Nagesh Adluru
Stroke Lesion Segmentation using Multi-Stage Cross-Scale Attention
Jan 26, 2025
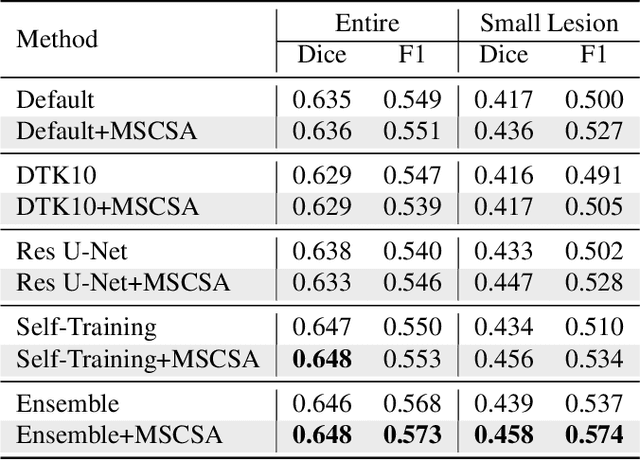
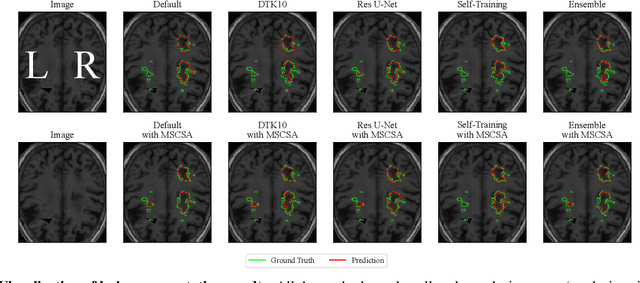
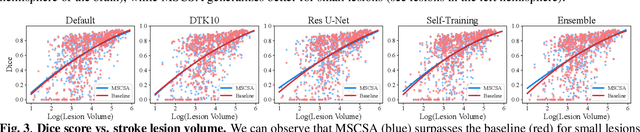
Abstract:Precise characterization of stroke lesions from MRI data has immense value in prognosticating clinical and cognitive outcomes following a stroke. Manual stroke lesion segmentation is time-consuming and requires the expertise of neurologists and neuroradiologists. Often, lesions are grossly characterized for their location and overall extent using bounding boxes without specific delineation of their boundaries. While such characterization provides some clinical value, to develop a precise mechanistic understanding of the impact of lesions on post-stroke vascular contributions to cognitive impairments and dementia (VCID), the stroke lesions need to be fully segmented with accurate boundaries. This work introduces the Multi-Stage Cross-Scale Attention (MSCSA) mechanism, applied to the U-Net family, to improve the mapping between brain structural features and lesions of varying sizes. Using the Anatomical Tracings of Lesions After Stroke (ATLAS) v2.0 dataset, MSCSA outperforms all baseline methods in both Dice and F1 scores on a subset focusing on small lesions, while maintaining competitive performance across the entire dataset. Notably, the ensemble strategy incorporating MSCSA achieves the highest scores for Dice and F1 on both the full dataset and the small lesion subset. These results demonstrate the effectiveness of MSCSA in segmenting small lesions and highlight its robustness across different training schemes for large stroke lesions. Our code is available at: https://github.com/nadluru/StrokeLesSeg.
MICCAI-CDMRI 2023 QuantConn Challenge Findings on Achieving Robust Quantitative Connectivity through Harmonized Preprocessing of Diffusion MRI
Nov 14, 2024



Abstract:White matter alterations are increasingly implicated in neurological diseases and their progression. International-scale studies use diffusion-weighted magnetic resonance imaging (DW-MRI) to qualitatively identify changes in white matter microstructure and connectivity. Yet, quantitative analysis of DW-MRI data is hindered by inconsistencies stemming from varying acquisition protocols. There is a pressing need to harmonize the preprocessing of DW-MRI datasets to ensure the derivation of robust quantitative diffusion metrics across acquisitions. In the MICCAI-CDMRI 2023 QuantConn challenge, participants were provided raw data from the same individuals collected on the same scanner but with two different acquisitions and tasked with preprocessing the DW-MRI to minimize acquisition differences while retaining biological variation. Submissions are evaluated on the reproducibility and comparability of cross-acquisition bundle-wise microstructure measures, bundle shape features, and connectomics. The key innovations of the QuantConn challenge are that (1) we assess bundles and tractography in the context of harmonization for the first time, (2) we assess connectomics in the context of harmonization for the first time, and (3) we have 10x additional subjects over prior harmonization challenge, MUSHAC and 100x over SuperMUDI. We find that bundle surface area, fractional anisotropy, connectome assortativity, betweenness centrality, edge count, modularity, nodal strength, and participation coefficient measures are most biased by acquisition and that machine learning voxel-wise correction, RISH mapping, and NeSH methods effectively reduce these biases. In addition, microstructure measures AD, MD, RD, bundle length, connectome density, efficiency, and path length are least biased by these acquisition differences.
* Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://melba-journal.org/2024/019
Segmenting Small Stroke Lesions with Novel Labeling Strategies
Aug 06, 2024



Abstract:Deep neural networks have demonstrated exceptional efficacy in stroke lesion segmentation. However, the delineation of small lesions, critical for stroke diagnosis, remains a challenge. In this study, we propose two straightforward yet powerful approaches that can be seamlessly integrated into a variety of networks: Multi-Size Labeling (MSL) and Distance-Based Labeling (DBL), with the aim of enhancing the segmentation accuracy of small lesions. MSL divides lesion masks into various categories based on lesion volume while DBL emphasizes the lesion boundaries. Experimental evaluations on the Anatomical Tracings of Lesions After Stroke (ATLAS) v2.0 dataset showcase that an ensemble of MSL and DBL achieves consistently better or equal performance on recall (3.6% and 3.7%), F1 (2.4% and 1.5%), and Dice scores (1.3% and 0.0%) compared to the top-1 winner of the 2022 MICCAI ATLAS Challenge on both the subset only containing small lesions and the entire dataset, respectively. Notably, on the mini-lesion subset, a single MSL model surpasses the previous best ensemble strategy, with enhancements of 1.0% and 0.3% on F1 and Dice scores, respectively. Our code is available at: https://github.com/nadluru/StrokeLesSeg.
Vision Backbone Enhancement via Multi-Stage Cross-Scale Attention
Aug 14, 2023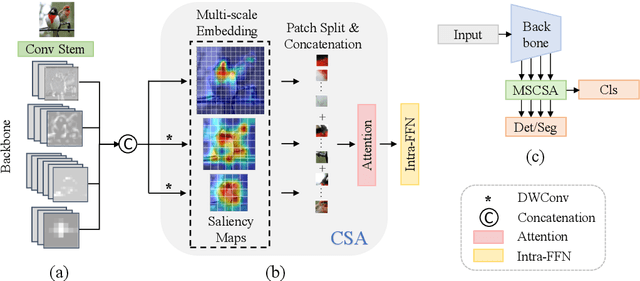



Abstract:Convolutional neural networks (CNNs) and vision transformers (ViTs) have achieved remarkable success in various vision tasks. However, many architectures do not consider interactions between feature maps from different stages and scales, which may limit their performance. In this work, we propose a simple add-on attention module to overcome these limitations via multi-stage and cross-scale interactions. Specifically, the proposed Multi-Stage Cross-Scale Attention (MSCSA) module takes feature maps from different stages to enable multi-stage interactions and achieves cross-scale interactions by computing self-attention at different scales based on the multi-stage feature maps. Our experiments on several downstream tasks show that MSCSA provides a significant performance boost with modest additional FLOPs and runtime.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Accurate Automatic Segmentation of Amygdala Subnuclei and Modeling of Uncertainty via Bayesian Fully Convolutional Neural Network
Feb 19, 2019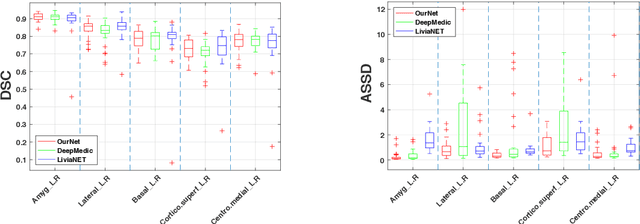
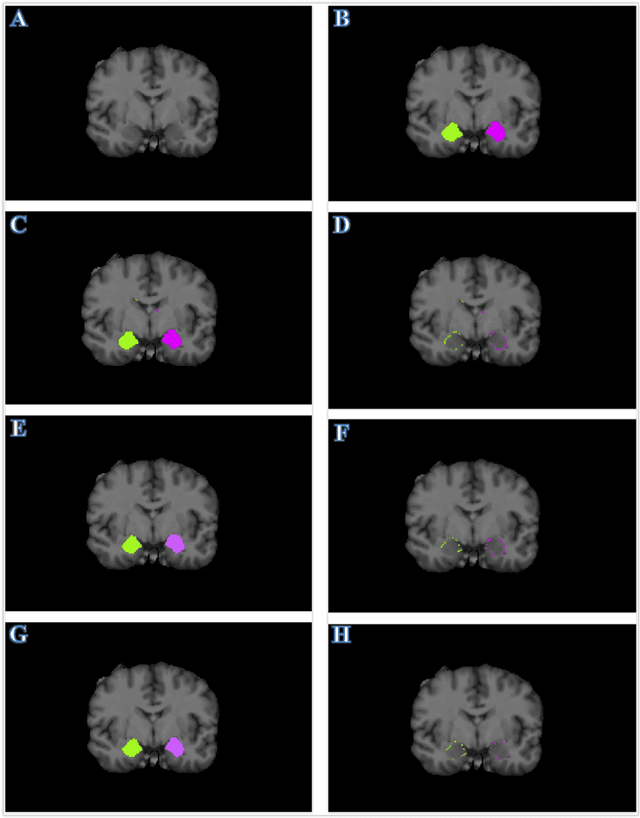
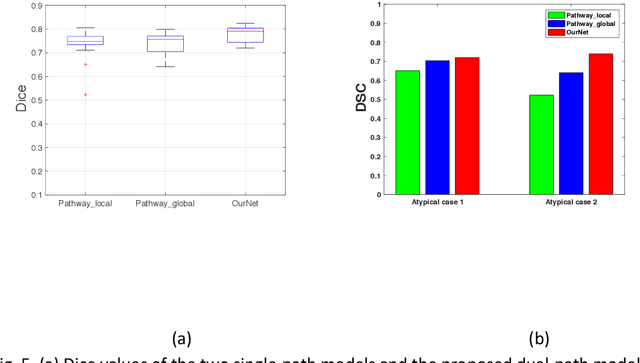
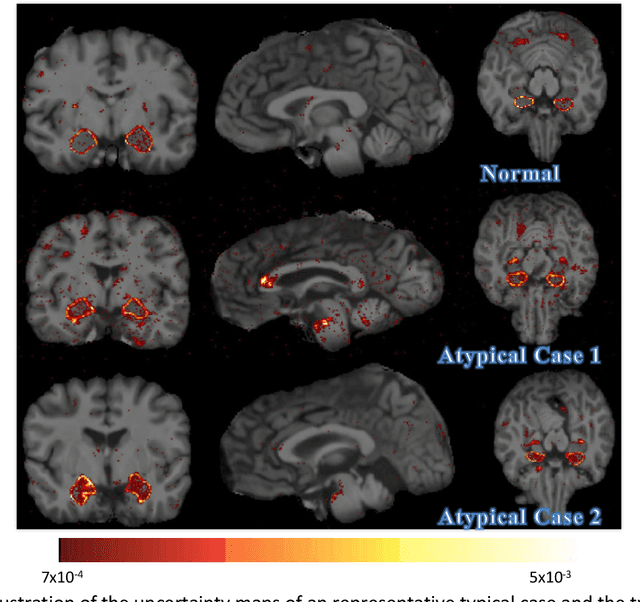
Abstract:Recent advances in deep learning have improved the segmentation accuracy of subcortical brain structures, which would be useful in neuroimaging studies of many neurological disorders. However, most of the previous deep learning work does not investigate the specific difficulties that exist in segmenting extremely small but important brain regions such as the amygdala and its subregions. To tackle this challenging task, a novel 3D Bayesian fully convolutional neural network was developed to apply a dilated dualpathway approach that retains fine details and utilizes both local and more global contextual information to automatically segment the amygdala and its subregions at high precision. The proposed method provides insights on network design and sampling strategy that target segmentations of small 3D structures. In particular, this study confirms that a large context, enabled by a large field of view, is beneficial for segmenting small objects; furthermore, precise contextual information enabled by dilated convolutions allows for better boundary localization, which is critical for examining the morphology of the structure. In addition, it is demonstrated that the uncertainty information estimated from our network may be leveraged to identify atypicality in data. Our method was compared with two state-of-the-art deep learning models and a traditional multi-atlas approach, and exhibited excellent performance as measured both by Dice overlap as well as average symmetric surface distance. To the best of our knowledge, this work is the first deep learning-based approach that targets the subregions of the amygdala.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge