Sandro Queirós
Point Tracking in Surgery--The 2024 Surgical Tattoos in Infrared (STIR) Challenge
Mar 31, 2025Abstract:Understanding tissue motion in surgery is crucial to enable applications in downstream tasks such as segmentation, 3D reconstruction, virtual tissue landmarking, autonomous probe-based scanning, and subtask autonomy. Labeled data are essential to enabling algorithms in these downstream tasks since they allow us to quantify and train algorithms. This paper introduces a point tracking challenge to address this, wherein participants can submit their algorithms for quantification. The submitted algorithms are evaluated using a dataset named surgical tattoos in infrared (STIR), with the challenge aptly named the STIR Challenge 2024. The STIR Challenge 2024 comprises two quantitative components: accuracy and efficiency. The accuracy component tests the accuracy of algorithms on in vivo and ex vivo sequences. The efficiency component tests the latency of algorithm inference. The challenge was conducted as a part of MICCAI EndoVis 2024. In this challenge, we had 8 total teams, with 4 teams submitting before and 4 submitting after challenge day. This paper details the STIR Challenge 2024, which serves to move the field towards more accurate and efficient algorithms for spatial understanding in surgery. In this paper we summarize the design, submissions, and results from the challenge. The challenge dataset is available here: https://zenodo.org/records/14803158 , and the code for baseline models and metric calculation is available here: https://github.com/athaddius/STIRMetrics
Exploring Optical Flow Inclusion into nnU-Net Framework for Surgical Instrument Segmentation
Mar 15, 2024Abstract:Surgical instrument segmentation in laparoscopy is essential for computer-assisted surgical systems. Despite the Deep Learning progress in recent years, the dynamic setting of laparoscopic surgery still presents challenges for precise segmentation. The nnU-Net framework excelled in semantic segmentation analyzing single frames without temporal information. The framework's ease of use, including its ability to be automatically configured, and its low expertise requirements, have made it a popular base framework for comparisons. Optical flow (OF) is a tool commonly used in video tasks to estimate motion and represent it in a single frame, containing temporal information. This work seeks to employ OF maps as an additional input to the nnU-Net architecture to improve its performance in the surgical instrument segmentation task, taking advantage of the fact that instruments are the main moving objects in the surgical field. With this new input, the temporal component would be indirectly added without modifying the architecture. Using CholecSeg8k dataset, three different representations of movement were estimated and used as new inputs, comparing them with a baseline model. Results showed that the use of OF maps improves the detection of classes with high movement, even when these are scarce in the dataset. To further improve performance, future work may focus on implementing other OF-preserving augmentations.
Deep Spatiotemporal Clutter Filtering of Transthoracic Echocardiographic Images Using a 3D Convolutional Auto-Encoder
Jan 23, 2024

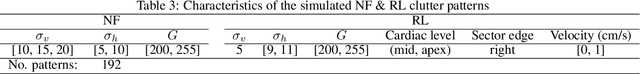

Abstract:This study presents a deep convolutional auto-encoder network for filtering reverberation artifacts, from transthoracic echocardiographic (TTE) image sequences. Given the spatiotemporal nature of these artifacts, the filtering network was built using 3D convolutional layers to suppress the clutter patterns throughout the cardiac cycle. The network was designed by taking advantage of: i) an attention mechanism to focus primarily on cluttered regions and ii) residual learning to preserve fine structures of the image frames. To train the deep network, a diverse set of artifact patterns was simulated and the simulated patterns were superimposed onto artifact-free ultra-realistic synthetic TTE sequences of six ultrasound vendors to generate input of the filtering network. The artifact-free sequences served as ground-truth. Performance of the filtering network was evaluated using unseen synthetic as well as in-vivo artifactual sequences. Satisfactory results obtained using the latter dataset confirmed the good generalization performance of the proposed network which was trained using the synthetic sequences and simulated artifact patterns. Suitability of the clutter-filtered sequences for further processing was assessed by computing segmental strain curves from them. The results showed that the large discrepancy between the strain profiles computed from the cluttered segments and their corresponding segments in the clutter-free images was significantly reduced after filtering the sequences using the proposed network. The trained deep network could process an artifactual TTE sequence in a fraction of a second and can be used for real-time clutter filtering. Moreover, it can improve the precision of the clinical indexes that are computed from the TTE sequences. The source code of the proposed method is available at: https://github.com/MahdiTabassian/Deep-Clutter-Filtering/tree/main.
High-Resolution Maps of Left Atrial Displacements and Strains Estimated with 3D CINE MRI and Unsupervised Neural Networks
Dec 14, 2023



Abstract:The functional analysis of the left atrium (LA) is important for evaluating cardiac health and understanding diseases like atrial fibrillation. Cine MRI is ideally placed for the detailed 3D characterisation of LA motion and deformation, but it is lacking appropriate acquisition and analysis tools. In this paper, we present Analysis for Left Atrial Displacements and Deformations using unsupervIsed neural Networks, \textit{Aladdin}, to automatically and reliably characterise regional LA deformations from high-resolution 3D Cine MRI. The tool includes: an online few-shot segmentation network (Aladdin-S), an online unsupervised image registration network (Aladdin-R), and a strain calculations pipeline tailored to the LA. We create maps of LA Displacement Vector Field (DVF) magnitude and LA principal strain values from images of 10 healthy volunteers and 8 patients with cardiovascular disease (CVD). We additionally create an atlas of these biomarkers using the data from the healthy volunteers. Aladdin is able to accurately track the LA wall across the cardiac cycle and characterize its motion and deformation. The overall DVF magnitude and principal strain values are significantly higher in the healthy group vs CVD patients: $2.85 \pm 1.59~mm$ and $0.09 \pm 0.05$ vs $1.96 \pm 0.74~mm$ and $0.03 \pm 0.04$, respectively. The time course of these metrics is also different in the two groups, with a more marked active contraction phase observed in the healthy cohort. Finally, utilizing the LA atlas allows us to identify regional deviations from the population distribution that may indicate focal tissue abnormalities. The proposed tool for the quantification of novel regional LA deformation biomarkers should have important clinical applications. The source code, anonymized images, generated maps and atlas are publicly available: https://github.com/cgalaz01/aladdin_cmr_la.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge