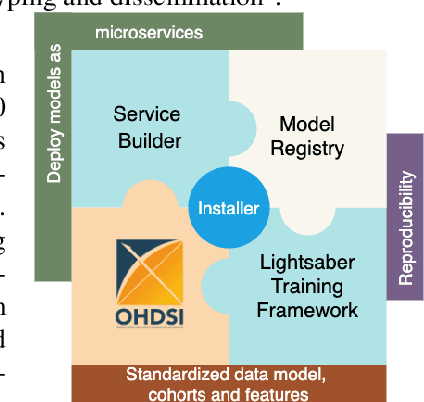
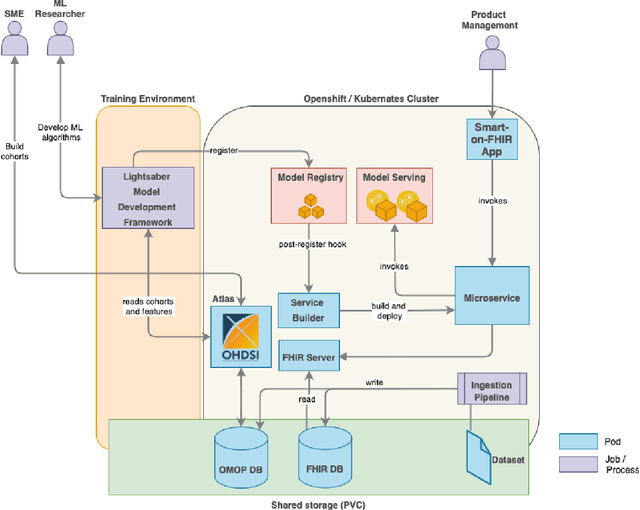
Abstract:In this work we introduce Disease Progression Modeling workbench 360 (DPM360) opensource clinical informatics framework for collaborative research and delivery of healthcare AI. DPM360, when fully developed, will manage the entire modeling life cycle, from data analysis (e.g., cohort identification) to machine learning algorithm development and prototyping. DPM360 augments the advantages of data model standardization and tooling (OMOP-CDM, Athena, ATLAS) provided by the widely-adopted OHDSI initiative with a powerful machine learning training framework, and a mechanism for rapid prototyping through automatic deployment of models as containerized services to a cloud environment.
Explicit-Blurred Memory Network for Analyzing Patient Electronic Health Records
Nov 15, 2019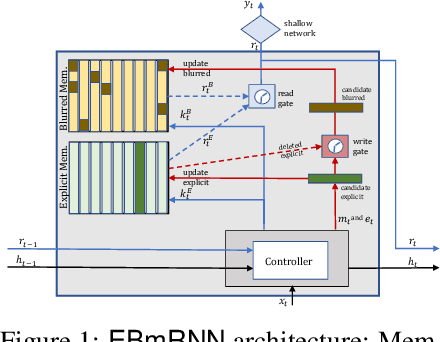
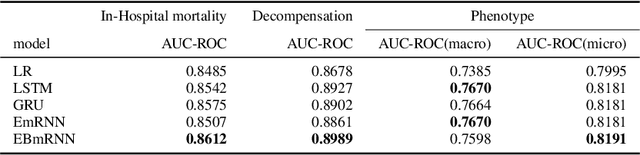
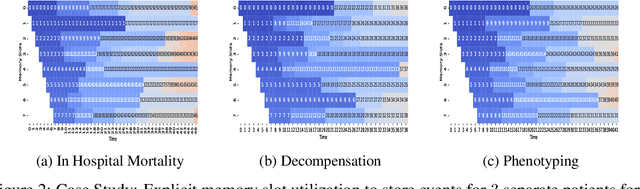
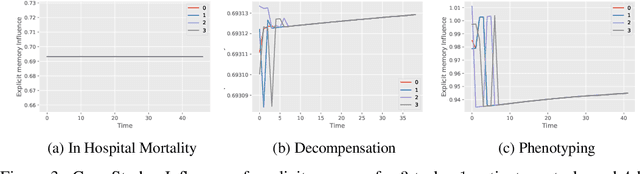
Abstract:In recent years, we have witnessed an increased interest in temporal modeling of patient records from large scale Electronic Health Records (EHR). While simpler RNN models have been used for such problems, memory networks, which in other domains were found to generalize well, are underutilized. Traditional memory networks involve diffused and non-linear operations where influence of past events on outputs are not readily quantifiable. We posit that this lack of interpretability makes such networks not applicable for EHR analysis. While networks with explicit memory have been proposed recently, the discontinuities imposed by the discrete operations make such networks harder to train and require more supervision. The problem is further exacerbated in the limited data setting of EHR studies. In this paper, we propose a novel memory architecture that is more interpretable than traditional memory networks while being easier to train than explicit memory banks. Inspired by well-known models of human cognition, we propose partitioning the external memory space into (a) a primary explicit memory block to store exact replicas of recent events to support interpretations, followed by (b) a secondary blurred memory block that accumulates salient aspects of past events dropped from the explicit block as higher level abstractions and allow training with less supervision by stabilize the gradients. We apply the model for 3 learning problems on ICU records from the MIMIC III database spanning millions of data points. Our model performs comparably to the state-of the art while also, crucially, enabling ready interpretation of the results.
Simultaneous Modeling of Multiple Complications for Risk Profiling in Diabetes Care
Feb 19, 2018
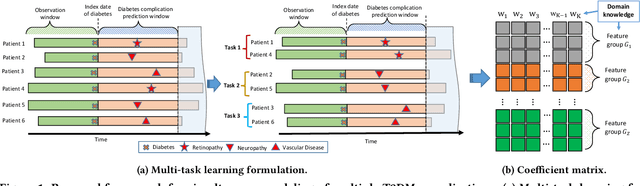
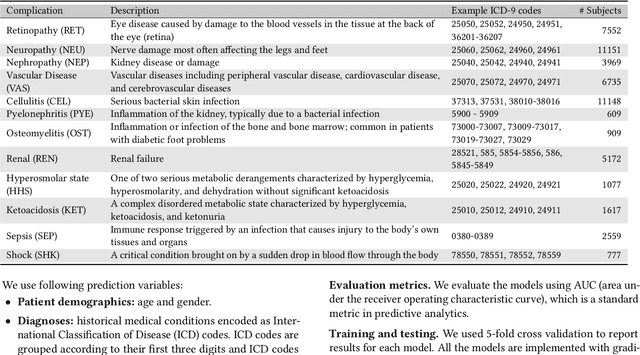
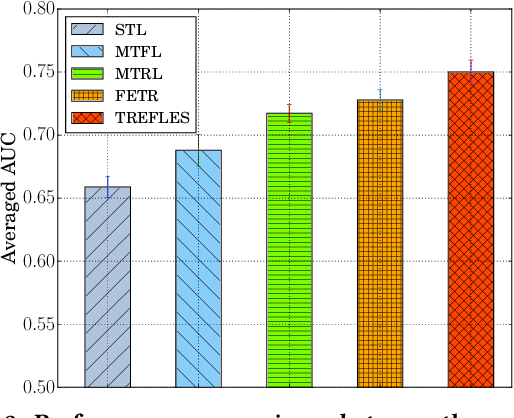
Abstract:Type 2 diabetes mellitus (T2DM) is a chronic disease that often results in multiple complications. Risk prediction and profiling of T2DM complications is critical for healthcare professionals to design personalized treatment plans for patients in diabetes care for improved outcomes. In this paper, we study the risk of developing complications after the initial T2DM diagnosis from longitudinal patient records. We propose a novel multi-task learning approach to simultaneously model multiple complications where each task corresponds to the risk modeling of one complication. Specifically, the proposed method strategically captures the relationships (1) between the risks of multiple T2DM complications, (2) between the different risk factors, and (3) between the risk factor selection patterns. The method uses coefficient shrinkage to identify an informative subset of risk factors from high-dimensional data, and uses a hierarchical Bayesian framework to allow domain knowledge to be incorporated as priors. The proposed method is favorable for healthcare applications because in additional to improved prediction performance, relationships among the different risks and risk factors are also identified. Extensive experimental results on a large electronic medical claims database show that the proposed method outperforms state-of-the-art models by a significant margin. Furthermore, we show that the risk associations learned and the risk factors identified lead to meaningful clinical insights.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge