Mauricio A. Álvarez
Cross-Granularity Representations for Biological Sequences: Insights from ESM and BiGCARP
Mar 21, 2026Abstract:Recent advances in general-purpose foundation models have stimulated the development of large biological sequence models. While natural language shows symbolic granularity (characters, words, sentences), biological sequences exhibit hierarchical granularity whose levels (nucleotides, amino acids, protein domains, genes) further encode biologically functional information. In this paper, we investigate the integration of cross-granularity knowledge from models through a case study of BiGCARP, a Pfam domain-level model for biosynthetic gene clusters, and ESM, an amino acid-level protein language model. Using representation analysis tools and a set of probe tasks, we first explain why a straightforward cross-model embedding initialization fails to improve downstream performance in BiGCARP, and show that deeper-layer embeddings capture a more contextual and faithful representation of the model's learned knowledge. Furthermore, we demonstrate that representations at different granularities encode complementary biological knowledge, and that combining them yields measurable performance gains in intermediate-level prediction tasks. Our findings highlight cross-granularity integration as a promising strategy for improving both the performance and interpretability of biological foundation models.
* 9 pages, 4 figures, published in 2025 IEEE International Conference on Bioinformatics and Biomedicine (BIBM)
Disentangling Similarity and Relatedness in Topic Models
Mar 11, 2026Abstract:The recent advancement of large language models has spurred a growing trend of integrating pre-trained language model (PLM) embeddings into topic models, fundamentally reshaping how topics capture semantic structure. Classical models such as Latent Dirichlet Allocation (LDA) derive topics from word co-occurrence statistics, whereas PLM-augmented models anchor these statistics to pre-trained embedding spaces, imposing a prior that also favours clustering of semantically similar words. This structural difference can be captured by the psycholinguistic dimensions of thematic relatedness and taxonomic similarity of the topic words. To disentangle these dimensions in topic models, we construct a large synthetic benchmark of word pairs using LLM-based annotation to train a neural scoring function. We apply this scorer to a comprehensive evaluation across multiple corpora and topic model families, revealing that different model families capture distinct semantic structure in their topics. We further demonstrate that similarity and relatedness scores successfully predict downstream task performance depending on task requirements. This paper establishes similarity and relatedness as essential axes for topic model evaluation and provides a reliable pipeline for characterising these across model families and corpora.
Counterfactual Credit Guided Bayesian Optimization
Oct 06, 2025Abstract:Bayesian optimization has emerged as a prominent methodology for optimizing expensive black-box functions by leveraging Gaussian process surrogates, which focus on capturing the global characteristics of the objective function. However, in numerous practical scenarios, the primary objective is not to construct an exhaustive global surrogate, but rather to quickly pinpoint the global optimum. Due to the aleatoric nature of the sequential optimization problem and its dependence on the quality of the surrogate model and the initial design, it is restrictive to assume that all observed samples contribute equally to the discovery of the optimum in this context. In this paper, we introduce Counterfactual Credit Guided Bayesian Optimization (CCGBO), a novel framework that explicitly quantifies the contribution of individual historical observations through counterfactual credit. By incorporating counterfactual credit into the acquisition function, our approach can selectively allocate resources in areas where optimal solutions are most likely to occur. We prove that CCGBO retains sublinear regret. Empirical evaluations on various synthetic and real-world benchmarks demonstrate that CCGBO consistently reduces simple regret and accelerates convergence to the global optimum.
Neighbour-Driven Gaussian Process Variational Autoencoders for Scalable Structured Latent Modelling
May 22, 2025Abstract:Gaussian Process (GP) Variational Autoencoders (VAEs) extend standard VAEs by replacing the fully factorised Gaussian prior with a GP prior, thereby capturing richer correlations among latent variables. However, performing exact GP inference in large-scale GPVAEs is computationally prohibitive, often forcing existing approaches to rely on restrictive kernel assumptions or large sets of inducing points. In this work, we propose a neighbour-driven approximation strategy that exploits local adjacencies in the latent space to achieve scalable GPVAE inference. By confining computations to the nearest neighbours of each data point, our method preserves essential latent dependencies, allowing more flexible kernel choices and mitigating the need for numerous inducing points. Through extensive experiments on tasks including representation learning, data imputation, and conditional generation, we demonstrate that our approach outperforms other GPVAE variants in both predictive performance and computational efficiency.
Adaptive RKHS Fourier Features for Compositional Gaussian Process Models
Jul 01, 2024Abstract:Deep Gaussian Processes (DGPs) leverage a compositional structure to model non-stationary processes. DGPs typically rely on local inducing point approximations across intermediate GP layers. Recent advances in DGP inference have shown that incorporating global Fourier features from Reproducing Kernel Hilbert Space (RKHS) can enhance the DGPs' capability to capture complex non-stationary patterns. This paper extends the use of these features to compositional GPs involving linear transformations. In particular, we introduce Ordinary Differential Equation (ODE) -based RKHS Fourier features that allow for adaptive amplitude and phase modulation through convolution operations. This convolutional formulation relates our work to recently proposed deep latent force models, a multi-layer structure designed for modelling nonlinear dynamical systems. By embedding these adjustable RKHS Fourier features within a doubly stochastic variational inference framework, our model exhibits improved predictive performance across various regression tasks.
Deep Latent Force Models: ODE-based Process Convolutions for Bayesian Deep Learning
Nov 24, 2023Abstract:Effectively modeling phenomena present in highly nonlinear dynamical systems whilst also accurately quantifying uncertainty is a challenging task, which often requires problem-specific techniques. We outline the deep latent force model (DLFM), a domain-agnostic approach to tackling this problem, which consists of a deep Gaussian process architecture where the kernel at each layer is derived from an ordinary differential equation using the framework of process convolutions. Two distinct formulations of the DLFM are presented which utilise weight-space and variational inducing points-based Gaussian process approximations, both of which are amenable to doubly stochastic variational inference. We provide evidence that our model is capable of capturing highly nonlinear behaviour in real-world multivariate time series data. In addition, we find that our approach achieves comparable performance to a number of other probabilistic models on benchmark regression tasks. We also empirically assess the negative impact of the inducing points framework on the extrapolation capabilities of LFM-based models.
Thin and Deep Gaussian Processes
Oct 17, 2023Abstract:Gaussian processes (GPs) can provide a principled approach to uncertainty quantification with easy-to-interpret kernel hyperparameters, such as the lengthscale, which controls the correlation distance of function values. However, selecting an appropriate kernel can be challenging. Deep GPs avoid manual kernel engineering by successively parameterizing kernels with GP layers, allowing them to learn low-dimensional embeddings of the inputs that explain the output data. Following the architecture of deep neural networks, the most common deep GPs warp the input space layer-by-layer but lose all the interpretability of shallow GPs. An alternative construction is to successively parameterize the lengthscale of a kernel, improving the interpretability but ultimately giving away the notion of learning lower-dimensional embeddings. Unfortunately, both methods are susceptible to particular pathologies which may hinder fitting and limit their interpretability. This work proposes a novel synthesis of both previous approaches: Thin and Deep GP (TDGP). Each TDGP layer defines locally linear transformations of the original input data maintaining the concept of latent embeddings while also retaining the interpretation of lengthscales of a kernel. Moreover, unlike the prior solutions, TDGP induces non-pathological manifolds that admit learning lower-dimensional representations. We show with theoretical and experimental results that i) TDGP is, unlike previous models, tailored to specifically discover lower-dimensional manifolds in the input data, ii) TDGP behaves well when increasing the number of layers, and iii) TDGP performs well in standard benchmark datasets.
Shallow and Deep Nonparametric Convolutions for Gaussian Processes
Jun 17, 2022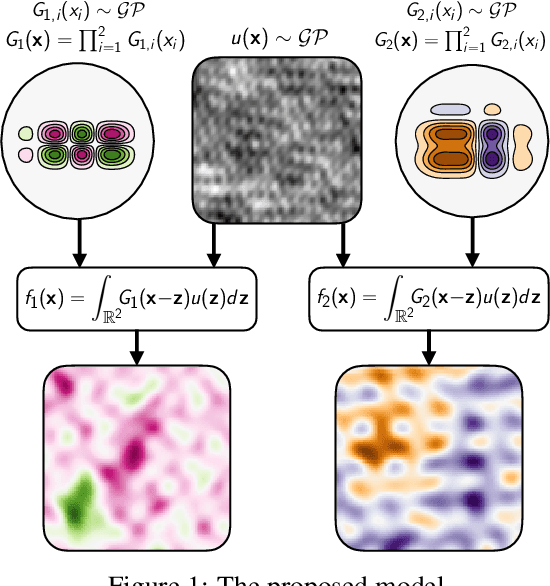
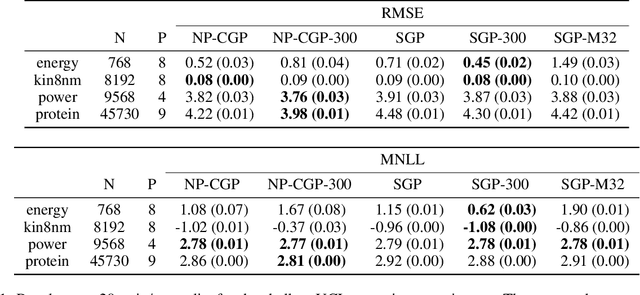
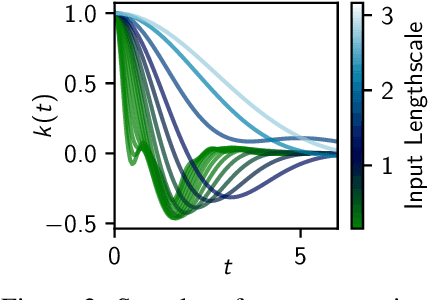

Abstract:A key challenge in the practical application of Gaussian processes (GPs) is selecting a proper covariance function. The moving average, or process convolutions, construction of GPs allows some additional flexibility, but still requires choosing a proper smoothing kernel, which is non-trivial. Previous approaches have built covariance functions by using GP priors over the smoothing kernel, and by extension the covariance, as a way to bypass the need to specify it in advance. However, such models have been limited in several ways: they are restricted to single dimensional inputs, e.g. time; they only allow modelling of single outputs and they do not scale to large datasets since inference is not straightforward. In this paper, we introduce a nonparametric process convolution formulation for GPs that alleviates these weaknesses by using a functional sampling approach based on Matheron's rule to perform fast sampling using interdomain inducing variables. Furthermore, we propose a composition of these nonparametric convolutions that serves as an alternative to classic deep GP models, and allows the covariance functions of the intermediate layers to be inferred from the data. We test the performance of our model on benchmarks for single output GPs, multiple output GPs and deep GPs and find that in many cases our approach can provide improvements over standard GP models.
Angular Super-Resolution in Diffusion MRI with a 3D Recurrent Convolutional Autoencoder
Mar 29, 2022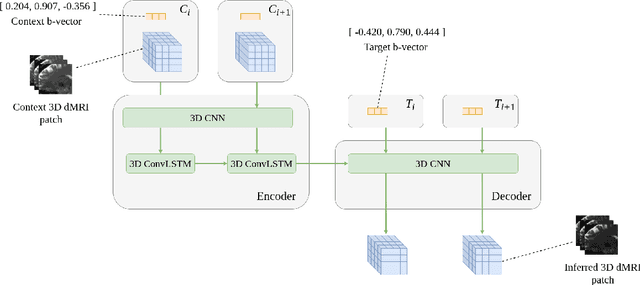
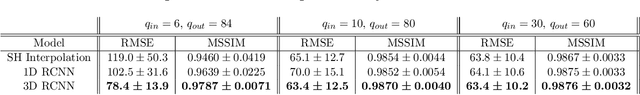

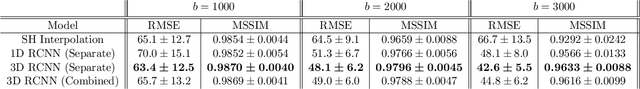
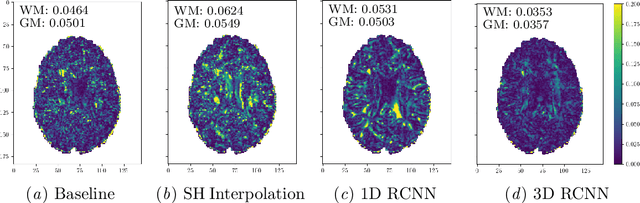
Abstract:High resolution diffusion MRI (dMRI) data is often constrained by limited scanning time in clinical settings, thus restricting the use of downstream analysis techniques that would otherwise be available. In this work we develop a 3D recurrent convolutional neural network (RCNN) capable of super-resolving dMRI volumes in the angular (q-space) domain. Our approach formulates the task of angular super-resolution as a patch-wise regression using a 3D autoencoder conditioned on target b-vectors. Within the network we use a convolutional long short term memory (ConvLSTM) cell to model the relationship between q-space samples. We compare model performance against a baseline spherical harmonic interpolation and a 1D variant of the model architecture. We show that the 3D model has the lowest error rates across different subsampling schemes and b-values. The relative performance of the 3D RCNN is greatest in the very low angular resolution domain. Code for this project is available at https://github.com/m-lyon/dMRI-RCNN.
Modular Gaussian Processes for Transfer Learning
Oct 26, 2021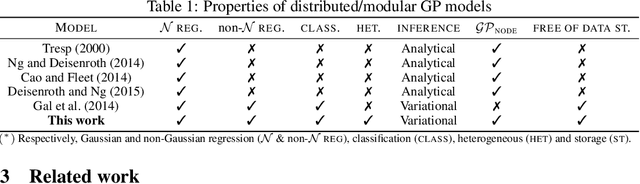
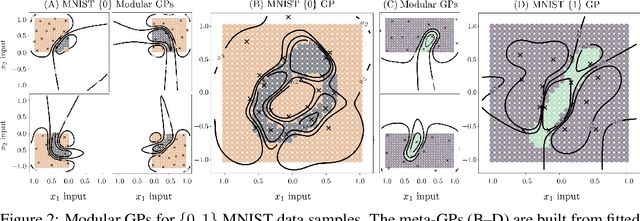
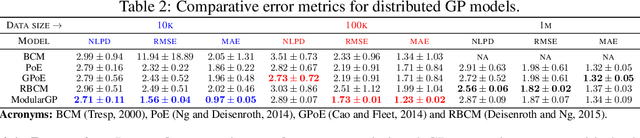
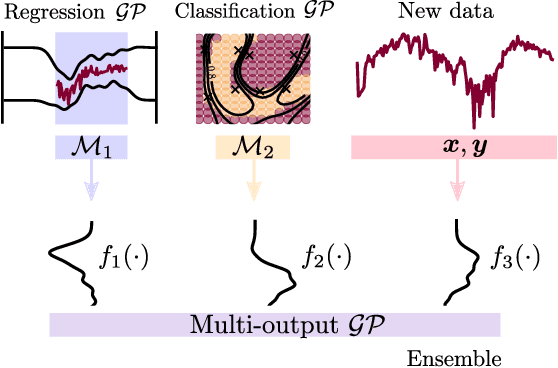
Abstract:We present a framework for transfer learning based on modular variational Gaussian processes (GP). We develop a module-based method that having a dictionary of well fitted GPs, one could build ensemble GP models without revisiting any data. Each model is characterised by its hyperparameters, pseudo-inputs and their corresponding posterior densities. Our method avoids undesired data centralisation, reduces rising computational costs and allows the transfer of learned uncertainty metrics after training. We exploit the augmentation of high-dimensional integral operators based on the Kullback-Leibler divergence between stochastic processes to introduce an efficient lower bound under all the sparse variational GPs, with different complexity and even likelihood distribution. The method is also valid for multi-output GPs, learning correlations a posteriori between independent modules. Extensive results illustrate the usability of our framework in large-scale and multi-task experiments, also compared with the exact inference methods in the literature.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge