Chunhao Wang
MCoT-MVS: Multi-level Vision Selection by Multi-modal Chain-of-Thought Reasoning for Composed Image Retrieval
Mar 18, 2026Abstract:Composed Image Retrieval (CIR) aims to retrieve target images based on a reference image and modified texts. However, existing methods often struggle to extract the correct semantic cues from the reference image that best reflect the user's intent under textual modification prompts, resulting in interference from irrelevant visual noise. In this paper, we propose a novel Multi-level Vision Selection by Multi-modal Chain-of-Thought Reasoning (MCoT-MVS) for CIR, integrating attention-aware multi-level vision features guided by reasoning cues from a multi-modal large language model (MLLM). Specifically, we leverage an MLLM to perform chain-of-thought reasoning on the multimodal composed input, generating the retained, removed, and target-inferred texts. These textual cues subsequently guide two reference visual attention selection modules to selectively extract discriminative patch-level and instance-level semantics from the reference image. Finally, to effectively fuse these multi-granular visual cues with the modified text and the imagined target description, we design a weighted hierarchical combination module to align the composed query with target images in a unified embedding space. Extensive experiments on two CIR benchmarks, namely CIRR and FashionIQ, demonstrate that our approach consistently outperforms existing methods and achieves new state-of-the-art performance. Code and trained models are publicly released.
Analysis of the MICCAI Brain Tumor Segmentation -- Metastases (BraTS-METS) 2025 Lighthouse Challenge: Brain Metastasis Segmentation on Pre- and Post-treatment MRI
Apr 16, 2025Abstract:Despite continuous advancements in cancer treatment, brain metastatic disease remains a significant complication of primary cancer and is associated with an unfavorable prognosis. One approach for improving diagnosis, management, and outcomes is to implement algorithms based on artificial intelligence for the automated segmentation of both pre- and post-treatment MRI brain images. Such algorithms rely on volumetric criteria for lesion identification and treatment response assessment, which are still not available in clinical practice. Therefore, it is critical to establish tools for rapid volumetric segmentations methods that can be translated to clinical practice and that are trained on high quality annotated data. The BraTS-METS 2025 Lighthouse Challenge aims to address this critical need by establishing inter-rater and intra-rater variability in dataset annotation by generating high quality annotated datasets from four individual instances of segmentation by neuroradiologists while being recorded on video (two instances doing "from scratch" and two instances after AI pre-segmentation). This high-quality annotated dataset will be used for testing phase in 2025 Lighthouse challenge and will be publicly released at the completion of the challenge. The 2025 Lighthouse challenge will also release the 2023 and 2024 segmented datasets that were annotated using an established pipeline of pre-segmentation, student annotation, two neuroradiologists checking, and one neuroradiologist finalizing the process. It builds upon its previous edition by including post-treatment cases in the dataset. Using these high-quality annotated datasets, the 2025 Lighthouse challenge plans to test benchmark algorithms for automated segmentation of pre-and post-treatment brain metastases (BM), trained on diverse and multi-institutional datasets of MRI images obtained from patients with brain metastases.
Embedding Radiomics into Vision Transformers for Multimodal Medical Image Classification
Apr 15, 2025Abstract:Background: Deep learning has significantly advanced medical image analysis, with Vision Transformers (ViTs) offering a powerful alternative to convolutional models by modeling long-range dependencies through self-attention. However, ViTs are inherently data-intensive and lack domain-specific inductive biases, limiting their applicability in medical imaging. In contrast, radiomics provides interpretable, handcrafted descriptors of tissue heterogeneity but suffers from limited scalability and integration into end-to-end learning frameworks. In this work, we propose the Radiomics-Embedded Vision Transformer (RE-ViT) that combines radiomic features with data-driven visual embeddings within a ViT backbone. Purpose: To develop a hybrid RE-ViT framework that integrates radiomics and patch-wise ViT embeddings through early fusion, enhancing robustness and performance in medical image classification. Methods: Following the standard ViT pipeline, images were divided into patches. For each patch, handcrafted radiomic features were extracted and fused with linearly projected pixel embeddings. The fused representations were normalized, positionally encoded, and passed to the ViT encoder. A learnable [CLS] token aggregated patch-level information for classification. We evaluated RE-ViT on three public datasets (including BUSI, ChestXray2017, and Retinal OCT) using accuracy, macro AUC, sensitivity, and specificity. RE-ViT was benchmarked against CNN-based (VGG-16, ResNet) and hybrid (TransMed) models. Results: RE-ViT achieved state-of-the-art results: on BUSI, AUC=0.950+/-0.011; on ChestXray2017, AUC=0.989+/-0.004; on Retinal OCT, AUC=0.986+/-0.001, which outperforms other comparison models. Conclusions: The RE-ViT framework effectively integrates radiomics with ViT architectures, demonstrating improved performance and generalizability across multimodal medical image classification tasks.
Quantum Speedups for Markov Chain Monte Carlo Methods with Application to Optimization
Apr 04, 2025Abstract:We propose quantum algorithms that provide provable speedups for Markov Chain Monte Carlo (MCMC) methods commonly used for sampling from probability distributions of the form $\pi \propto e^{-f}$, where $f$ is a potential function. Our first approach considers Gibbs sampling for finite-sum potentials in the stochastic setting, employing an oracle that provides gradients of individual functions. In the second setting, we consider access only to a stochastic evaluation oracle, allowing simultaneous queries at two points of the potential function under the same stochastic parameter. By introducing novel techniques for stochastic gradient estimation, our algorithms improve the gradient and evaluation complexities of classical samplers, such as Hamiltonian Monte Carlo (HMC) and Langevin Monte Carlo (LMC) in terms of dimension, precision, and other problem-dependent parameters. Furthermore, we achieve quantum speedups in optimization, particularly for minimizing non-smooth and approximately convex functions that commonly appear in empirical risk minimization problems.
An Explainable Neural Radiomic Sequence Model with Spatiotemporal Continuity for Quantifying 4DCT-based Pulmonary Ventilation
Mar 31, 2025Abstract:Accurate evaluation of regional lung ventilation is essential for the management and treatment of lung cancer patients, supporting assessments of pulmonary function, optimization of therapeutic strategies, and monitoring of treatment response. Currently, ventilation scintigraphy using nuclear medicine techniques is widely employed in clinical practice; however, it is often time-consuming, costly, and entails additional radiation exposure. In this study, we propose an explainable neural radiomic sequence model to identify regions of compromised pulmonary ventilation based on four-dimensional computed tomography (4DCT). A cohort of 45 lung cancer patients from the VAMPIRE dataset was analyzed. For each patient, lung volumes were segmented from 4DCT, and voxel-wise radiomic features (56-dimensional) were extracted across the respiratory cycle to capture local intensity and texture dynamics, forming temporal radiomic sequences. Ground truth ventilation defects were delineated voxel-wise using Galligas-PET and DTPA-SPECT. To identify compromised regions, we developed a temporal saliency-enhanced explainable long short-term memory (LSTM) network trained on the radiomic sequences. Temporal saliency maps were generated to highlight key features contributing to the model's predictions. The proposed model demonstrated robust performance, achieving average (range) Dice similarity coefficients of 0.78 (0.74-0.79) for 25 PET cases and 0.78 (0.74-0.82) for 20 SPECT cases. The temporal saliency map explained three key radiomic sequences in ventilation quantification: during lung exhalation, compromised pulmonary function region typically exhibits (1) an increasing trend of intensity and (2) a decreasing trend of homogeneity, in contrast to healthy lung tissue.
BraTS-PEDs: Results of the Multi-Consortium International Pediatric Brain Tumor Segmentation Challenge 2023
Jul 11, 2024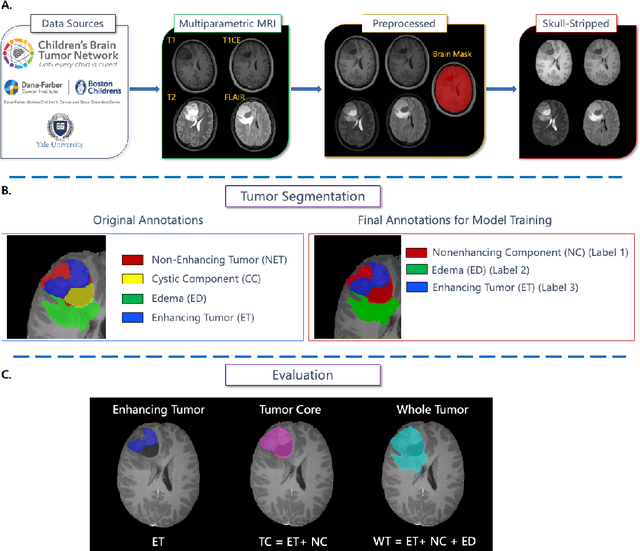

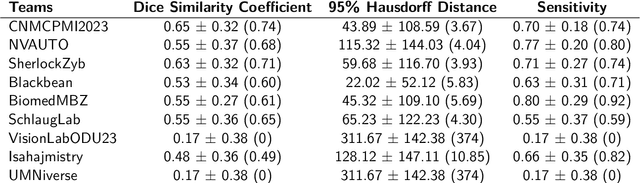
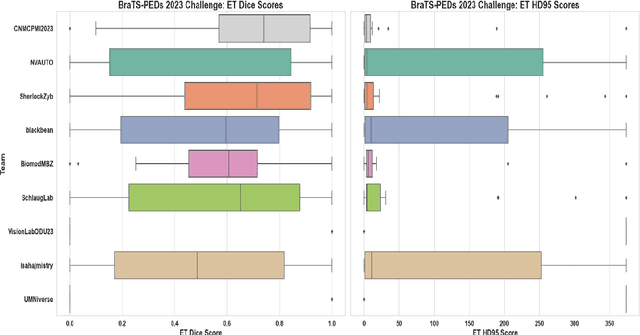
Abstract:Pediatric central nervous system tumors are the leading cause of cancer-related deaths in children. The five-year survival rate for high-grade glioma in children is less than 20%. The development of new treatments is dependent upon multi-institutional collaborative clinical trials requiring reproducible and accurate centralized response assessment. We present the results of the BraTS-PEDs 2023 challenge, the first Brain Tumor Segmentation (BraTS) challenge focused on pediatric brain tumors. This challenge utilized data acquired from multiple international consortia dedicated to pediatric neuro-oncology and clinical trials. BraTS-PEDs 2023 aimed to evaluate volumetric segmentation algorithms for pediatric brain gliomas from magnetic resonance imaging using standardized quantitative performance evaluation metrics employed across the BraTS 2023 challenges. The top-performing AI approaches for pediatric tumor analysis included ensembles of nnU-Net and Swin UNETR, Auto3DSeg, or nnU-Net with a self-supervised framework. The BraTSPEDs 2023 challenge fostered collaboration between clinicians (neuro-oncologists, neuroradiologists) and AI/imaging scientists, promoting faster data sharing and the development of automated volumetric analysis techniques. These advancements could significantly benefit clinical trials and improve the care of children with brain tumors.
Brain Tumor Segmentation (BraTS) Challenge 2024: Meningioma Radiotherapy Planning Automated Segmentation
May 28, 2024Abstract:The 2024 Brain Tumor Segmentation Meningioma Radiotherapy (BraTS-MEN-RT) challenge aims to advance automated segmentation algorithms using the largest known multi-institutional dataset of radiotherapy planning brain MRIs with expert-annotated target labels for patients with intact or post-operative meningioma that underwent either conventional external beam radiotherapy or stereotactic radiosurgery. Each case includes a defaced 3D post-contrast T1-weighted radiotherapy planning MRI in its native acquisition space, accompanied by a single-label "target volume" representing the gross tumor volume (GTV) and any at-risk post-operative site. Target volume annotations adhere to established radiotherapy planning protocols, ensuring consistency across cases and institutions. For pre-operative meningiomas, the target volume encompasses the entire GTV and associated nodular dural tail, while for post-operative cases, it includes at-risk resection cavity margins as determined by the treating institution. Case annotations were reviewed and approved by expert neuroradiologists and radiation oncologists. Participating teams will develop, containerize, and evaluate automated segmentation models using this comprehensive dataset. Model performance will be assessed using the lesion-wise Dice Similarity Coefficient and the 95% Hausdorff distance. The top-performing teams will be recognized at the Medical Image Computing and Computer Assisted Intervention Conference in October 2024. BraTS-MEN-RT is expected to significantly advance automated radiotherapy planning by enabling precise tumor segmentation and facilitating tailored treatment, ultimately improving patient outcomes.
Analysis of the BraTS 2023 Intracranial Meningioma Segmentation Challenge
May 16, 2024



Abstract:We describe the design and results from the BraTS 2023 Intracranial Meningioma Segmentation Challenge. The BraTS Meningioma Challenge differed from prior BraTS Glioma challenges in that it focused on meningiomas, which are typically benign extra-axial tumors with diverse radiologic and anatomical presentation and a propensity for multiplicity. Nine participating teams each developed deep-learning automated segmentation models using image data from the largest multi-institutional systematically expert annotated multilabel multi-sequence meningioma MRI dataset to date, which included 1000 training set cases, 141 validation set cases, and 283 hidden test set cases. Each case included T2, T2/FLAIR, T1, and T1Gd brain MRI sequences with associated tumor compartment labels delineating enhancing tumor, non-enhancing tumor, and surrounding non-enhancing T2/FLAIR hyperintensity. Participant automated segmentation models were evaluated and ranked based on a scoring system evaluating lesion-wise metrics including dice similarity coefficient (DSC) and 95% Hausdorff Distance. The top ranked team had a lesion-wise median dice similarity coefficient (DSC) of 0.976, 0.976, and 0.964 for enhancing tumor, tumor core, and whole tumor, respectively and a corresponding average DSC of 0.899, 0.904, and 0.871, respectively. These results serve as state-of-the-art benchmarks for future pre-operative meningioma automated segmentation algorithms. Additionally, we found that 1286 of 1424 cases (90.3%) had at least 1 compartment voxel abutting the edge of the skull-stripped image edge, which requires further investigation into optimal pre-processing face anonymization steps.
The Brain Tumor Segmentation in Pediatrics (BraTS-PEDs) Challenge: Focus on Pediatrics (CBTN-CONNECT-DIPGR-ASNR-MICCAI BraTS-PEDs)
Apr 29, 2024Abstract:Pediatric tumors of the central nervous system are the most common cause of cancer-related death in children. The five-year survival rate for high-grade gliomas in children is less than 20%. Due to their rarity, the diagnosis of these entities is often delayed, their treatment is mainly based on historic treatment concepts, and clinical trials require multi-institutional collaborations. Here we present the CBTN-CONNECT-DIPGR-ASNR-MICCAI BraTS-PEDs challenge, focused on pediatric brain tumors with data acquired across multiple international consortia dedicated to pediatric neuro-oncology and clinical trials. The CBTN-CONNECT-DIPGR-ASNR-MICCAI BraTS-PEDs challenge brings together clinicians and AI/imaging scientists to lead to faster development of automated segmentation techniques that could benefit clinical trials, and ultimately the care of children with brain tumors.
Stochastic Quantum Sampling for Non-Logconcave Distributions and Estimating Partition Functions
Oct 17, 2023Abstract:We present quantum algorithms for sampling from non-logconcave probability distributions in the form of $\pi(x) \propto \exp(-\beta f(x))$. Here, $f$ can be written as a finite sum $f(x):= \frac{1}{N}\sum_{k=1}^N f_k(x)$. Our approach is based on quantum simulated annealing on slowly varying Markov chains derived from unadjusted Langevin algorithms, removing the necessity for function evaluations which can be computationally expensive for large data sets in mixture modeling and multi-stable systems. We also incorporate a stochastic gradient oracle that implements the quantum walk operators inexactly by only using mini-batch gradients. As a result, our stochastic gradient based algorithm only accesses small subsets of data points in implementing the quantum walk. One challenge of quantizing the resulting Markov chains is that they do not satisfy the detailed balance condition in general. Consequently, the mixing time of the algorithm cannot be expressed in terms of the spectral gap of the transition density, making the quantum algorithms nontrivial to analyze. To overcome these challenges, we first build a hypothetical Markov chain that is reversible, and also converges to the target distribution. Then, we quantified the distance between our algorithm's output and the target distribution by using this hypothetical chain as a bridge to establish the total complexity. Our quantum algorithms exhibit polynomial speedups in terms of both dimension and precision dependencies when compared to the best-known classical algorithms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge