Annette Birkhold
Siemens Healthcare GmbH, Forchheim, Germany
ProPLIKS: Probablistic 3D human body pose estimation
Dec 05, 2024
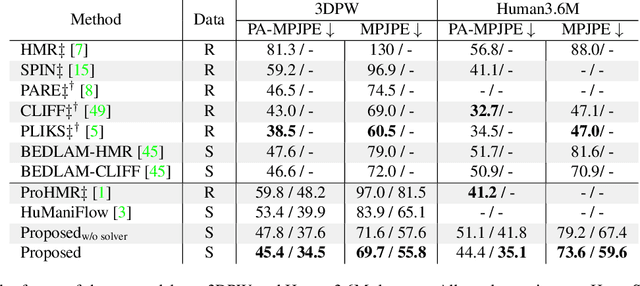

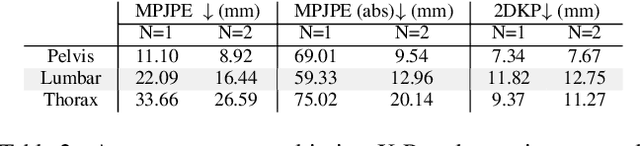
Abstract:We present a novel approach for 3D human pose estimation by employing probabilistic modeling. This approach leverages the advantages of normalizing flows in non-Euclidean geometries to address uncertain poses. Specifically, our method employs normalizing flow tailored to the SO(3) rotational group, incorporating a coupling mechanism based on the M\"obius transformation. This enables the framework to accurately represent any distribution on SO(3), effectively addressing issues related to discontinuities. Additionally, we reinterpret the challenge of reconstructing 3D human figures from 2D pixel-aligned inputs as the task of mapping these inputs to a range of probable poses. This perspective acknowledges the intrinsic ambiguity of the task and facilitates a straightforward integration method for multi-view scenarios. The combination of these strategies showcases the effectiveness of probabilistic models in complex scenarios for human pose estimation techniques. Our approach notably surpasses existing methods in the field of pose estimation. We also validate our methodology on human pose estimation from RGB images as well as medical X-Ray datasets.
Physics-Informed Learning for Time-Resolved Angiographic Contrast Agent Concentration Reconstruction
Mar 04, 2024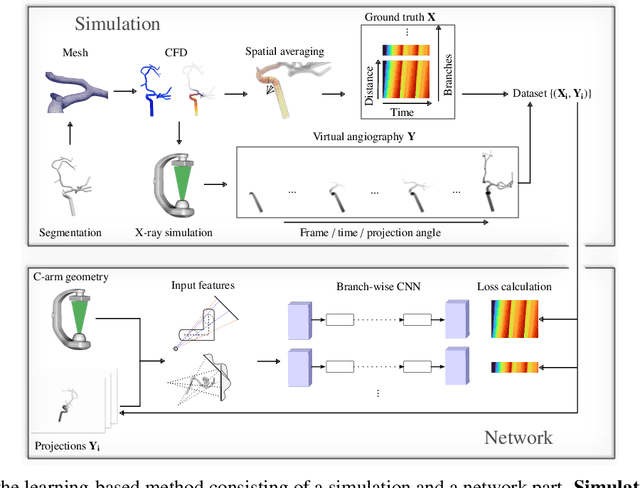
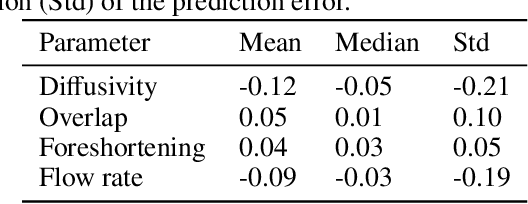
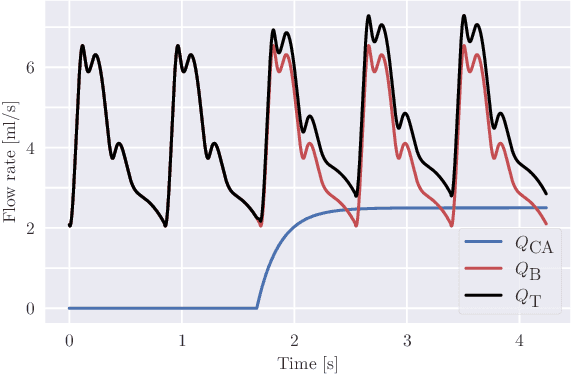
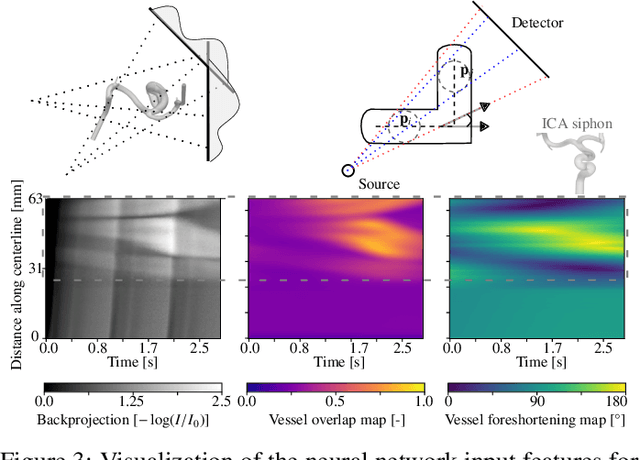
Abstract:Three-dimensional Digital Subtraction Angiography (3D-DSA) is a well-established X-ray-based technique for visualizing vascular anatomy. Recently, four-dimensional DSA (4D-DSA) reconstruction algorithms have been developed to enable the visualization of volumetric contrast flow dynamics through time-series of volumes. . This reconstruction problem is ill-posed mainly due to vessel overlap in the projection direction and geometric vessel foreshortening, which leads to information loss in the recorded projection images. However, knowledge about the underlying fluid dynamics can be leveraged to constrain the solution space. In our work, we implicitly include this information in a neural network-based model that is trained on a dataset of image-based blood flow simulations. The model predicts the spatially averaged contrast agent concentration for each centerline point of the vasculature over time, lowering the overall computational demand. The trained network enables the reconstruction of relative contrast agent concentrations with a mean absolute error of 0.02 $\pm$ 0.02 and a mean absolute percentage error of 5.31 % $\pm$ 9.25 %. Moreover, the network is robust to varying degrees of vessel overlap and vessel foreshortening. Our approach demonstrates the potential of the integration of machine learning and blood flow simulations in time-resolved angiographic flow reconstruction.
BOSS: Bones, Organs and Skin Shape Model
Mar 08, 2023Abstract:Objective: A digital twin of a patient can be a valuable tool for enhancing clinical tasks such as workflow automation, patient-specific X-ray dose optimization, markerless tracking, positioning, and navigation assistance in image-guided interventions. However, it is crucial that the patient's surface and internal organs are of high quality for any pose and shape estimates. At present, the majority of statistical shape models (SSMs) are restricted to a small number of organs or bones or do not adequately represent the general population. Method: To address this, we propose a deformable human shape and pose model that combines skin, internal organs, and bones, learned from CT images. By modeling the statistical variations in a pose-normalized space using probabilistic PCA while also preserving joint kinematics, our approach offers a holistic representation of the body that can benefit various medical applications. Results: We assessed our model's performance on a registered dataset, utilizing the unified shape space, and noted an average error of 3.6 mm for bones and 8.8 mm for organs. To further verify our findings, we conducted additional tests on publicly available datasets with multi-part segmentations, which confirmed the effectiveness of our model. Conclusion: This works shows that anatomically parameterized statistical shape models can be created accurately and in a computationally efficient manner. Significance: The proposed approach enables the construction of shape models that can be directly applied to various medical applications, including biomechanics and reconstruction.
Transient Hemodynamics Prediction Using an Efficient Octree-Based Deep Learning Model
Feb 13, 2023
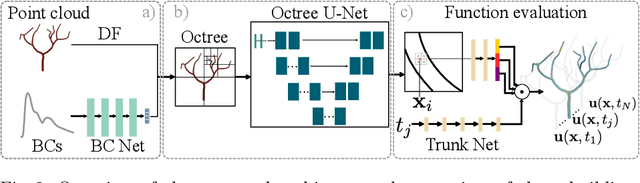
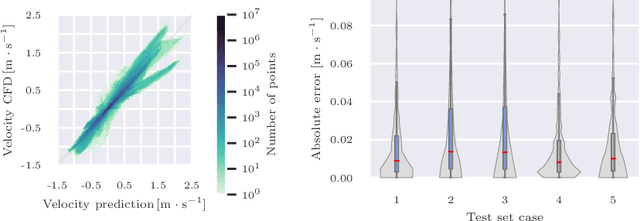
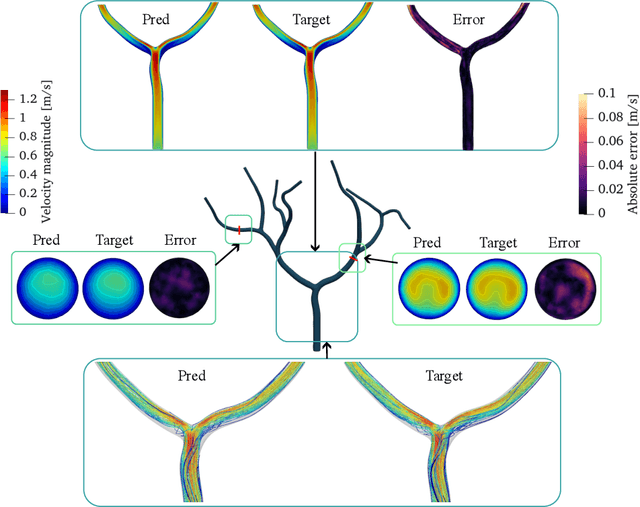
Abstract:Patient-specific hemodynamics assessment could support diagnosis and treatment of neurovascular diseases. Currently, conventional medical imaging modalities are not able to accurately acquire high-resolution hemodynamic information that would be required to assess complex neurovascular pathologies. Therefore, computational fluid dynamics (CFD) simulations can be applied to tomographic reconstructions to obtain clinically relevant information. However, three-dimensional (3D) CFD simulations require enormous computational resources and simulation-related expert knowledge that are usually not available in clinical environments. Recently, deep-learning-based methods have been proposed as CFD surrogates to improve computational efficiency. Nevertheless, the prediction of high-resolution transient CFD simulations for complex vascular geometries poses a challenge to conventional deep learning models. In this work, we present an architecture that is tailored to predict high-resolution (spatial and temporal) velocity fields for complex synthetic vascular geometries. For this, an octree-based spatial discretization is combined with an implicit neural function representation to efficiently handle the prediction of the 3D velocity field for each time step. The presented method is evaluated for the task of cerebral hemodynamics prediction before and during the injection of contrast agent in the internal carotid artery (ICA). Compared to CFD simulations, the velocity field can be estimated with a mean absolute error of 0.024 m/s, whereas the run time reduces from several hours on a high-performance cluster to a few seconds on a consumer graphical processing unit.
PLIKS: A Pseudo-Linear Inverse Kinematic Solver for 3D Human Body Estimation
Nov 21, 2022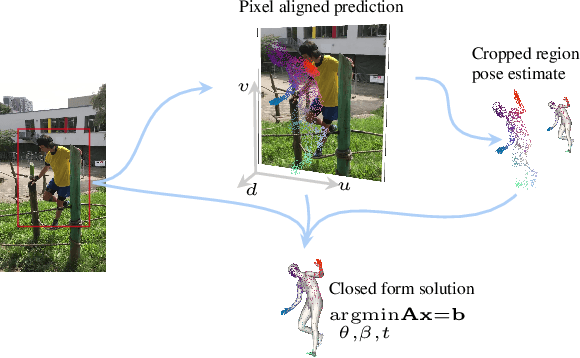
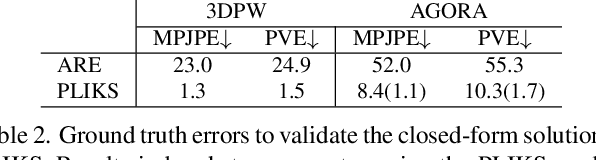
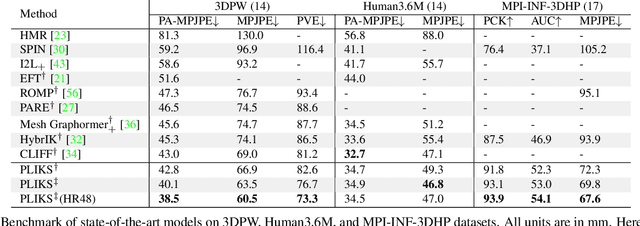
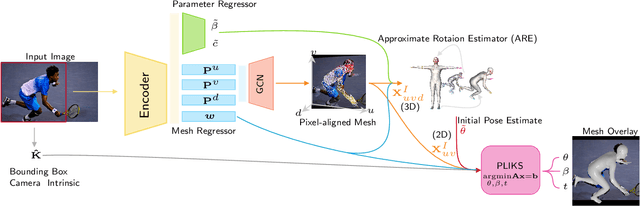

Abstract:We consider the problem of reconstructing a 3D mesh of the human body from a single 2D image as a model-in-the-loop optimization problem. Existing approaches often regress the shape, pose, and translation parameters of a parametric statistical model assuming a weak-perspective camera. In contrast, we first estimate 2D pixel-aligned vertices in image space and propose PLIKS (Pseudo-Linear Inverse Kinematic Solver) to regress the model parameters by minimizing a linear least squares problem. PLIKS is a linearized formulation of the parametric SMPL model, which provides an optimal pose and shape solution from an adequate initialization. Our method is based on analytically calculating an initial pose estimate from the network predicted 3D mesh followed by PLIKS to obtain an optimal solution for the given constraints. As our framework makes use of 2D pixel-aligned maps, it is inherently robust to partial occlusion. To demonstrate the performance of the proposed approach, we present quantitative evaluations which confirm that PLIKS achieves more accurate reconstruction with greater than 10% improvement compared to other state-of-the-art methods with respect to the standard 3D human pose and shape benchmarks while also obtaining a reconstruction error improvement of 12.9 mm on the newer AGORA dataset.
Deep Learning compatible Differentiable X-ray Projections for Inverse Rendering
Feb 04, 2021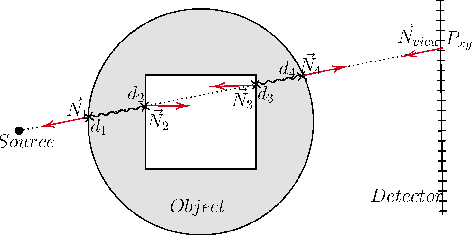
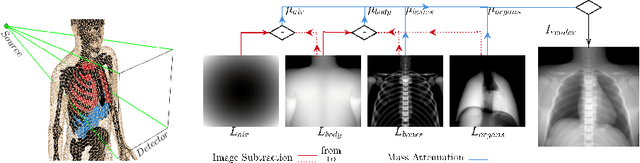

Abstract:Many minimally invasive interventional procedures still rely on 2D fluoroscopic imaging. Generating a patient-specific 3D model from these X-ray projection data would allow to improve the procedural workflow, e.g. by providing assistance functions such as automatic positioning. To accomplish this, two things are required. First, a statistical human shape model of the human anatomy and second, a differentiable X-ray renderer. In this work, we propose a differentiable renderer by deriving the distance travelled by a ray inside mesh structures to generate a distance map. To demonstrate its functioning, we use it for simulating X-ray images from human shape models. Then we show its application by solving the inverse problem, namely reconstructing 3D models from real 2D fluoroscopy images of the pelvis, which is an ideal anatomical structure for patient registration. This is accomplished by an iterative optimization strategy using gradient descent. With the majority of the pelvis being in the fluoroscopic field of view, we achieve a mean Hausdorff distance of 30 mm between the reconstructed model and the ground truth segmentation.
X-ray Scatter Estimation Using Deep Splines
Jan 22, 2021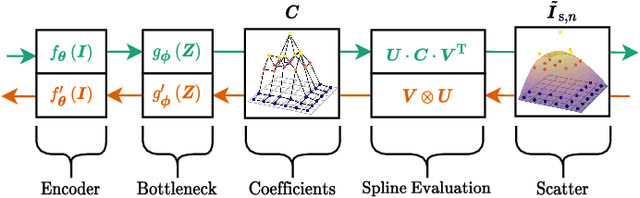
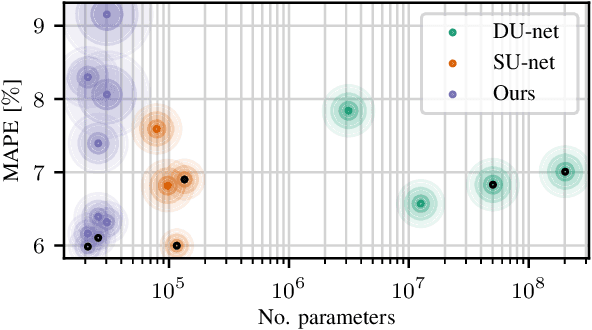
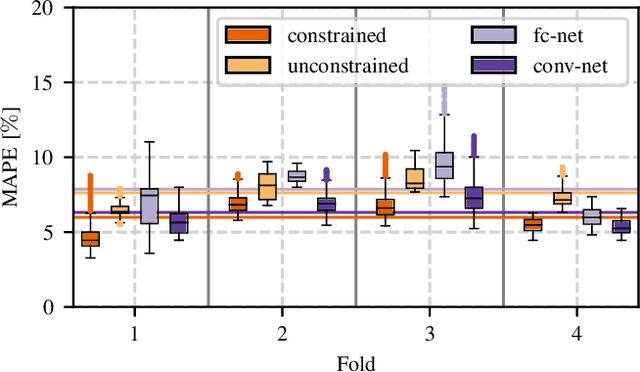
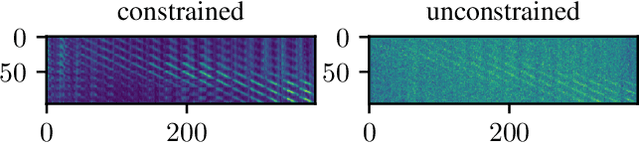
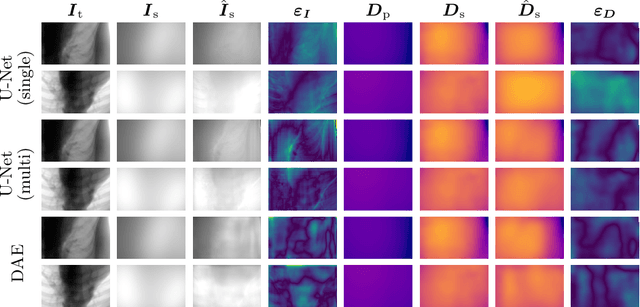
Abstract:Algorithmic X-ray scatter compensation is a desirable technique in flat-panel X-ray imaging and cone-beam computed tomography. State-of-the-art U-net based image translation approaches yielded promising results. As there are no physics constraints applied to the output of the U-Net, it cannot be ruled out that it yields spurious results. Unfortunately, those may be misleading in the context of medical imaging. To overcome this problem, we propose to embed B-splines as a known operator into neural networks. This inherently limits their predictions to well-behaved and smooth functions. In a study using synthetic head and thorax data as well as real thorax phantom data, we found that our approach performed on par with U-net when comparing both algorithms based on quantitative performance metrics. However, our approach not only reduces runtime and parameter complexity, but we also found it much more robust to unseen noise levels. While the U-net responded with visible artifacts, our approach preserved the X-ray signal's frequency characteristics.
Simultaneous Estimation of X-ray Back-Scatter and Forward-Scatter using Multi-Task Learning
Jul 08, 2020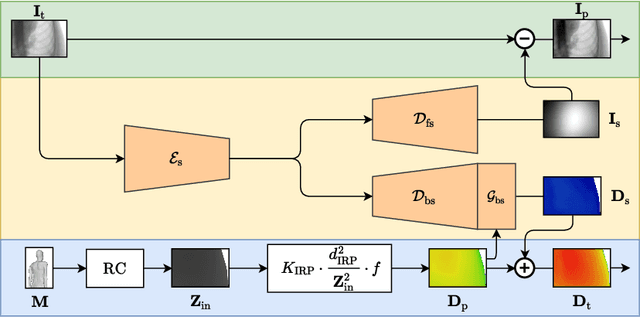
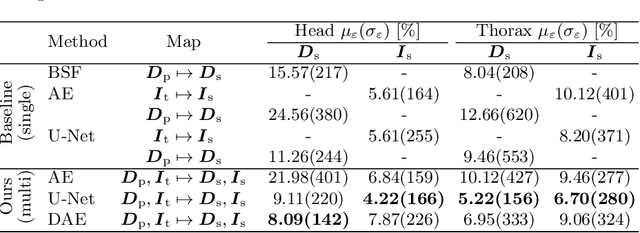
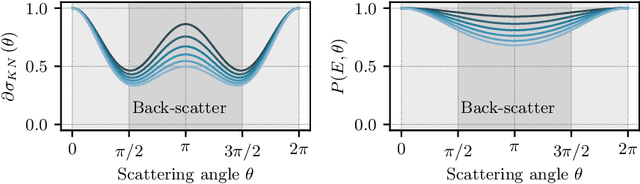

Abstract:Scattered radiation is a major concern impacting X-ray image-guided procedures in two ways. First, back-scatter significantly contributes to patient (skin) dose during complicated interventions. Second, forward-scattered radiation reduces contrast in projection images and introduces artifacts in 3-D reconstructions. While conventionally employed anti-scatter grids improve image quality by blocking X-rays, the additional attenuation due to the anti-scatter grid at the detector needs to be compensated for by a higher patient entrance dose. This also increases the room dose affecting the staff caring for the patient. For skin dose quantification, back-scatter is usually accounted for by applying pre-determined scalar back-scatter factors or linear point spread functions to a primary kerma forward projection onto a patient surface point. However, as patients come in different shapes, the generalization of conventional methods is limited. Here, we propose a novel approach combining conventional techniques with learning-based methods to simultaneously estimate the forward-scatter reaching the detector as well as the back-scatter affecting the patient skin dose. Knowing the forward-scatter, we can correct X-ray projections, while a good estimate of the back-scatter component facilitates an improved skin dose assessment. To simultaneously estimate forward-scatter as well as back-scatter, we propose a multi-task approach for joint back- and forward-scatter estimation by combining X-ray physics with neural networks. We show that, in theory, highly accurate scatter estimation in both cases is possible. In addition, we identify research directions for our multi-task framework and learning-based scatter estimation in general.
Action Learning for 3D Point Cloud Based Organ Segmentation
Jun 14, 2018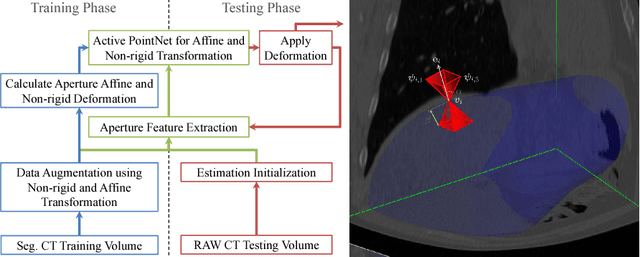
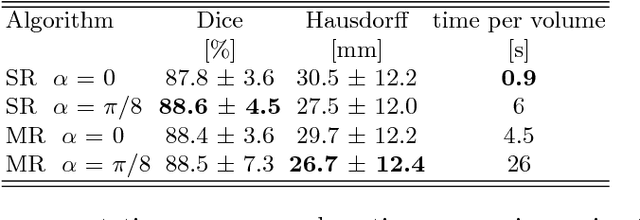

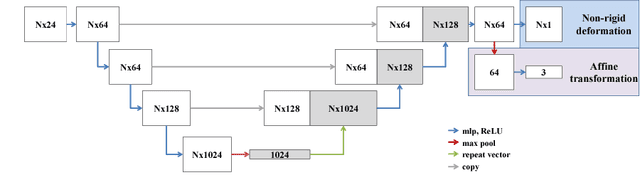
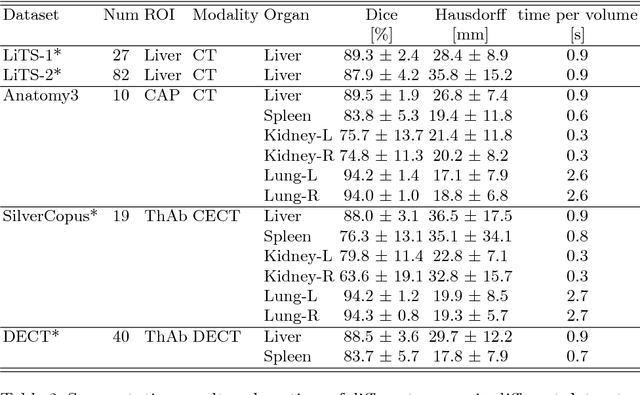
Abstract:We propose a novel point cloud based 3D organ segmentation pipeline utilizing deep Q-learning. In order to preserve shape properties, the learning process is guided using a statistical shape model. The trained agent directly predicts piece-wise linear transformations for all vertices in each iteration. This mapping between the ideal transformation for an object outline estimation is learned based on image features. To this end, we introduce aperture features that extract gray values by sampling the 3D volume within the cone centered around the associated vertex and its normal vector. Our approach is also capable of estimating a hierarchical pyramid of non rigid deformations for multi-resolution meshes. In the application phase, we use a marginal approach to gradually estimate affine as well as non-rigid transformations. We performed extensive evaluations to highlight the robust performance of our approach on a variety of challenge data as well as clinical data. Additionally, our method has a run time ranging from 0.3 to 2.7 seconds to segment each organ. In addition, we show that the proposed method can be applied to different organs, X-ray based modalities, and scanning protocols without the need of transfer learning. As we learn actions, even unseen reference meshes can be processed as demonstrated in an example with the Visible Human. From this we conclude that our method is robust, and we believe that our method can be successfully applied to many more applications, in particular, in the interventional imaging space.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge