Yuhong Wen
Empowering Federated Learning for Massive Models with NVIDIA FLARE
Feb 12, 2024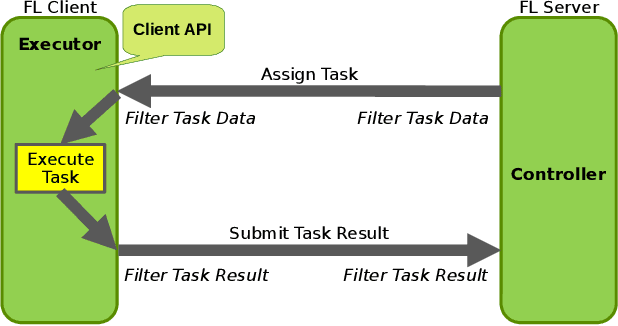

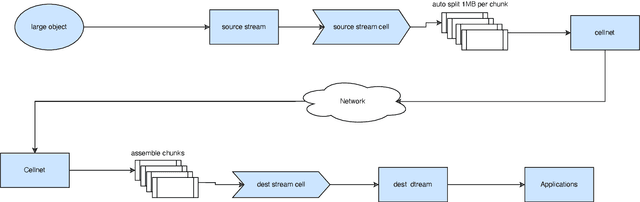
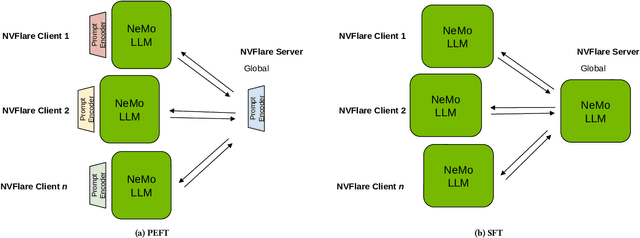
Abstract:In the ever-evolving landscape of artificial intelligence (AI) and large language models (LLMs), handling and leveraging data effectively has become a critical challenge. Most state-of-the-art machine learning algorithms are data-centric. However, as the lifeblood of model performance, necessary data cannot always be centralized due to various factors such as privacy, regulation, geopolitics, copyright issues, and the sheer effort required to move vast datasets. In this paper, we explore how federated learning enabled by NVIDIA FLARE can address these challenges with easy and scalable integration capabilities, enabling parameter-efficient and full supervised fine-tuning of LLMs for natural language processing and biopharmaceutical applications to enhance their accuracy and robustness.
NVIDIA FLARE: Federated Learning from Simulation to Real-World
Oct 24, 2022Abstract:Federated learning (FL) enables building robust and generalizable AI models by leveraging diverse datasets from multiple collaborators without centralizing the data. We created NVIDIA FLARE as an open-source software development kit (SDK) to make it easier for data scientists to use FL in their research and real-world applications. The SDK includes solutions for state-of-the-art FL algorithms and federated machine learning approaches, which facilitate building workflows for distributed learning across enterprises and enable platform developers to create a secure, privacy-preserving offering for multiparty collaboration utilizing homomorphic encryption or differential privacy. The SDK is a lightweight, flexible, and scalable Python package, and allows researchers to bring their data science workflows implemented in any training libraries (PyTorch, TensorFlow, XGBoost, or even NumPy) and apply them in real-world FL settings. This paper introduces the key design principles of FLARE and illustrates some use cases (e.g., COVID analysis) with customizable FL workflows that implement different privacy-preserving algorithms. Code is available at https://github.com/NVIDIA/NVFlare.
Federated Learning for Breast Density Classification: A Real-World Implementation
Sep 17, 2020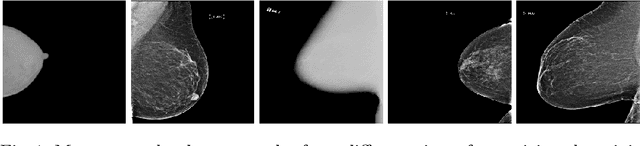
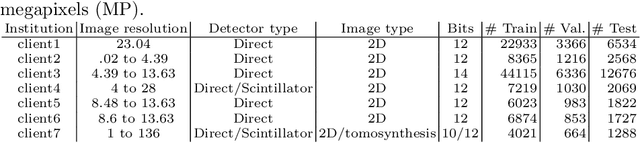

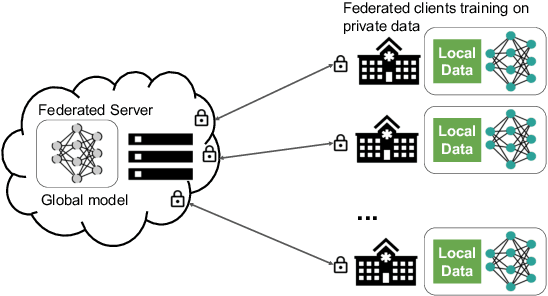
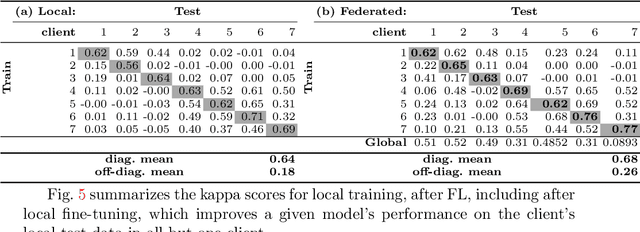
Abstract:Building robust deep learning-based models requires large quantities of diverse training data. In this study, we investigate the use of federated learning (FL) to build medical imaging classification models in a real-world collaborative setting. Seven clinical institutions from across the world joined this FL effort to train a model for breast density classification based on Breast Imaging, Reporting & Data System (BI-RADS). We show that despite substantial differences among the datasets from all sites (mammography system, class distribution, and data set size) and without centralizing data, we can successfully train AI models in federation. The results show that models trained using FL perform 6.3% on average better than their counterparts trained on an institute's local data alone. Furthermore, we show a 45.8% relative improvement in the models' generalizability when evaluated on the other participating sites' testing data.
Weakly supervised one-stage vision and language disease detection using large scale pneumonia and pneumothorax studies
Jul 31, 2020

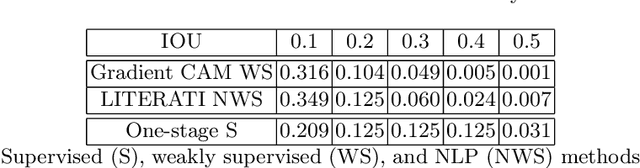

Abstract:Detecting clinically relevant objects in medical images is a challenge despite large datasets due to the lack of detailed labels. To address the label issue, we utilize the scene-level labels with a detection architecture that incorporates natural language information. We present a challenging new set of radiologist paired bounding box and natural language annotations on the publicly available MIMIC-CXR dataset especially focussed on pneumonia and pneumothorax. Along with the dataset, we present a joint vision language weakly supervised transformer layer-selected one-stage dual head detection architecture (LITERATI) alongside strong baseline comparisons with class activation mapping (CAM), gradient CAM, and relevant implementations on the NIH ChestXray-14 and MIMIC-CXR dataset. Borrowing from advances in vision language architectures, the LITERATI method demonstrates joint image and referring expression (objects localized in the image using natural language) input for detection that scales in a purely weakly supervised fashion. The architectural modifications address three obstacles -- implementing a supervised vision and language detection method in a weakly supervised fashion, incorporating clinical referring expression natural language information, and generating high fidelity detections with map probabilities. Nevertheless, the challenging clinical nature of the radiologist annotations including subtle references, multi-instance specifications, and relatively verbose underlying medical reports, ensures the vision language detection task at scale remains stimulating for future investigation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge