Swapnil Mishra
medR: Reward Engineering for Clinical Offline Reinforcement Learning via Tri-Drive Potential Functions
Feb 03, 2026Abstract:Reinforcement Learning (RL) offers a powerful framework for optimizing dynamic treatment regimes (DTRs). However, clinical RL is fundamentally bottlenecked by reward engineering: the challenge of defining signals that safely and effectively guide policy learning in complex, sparse offline environments. Existing approaches often rely on manual heuristics that fail to generalize across diverse pathologies. To address this, we propose an automated pipeline leveraging Large Language Models (LLMs) for offline reward design and verification. We formulate the reward function using potential functions consisted of three core components: survival, confidence, and competence. We further introduce quantitative metrics to rigorously evaluate and select the optimal reward structure prior to deployment. By integrating LLM-driven domain knowledge, our framework automates the design of reward functions for specific diseases while significantly enhancing the performance of the resulting policies.
A Foundation Model for Chest X-ray Interpretation with Grounded Reasoning via Online Reinforcement Learning
Sep 04, 2025Abstract:Medical foundation models (FMs) have shown tremendous promise amid the rapid advancements in artificial intelligence (AI) technologies. However, current medical FMs typically generate answers in a black-box manner, lacking transparent reasoning processes and locally grounded interpretability, which hinders their practical clinical deployments. To this end, we introduce DeepMedix-R1, a holistic medical FM for chest X-ray (CXR) interpretation. It leverages a sequential training pipeline: initially fine-tuned on curated CXR instruction data to equip with fundamental CXR interpretation capabilities, then exposed to high-quality synthetic reasoning samples to enable cold-start reasoning, and finally refined via online reinforcement learning to enhance both grounded reasoning quality and generation performance. Thus, the model produces both an answer and reasoning steps tied to the image's local regions for each query. Quantitative evaluation demonstrates substantial improvements in report generation (e.g., 14.54% and 31.32% over LLaVA-Rad and MedGemma) and visual question answering (e.g., 57.75% and 23.06% over MedGemma and CheXagent) tasks. To facilitate robust assessment, we propose Report Arena, a benchmarking framework using advanced language models to evaluate answer quality, further highlighting the superiority of DeepMedix-R1. Expert review of generated reasoning steps reveals greater interpretability and clinical plausibility compared to the established Qwen2.5-VL-7B model (0.7416 vs. 0.2584 overall preference). Collectively, our work advances medical FM development toward holistic, transparent, and clinically actionable modeling for CXR interpretation.
Uncertainty-Aware Regression for Socio-Economic Estimation via Multi-View Remote Sensing
Nov 21, 2024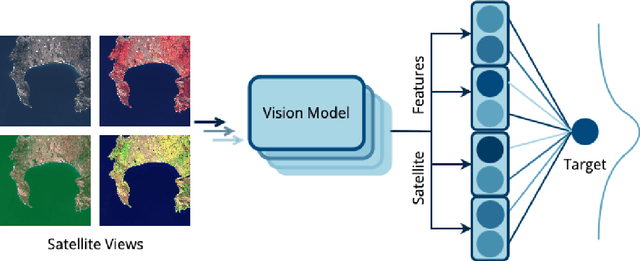
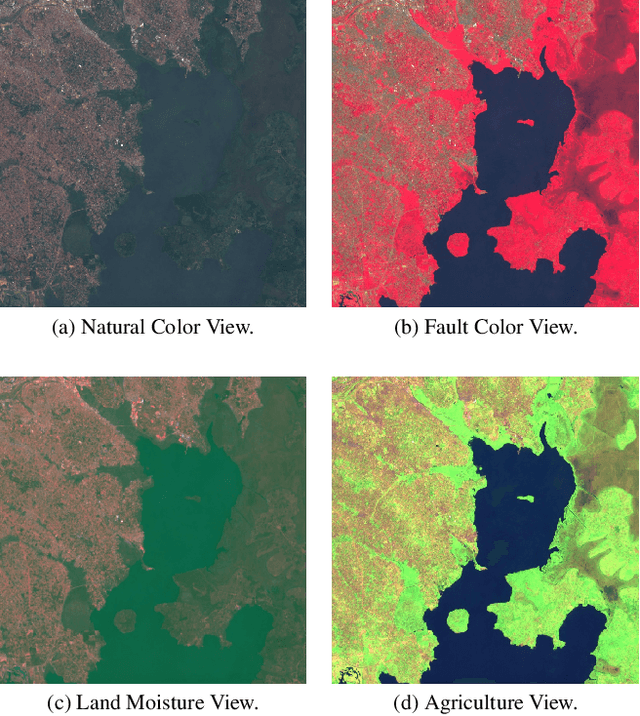


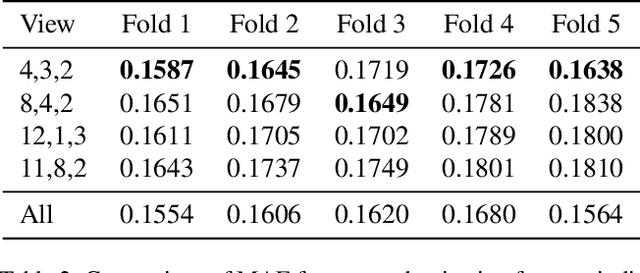
Abstract:Remote sensing imagery offers rich spectral data across extensive areas for Earth observation. Many attempts have been made to leverage these data with transfer learning to develop scalable alternatives for estimating socio-economic conditions, reducing reliance on expensive survey-collected data. However, much of this research has primarily focused on daytime satellite imagery due to the limitation that most pre-trained models are trained on 3-band RGB images. Consequently, modeling techniques for spectral bands beyond the visible spectrum have not been thoroughly investigated. Additionally, quantifying uncertainty in remote sensing regression has been less explored, yet it is essential for more informed targeting and iterative collection of ground truth survey data. In this paper, we introduce a novel framework that leverages generic foundational vision models to process remote sensing imagery using combinations of three spectral bands to exploit multi-spectral data. We also employ methods such as heteroscedastic regression and Bayesian modeling to generate uncertainty estimates for the predictions. Experimental results demonstrate that our method outperforms existing models that use RGB or multi-spectral models with unstructured band usage. Moreover, our framework helps identify uncertain predictions, guiding future ground truth data acquisition.
Evidential time-to-event prediction model with well-calibrated uncertainty estimation
Nov 12, 2024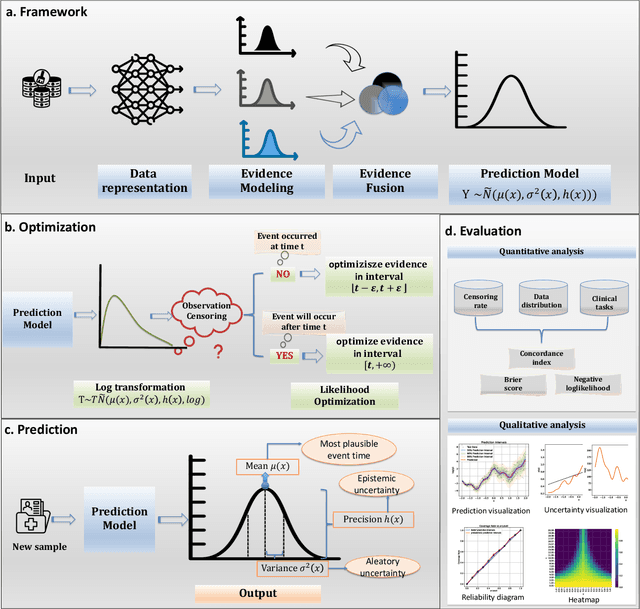

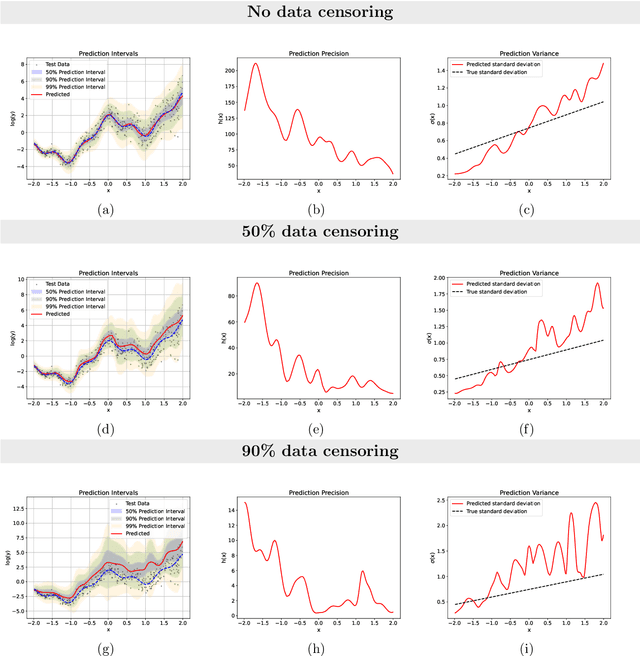
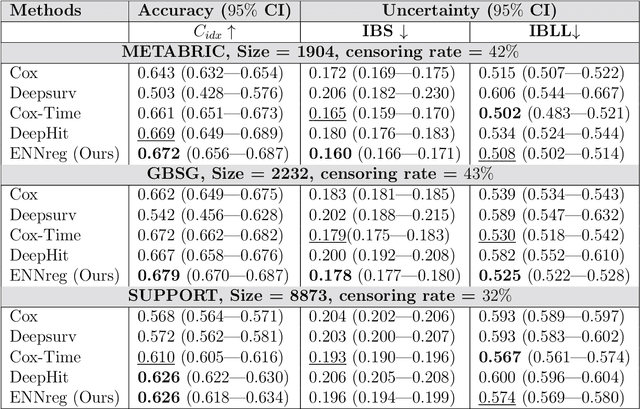
Abstract:Time-to-event analysis, or Survival analysis, provides valuable insights into clinical prognosis and treatment recommendations. However, this task is typically more challenging than other regression tasks due to the censored observations. Moreover, concerns regarding the reliability of predictions persist among clinicians, mainly attributed to the absence of confidence assessment, robustness, and calibration of prediction. To address those challenges, we introduce an evidential regression model designed especially for time-to-event prediction tasks, with which the most plausible event time, is directly quantified by aggregated Gaussian random fuzzy numbers (GRFNs). The GRFNs are a newly introduced family of random fuzzy subsets of the real line that generalizes both Gaussian random variables and Gaussian possibility distributions. Different from conventional methods that construct models based on strict data distribution, e.g., proportional hazard function, our model only assumes the event time is encoded in a real line GFRN without any strict distribution assumption, therefore offering more flexibility in complex data scenarios. Furthermore, the epistemic and aleatory uncertainty regarding the event time is quantified within the aggregated GRFN as well. Our model can, therefore, provide more detailed clinical decision-making guidance with two more degrees of information. The model is fit by minimizing a generalized negative log-likelihood function that accounts for data censoring based on uncertainty evidence reasoning. Experimental results on simulated datasets with varying data distributions and censoring scenarios, as well as on real-world datasets across diverse clinical settings and tasks, demonstrate that our model achieves both accurate and reliable performance, outperforming state-of-the-art methods.
KidSat: satellite imagery to map childhood poverty dataset and benchmark
Jul 08, 2024


Abstract:Satellite imagery has emerged as an important tool to analyse demographic, health, and development indicators. While various deep learning models have been built for these tasks, each is specific to a particular problem, with few standard benchmarks available. We propose a new dataset pairing satellite imagery and high-quality survey data on child poverty to benchmark satellite feature representations. Our dataset consists of 33,608 images, each 10 km $\times$ 10 km, from 19 countries in Eastern and Southern Africa in the time period 1997-2022. As defined by UNICEF, multidimensional child poverty covers six dimensions and it can be calculated from the face-to-face Demographic and Health Surveys (DHS) Program . As part of the benchmark, we test spatial as well as temporal generalization, by testing on unseen locations, and on data after the training years. Using our dataset we benchmark multiple models, from low-level satellite imagery models such as MOSAIKS , to deep learning foundation models, which include both generic vision models such as Self-Distillation with no Labels (DINOv2) models and specific satellite imagery models such as SatMAE. We provide open source code for building the satellite dataset, obtaining ground truth data from DHS and running various models assessed in our work.
Deep learning and MCMC with aggVAE for shifting administrative boundaries: mapping malaria prevalence in Kenya
May 31, 2023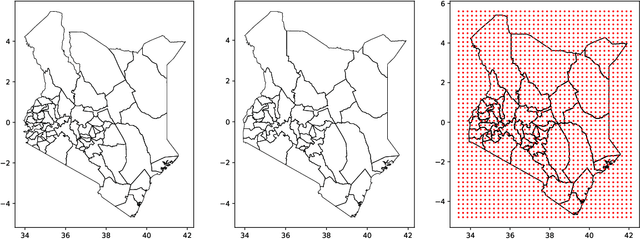
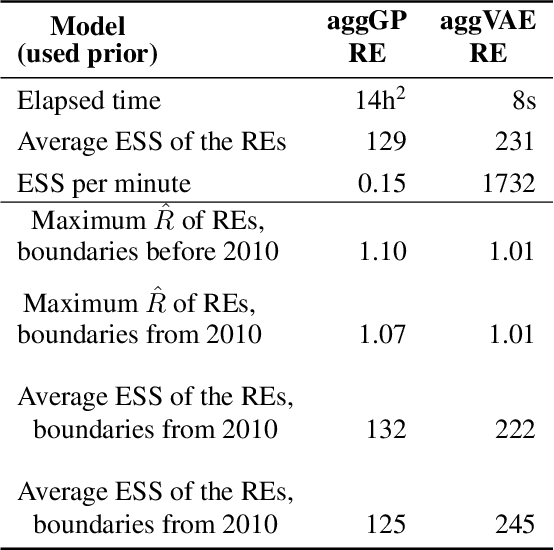
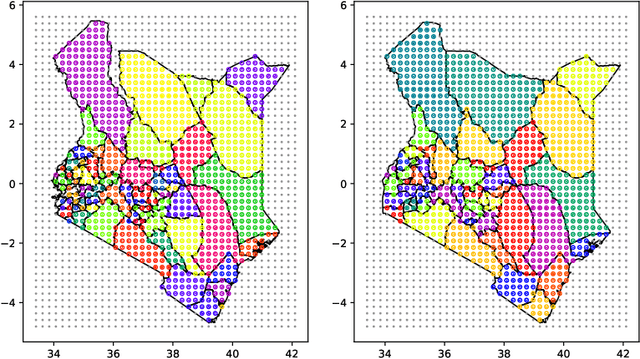

Abstract:Model-based disease mapping remains a fundamental policy-informing tool in public health and disease surveillance with hierarchical Bayesian models being the current state-of-the-art approach. When working with areal data, e.g. aggregates at the administrative unit level such as district or province, routinely used models rely on the adjacency structure of areal units to account for spatial correlations. The goal of disease surveillance systems is to track disease outcomes over time, but this provides challenging in situations of crises, such as political changes, leading to changes of administrative boundaries. Kenya is an example of such country. Moreover, adjacency-based approach ignores the continuous nature of spatial processes and cannot solve the change-of-support problem, i.e. when administrative boundaries change. We present a novel, practical, and easy to implement solution relying on a methodology combining deep generative modelling and fully Bayesian inference. We build on the recent work of PriorVAE able to encode spatial priors over small areas with variational autoencoders, to map malaria prevalence in Kenya. We solve the change-of-support problem arising from Kenya changing its district boundaries in 2010. We draw realisations of the Gaussian Process (GP) prior over a fine artificial spatial grid representing continuous space and then aggregate these realisations to the level of administrative boundaries. The aggregated values are then encoded using the PriorVAE technique. The trained priors (aggVAE) are then used at the inference stage instead of the GP priors within a Markov chain Monte Carlo (MCMC) scheme. We demonstrate that it is possible to use the flexible and appropriate model for areal data based on aggregation of continuous priors, and that inference is orders of magnitude faster when using aggVAE than combining the original GP priors and the aggregation step.
A comparison of short-term probabilistic forecasts for the incidence of COVID-19 using mechanistic and statistical time series models
May 01, 2023Abstract:Short-term forecasts of infectious disease spread are a critical component in risk evaluation and public health decision making. While different models for short-term forecasting have been developed, open questions about their relative performance remain. Here, we compare short-term probabilistic forecasts of popular mechanistic models based on the renewal equation with forecasts of statistical time series models. Our empirical comparison is based on data of the daily incidence of COVID-19 across six large US states over the first pandemic year. We find that, on average, probabilistic forecasts from statistical time series models are overall at least as accurate as forecasts from mechanistic models. Moreover, statistical time series models better capture volatility. Our findings suggest that domain knowledge, which is integrated into mechanistic models by making assumptions about disease dynamics, does not improve short-term forecasts of disease incidence. We note, however, that forecasting is often only one of many objectives and thus mechanistic models remain important, for example, to model the impact of vaccines or the emergence of new variants.
The interaction of transmission intensity, mortality, and the economy: a retrospective analysis of the COVID-19 pandemic
Oct 31, 2022



Abstract:The COVID-19 pandemic has caused over 6.4 million registered deaths to date, and has had a profound impact on economic activity. Here, we study the interaction of transmission, mortality, and the economy during the SARS-CoV-2 pandemic from January 2020 to December 2022 across 25 European countries. We adopt a Bayesian vector autoregressive model with both fixed and random effects. We find that increases in disease transmission intensity decreases Gross domestic product (GDP) and increases daily excess deaths, with a longer lasting impact on excess deaths in comparison to GDP, which recovers more rapidly. Broadly, our results reinforce the intuitive phenomenon that significant economic activity arises from diverse person-to-person interactions. We report on the effectiveness of non-pharmaceutical interventions (NPIs) on transmission intensity, excess deaths and changes in GDP, and resulting implications for policy makers. Our results highlight a complex cost-benefit trade off from individual NPIs. For example, banning international travel increases GDP however reduces excess deaths. We consider country random effects and their associations with excess changes in GDP and excess deaths. For example, more developed countries in Europe typically had more cautious approaches to the COVID-19 pandemic, prioritising healthcare and excess deaths over economic performance. Long term economic impairments are not fully captured by our model, as well as long term disease effects (Long Covid). Our results highlight that the impact of disease on a country is complex and multifaceted, and simple heuristic conclusions to extract the best outcome from the economy and disease burden are challenging.
Encoding spatiotemporal priors with VAEs for small-area estimation
Oct 20, 2021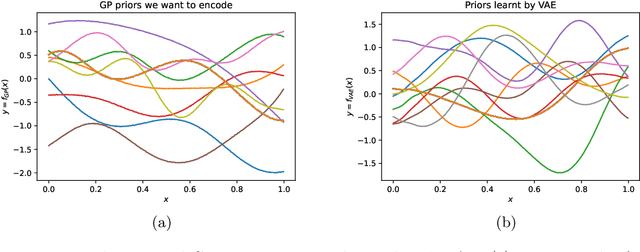
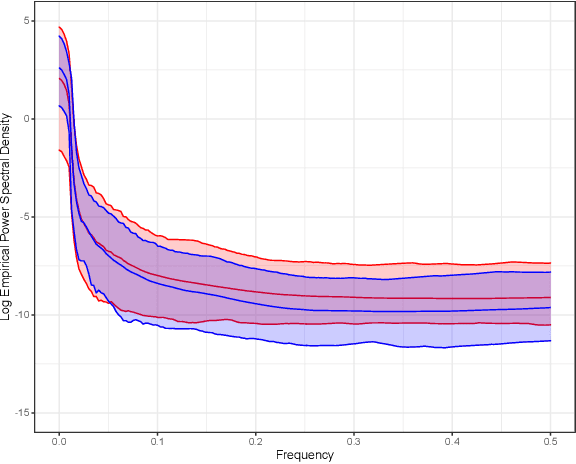
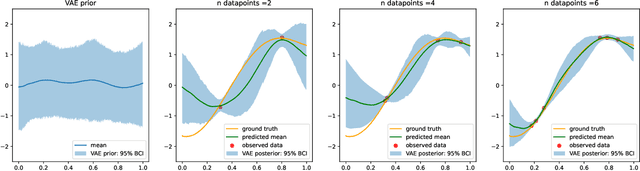
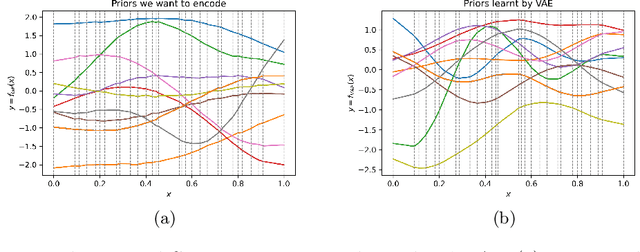
Abstract:Gaussian processes (GPs), implemented through multivariate Gaussian distributions for a finite collection of data, are the most popular approach in small-area spatiotemporal statistical modelling. In this context they are used to encode correlation structures over space and time and can generalise well in interpolation tasks. Despite their flexibility, off-the-shelf GPs present serious computational challenges which limit their scalability and practical usefulness in applied settings. Here, we propose a novel, deep generative modelling approach to tackle this challenge: for a particular spatiotemporal setting, we approximate a class of GP priors through prior sampling and subsequent fitting of a variational autoencoder (VAE). Given a trained VAE, the resultant decoder allows spatiotemporal inference to become incredibly efficient due to the low dimensional, independently distributed latent Gaussian space representation of the VAE. Once trained, inference using the VAE decoder replaces the GP within a Bayesian sampling framework. This approach provides tractable and easy-to-implement means of approximately encoding spatiotemporal priors and facilitates efficient statistical inference. We demonstrate the utility of our VAE two stage approach on Bayesian, small-area estimation tasks.
Gaussian Process Nowcasting: Application to COVID-19 Mortality Reporting
Feb 22, 2021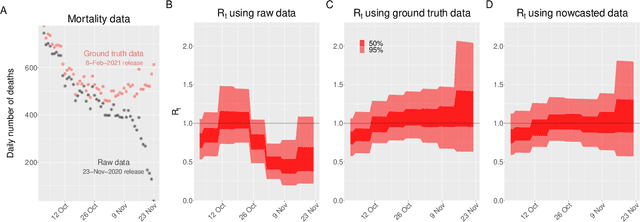
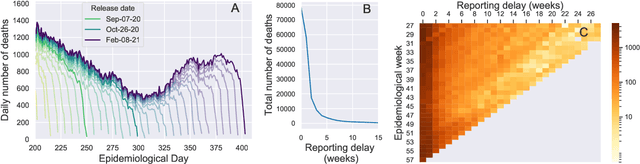
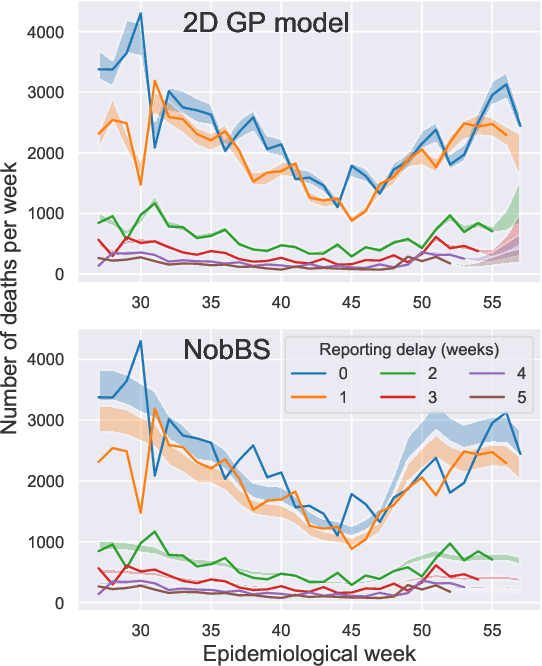
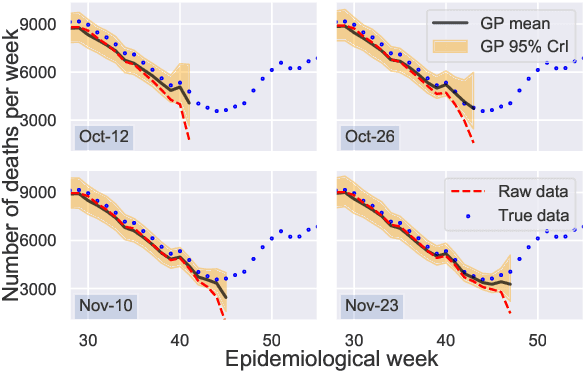
Abstract:Updating observations of a signal due to the delays in the measurement process is a common problem in signal processing, with prominent examples in a wide range of fields. An important example of this problem is the nowcasting of COVID-19 mortality: given a stream of reported counts of daily deaths, can we correct for the delays in reporting to paint an accurate picture of the present, with uncertainty? Without this correction, raw data will often mislead by suggesting an improving situation. We present a flexible approach using a latent Gaussian process that is capable of describing the changing auto-correlation structure present in the reporting time-delay surface. This approach also yields robust estimates of uncertainty for the estimated nowcasted numbers of deaths. We test assumptions in model specification such as the choice of kernel or hyper priors, and evaluate model performance on a challenging real dataset from Brazil. Our experiments show that Gaussian process nowcasting performs favourably against both comparable methods, and a small sample of expert human predictions. Our approach has substantial practical utility in disease modelling -- by applying our approach to COVID-19 mortality data from Brazil, where reporting delays are large, we can make informative predictions on important epidemiological quantities such as the current effective reproduction number.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge